Abstract
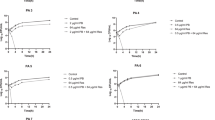
Bacterial infections caused by multidrug-resistant (MDR) gram-negative strains carrying the mobile colistin resistance gene mcr-1 are serious threats to world public health due to the lack of effective treatments. Inhibition of the ATP synthase makes bacteria such as Staphylococcus aureus and Klebsiella pneumoniae more sensitive to polymyxin. This provides new strategies for treating infections caused by polymyxins-resistant bacteria carrying mcr-1. Six mcr-1-positive strains were isolated from clinical samples, and all were identified as Escherichia coli. Here we investigated several ATP synthase inhibitors, N,N’-dicyclohexylcarbodiimide (DCCD), resveratrol, and piceatannol, for their antibacterial effects against the mcr-1-positive strains combined with polymyxin B (POL). Checkerboard assay, time-kill assay, biofilm inhibition and eradication assay indicated the significant synergistic effect of ATP synthase inhibitors/POL combination in vitro. Meanwhile, mouse infection model experiment was also performed, showing a 5 log10 reduction of the pathogen after treatment with the resveratrol/POL combination. Moreover, adding adenosine disodium triphosphate (Na2ATP) could inhibit the antibacterial effect of the ATP synthase inhibitors/POL combination. In conclusion, our study confirmed that inhibition of ATP production could increase the susceptibility of bacteria carrying mcr-1 to polymyxins. This provides a new strategy against polymyxins-resistant bacteria infection.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The datasets used and analyzed during the current study are available from the corresponding author upon reasonable request. Most of the data is included in this published article.
References
Domingues MM, Inacio RG, Raimundo JM, Martins M, Castanho MA, Santos NC. Biophysical characterization of polymyxin B interaction with LPS aggregates and membrane model systems. Biopolymers. 2012;98:338–44. https://doi.org/10.1002/bip.22095.
Moffatt JH, Harper M, Boyce JD. Mechanisms of polymyxin resistance. Adv Exp Med Biol. 2019;1145:55–71. https://doi.org/10.1007/978-3-030-16373-0_5.
Falagas ME, Kasiakou SK. Toxicity of polymyxins: a systematic review of the evidence from old and recent studies. Crit Care. 2006;10:R27. https://doi.org/10.1186/cc3995.
Xiaomin S, Yiming L, Yuying Y, Zhangqi S, Yongning W, Shaolin W. Global impact of mcr-1-positive Enterobacteriaceae bacteria on "one health". Crit Rev Microbiol. 2020;46:565–77. https://doi.org/10.1080/1040841X.2020.1812510.
Nang SC, Li J, Velkov T. The rise and spread of mcr plasmid-mediated polymyxin resistance. Crit Rev Microbiol. 2019;45:131–61. https://doi.org/10.1080/1040841X.2018.1492902.
Olaitan AO, Morand S, Rolain JM. Mechanisms of polymyxin resistance: acquired and intrinsic resistance in bacteria. Front Microbiol. 2014;5:643. https://doi.org/10.3389/fmicb.2014.00643.
Baron S, Hadjadj L, Rolain JM, Olaitan AO. Molecular mechanisms of polymyxin resistance: knowns and unknowns. Int J Antimicrob Agents. 2016;48:583–91. https://doi.org/10.1016/j.ijantimicag.2016.06.023.
Liu YY, Wang Y, Walsh TR, Yi LX, Zhang R, Spencer J, Doi Y, Tian G, Dong B, Huang X, et al. Emergence of plasmid-mediated colistin resistance mechanism MCR-1 in animals and human beings in China: a microbiological and molecular biological study. Lancet Infect Dis. 2016;16:161–8. https://doi.org/10.1016/S1473-3099(15)00424-7.
Hussein NH, Al-Kadmy I, Taha BM, Hussein JD. Mobilized colistin resistance (mcr) genes from 1 to 10: a comprehensive review. Mol Biol Rep. 2021;48:2897–907. https://doi.org/10.1007/s11033-021-06307-y.
Vestergaard M, Bald D, Ingmer H. Targeting the ATP synthase in bacterial and fungal pathogens: beyond Mycobacterium tuberculosis. J Glob Antimicrob Resist. 2022;29:29–41. https://doi.org/10.1016/j.jgar.2022.01.026.
Chan B, Khadem TM, Brown J. A review of tuberculosis: focus on bedaquiline. Am J Health Syst Pharm. 2013;70:1984–94. https://doi.org/10.2146/ajhp130199.
Vestergaard M, Nohr-Meldgaard K, Bojer MS, Krogsgard NC, Meyer RL, Slavetinsky C, Peschel A, Ingmer H. Inhibition of the ATP synthase eliminates the intrinsic resistance of Staphylococcus aureus towards polymyxins. mBio. 2017;8. https://doi.org/10.1128/mBio.01114-17.
Liu A, Tran L, Becket E, Lee K, Chinn L, Park E, Tran K, Miller JH. Antibiotic sensitivity profiles determined with an Escherichia coli gene knockout collection: generating an antibiotic bar code. Antimicrob Agents Chemother. 2010;54:1393–403. https://doi.org/10.1128/AAC.00906-09.
Yu WB, Pan Q, Ye BC. Glucose-induced cyclic lipopeptides resistance in bacteria via ATP maintenance through enhanced glycolysis. iScience. 2019;21:135–44. https://doi.org/10.1016/j.isci.2019.10.009.
Deris ZZ, Akter J, Sivanesan S, Roberts KD, Thompson PE, Nation RL, Li J, Velkov T. A secondary mode of action of polymyxins against Gram-negative bacteria involves the inhibition of NADH-quinone oxidoreductase activity. J Antibiot. 2014;67:147–51. https://doi.org/10.1038/ja.2013.111.
Humphries R, Bobenchik AM, Hindler JA, Schuetz AN. Overview of changes to the clinical and laboratory standards institute performance standards for antimicrobial susceptibility testing, M100, 31st edition. J Clin Microbiol. 2021;59:e21321. https://doi.org/10.1128/JCM.00213-21.
Lehar J, Krueger AS, Avery W, Heilbut AM, Johansen LM, Price ER, Rickles RJ, Short GR, Staunton JE, Jin X, et al. Synergistic drug combinations tend to improve therapeutically relevant selectivity. Nat Biotechnol. 2009;27:659–66. https://doi.org/10.1038/nbt.1549.
Qu S, Dai C, Shen Z, Tang Q, Wang H, Zhai B, Zhao L, Hao Z. Mechanism of synergy between tetracycline and quercetin against antibiotic resistant Escherichia coli. Front Microbiol. 2019;10:2536. https://doi.org/10.3389/fmicb.2019.02536.
Odds FC. Synergy, antagonism, and what the chequerboard puts between them. J Antimicrob Chemother. 2003;52:1. https://doi.org/10.1093/jac/dkg301.
Toei M, Noji H. Single-molecule analysis of F0F1-ATP synthase inhibited by N,N-dicyclohexylcarbodiimide. J Biol Chem. 2013;288:25717–26. https://doi.org/10.1074/jbc.M113.482455.
Dadi PK, Ahmad M, Ahmad Z. Inhibition of ATPase activity of Escherichia coli ATP synthase by polyphenols. Int J Biol Macromol. 2009;45:72–79. https://doi.org/10.1016/j.ijbiomac.2009.04.004.
Balemans W, Vranckx L, Lounis N, Pop O, Guillemont J, Vergauwen K, Mol S, Gilissen R, Motte M, Lancois D, et al. Novel antibiotics targeting respiratory ATP synthesis in Gram-positive pathogenic bacteria. Antimicrob Agents Chemother. 2012;56:4131–9. https://doi.org/10.1128/AAC.00273-12.
Cheah SE, Li J, Tsuji BT, Forrest A, Bulitta JB, Nation RL. Colistin and polymyxin B dosage regimens against Acinetobacter baumannii: differences in activity and the emergence of resistance. Antimicrob Agents Chemother. 2016;60:3921–33. https://doi.org/10.1128/AAC.02927-15.
Kim JS, Yu JK, Jeon SJ, Park SH, Han S, Park SH, Kang M, Jang JI, Shin EK, Kim J, et al. Distribution of mcr genes among carbapenem-resistant Enterobacterales clinical isolates: high prevalence of mcr-positive Enterobacter cloacae complex in Seoul, Republic of Korea. Int J Antimicrob Agents. 2021;58:106418. https://doi.org/10.1016/j.ijantimicag.2021.106418.
Tran TB, Wang J, Doi Y, Velkov T, Bergen PJ, Li J. Novel polymyxin combination with antineoplastic mitotane improved the bacterial killing against polymyxin-resistant multidrug-resistant Gram-negative pathogens. Front Microbiol. 2018;9:721. https://doi.org/10.3389/fmicb.2018.00721.
Ayerbe-Algaba R, Gil-Marques ML, Miro-Canturri A, Parra-Millan R, Pachon-Ibanez ME, Jimenez-Mejias ME, Pachon J, Smani Y. The anthelmintic oxyclozanide restores the activity of colistin against colistin-resistant Gram-negative bacilli. Int J Antimicrob Agents. 2019;54:507–12. https://doi.org/10.1016/j.ijantimicag.2019.07.006.
Zhang Y, Wang X, Li X, Dong L, Hu X, Nie T, Lu Y, Lu X, Pang J, Li G, et al. Synergistic effect of colistin combined with PFK-158 against colistin-resistant Enterobacteriaceae. Antimicrob Agents Chemother. 2019;63. https://doi.org/10.1128/AAC.00271-19.
Wahdan SA, Azab SS, Elsherbiny DA, El-Demerdash E. Piceatannol protects against cisplatin nephrotoxicity via activation of Nrf2/HO-1 pathway and hindering NF-kappaB inflammatory cascade. Naunyn Schmiedebergs Arch Pharm. 2019;392:1331–45. https://doi.org/10.1007/s00210-019-01673-8.
Breuss JM, Atanasov AG, Uhrin P. Resveratrol and its effects on the vascular system. Int J Mol Sci. 2019;20. https://doi.org/10.3390/ijms20071523.
Meng T, Xiao D, Muhammed A, Deng J, Chen L, He J. Anti-inflammatory action and mechanisms of resveratrol. Molecules. 2021;26. https://doi.org/10.3390/molecules26010229.
Banik K, Ranaware AM, Harsha C, Nitesh T, Girisa S, Deshpande V, Fan L, Nalawade SP, Sethi G, Kunnumakkara AB. Piceatannol: A natural stilbene for the prevention and treatment of cancer. Pharm Res. 2020;153:104635. https://doi.org/10.1016/j.phrs.2020.104635.
Catalgol B, Batirel S, Taga Y, Ozer NK. Resveratrol: French paradox revisited. Front Pharm. 2012;3:141. https://doi.org/10.3389/fphar.2012.00141.
Wang L, Zhang Y, Lin Y, Cao J, Xu C, Chen L, Wang Y, Sun Y, Zheng X, Liu Y, et al. Resveratrol increases sensitivity of clinical colistin-resistant pseudomonas aeruginosa to colistin in vitro and in vivo. Microbiol Spectr. 2023;11:e199222. https://doi.org/10.1128/spectrum.01992-22.
Liu L, Yu J, Shen X, Cao X, Zhan Q, Guo Y, Yu F. Resveratrol enhances the antimicrobial effect of polymyxin B on Klebsiella pneumoniae and Escherichia coli isolates with polymyxin B resistance. BMC Microbiol. 2020;20:306. https://doi.org/10.1186/s12866-020-01995-1.
Fukuhara K, Miyata N. Resveratrol as a new type of DNA-cleaving agent. Bioorg Med Chem Lett. 1998;8:3187–92. https://doi.org/10.1016/s0960-894x(98)00585-x.
Hwang D, Lim YH. Resveratrol antibacterial activity against Escherichia coli is mediated by Z-ring formation inhibition via suppression of FtsZ expression. Sci Rep. 2015;5:10029. https://doi.org/10.1038/srep10029.
Yeaman MR, Yount NY. Mechanisms of antimicrobial peptide action and resistance. Pharm Rev. 2003;55:27–55. https://doi.org/10.1124/pr.55.1.2.
Alteri CJ, Lindner JR, Reiss DJ, Smith SN, Mobley HL. The broadly conserved regulator PhoP links pathogen virulence and membrane potential in Escherichia coli. Mol Microbiol. 2011;82:145–63. https://doi.org/10.1111/j.1365-2958.2011.07804.x.
Funding
This work was supported by grants from the National Natural Science Foundation of China (32200159, 82272352, 82130065).
Author information
Authors and Affiliations
Contributions
JY, ZF, and ZL designed the experiments. ZF, ZL, TF, YF, BD, XC, RZ, and HZ performed the experiments. The other authors analyzed the results. ZF and ZL wrote the manuscript. JY and ZF revised the manuscript. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Ethics approval and consent to participate
We declared that all animal experiments complied with the US and Chinese national guidelines for the use of animals in research. The protocol was approved by the Capital Institute of Pediatrics (permission number DWLL2023011).
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Fan, Z., Li, Z., Fu, T. et al. Inhibition of the ATP synthase increases sensitivity of Escherichia coli carrying mcr-1 to polymyxin B. J Antibiot 77, 685–696 (2024). https://doi.org/10.1038/s41429-024-00753-z
Received:
Revised:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41429-024-00753-z
This article is cited by
-
Natural compounds: a promising therapeutic option for managing colistin-resistant bacteria and improving colistin activity
Molecular Biology Reports (2026)
-
The interactions of natural compounds with Escherichia coli motility, attachment, communication systems, and mature biofilm
Archives of Microbiology (2025)