Abstract
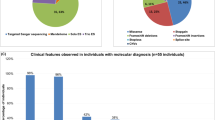
High-throughput sequencing (HTS) of human genome coding regions allows the simultaneous screen of a large number of genes, significantly improving the diagnosis of non-syndromic intellectual disabilities (ID). HTS studies permit the redefinition of the phenotypical spectrum of known disease-causing genes, escaping the clinical inclusion bias of gene-by-gene Sanger sequencing. We studied a cohort of 903 patients with ID not reminiscent of a well-known syndrome, using an ID-targeted HTS of several hundred genes and found de novo heterozygous variants in TCF4 (transcription factor 4) in eight novel patients. Piecing together the patients from this study and those from previous large-scale unbiased HTS studies, we estimated the rate of individuals with ID carrying a disease-causing TCF4 mutation to 0.7%. So far, TCF4 molecular abnormalities were known to cause a syndromic form of ID, Pitt–Hopkins syndrome (PTHS), which combines severe ID, developmental delay, absence of speech, behavioral and ventilation disorders, and a distinctive facial gestalt. Therefore, we reevaluated ten patients carrying a pathogenic or likely pathogenic variant in TCF4 (eight patients included in this study and two from our previous ID-HTS study) for PTHS criteria defined by Whalen and Marangi. A posteriori, five patients had a score highly evocative of PTHS, three were possibly consistent with this diagnosis, and two had a score below the defined PTHS threshold. In conclusion, these results highlight TCF4 as a frequent cause of moderate to profound ID and broaden the clinical spectrum associated to TCF4 mutations to nonspecific ID.
Similar content being viewed by others
Log in or create a free account to read this content
Gain free access to this article, as well as selected content from this journal and more on nature.com
or
References
Redin C, Gerard B, Lauer J, et al. Efficient strategy for the molecular diagnosis of intellectual disability using targeted high-throughput sequencing. J Med Genet. 2014;51:724–36.
Amiel J, Rio M, de Pontual L, et al. Mutations in TCF4, encoding a class I basic helix-loop-helix transcription factor, are responsible for Pitt-Hopkins syndrome, a severe epileptic encephalopathy associated with autonomic dysfunction. Am J Hum Genet. 2007;80:988–93.
Brockschmidt A, Todt U, Ryu S, et al. Severe mental retardation with breathing abnormalities (Pitt-Hopkins syndrome) is caused by haploinsufficiency of the neuronal bHLH transcription factor TCF4. Hum Mol Genet. 2007;16:1488–94.
Zweier C, Peippo MM, Hoyer J, et al. Haploinsufficiency of TCF4 causes syndromal mental retardation with intermittent hyperventilation (Pitt-Hopkins syndrome). Am J Hum Genet. 2007;80:994–1001.
Zhuang Y, Cheng P, Weintraub H. B-lymphocyte development is regulated by the combined dosage of three basic helix-loop-helix genes, E2A, E2-2, and HEB. Mol Cell Biol. 1996;16:2898–905.
Whalen S, Heron D, Gaillon T, et al. Novel comprehensive diagnostic strategy in Pitt-Hopkins syndrome: clinical score and further delineation of the TCF4 mutational spectrum. Hum Mutat. 2012;33:64–72.
Marangi G, Ricciardi S, Orteschi D, et al. Proposal of a clinical score for the molecular test for Pitt-Hopkins syndrome. Am J Med Genet A. 2012;158A:1604–11.
Hamdan FF, Daoud H, Patry L, et al. Parent-child exome sequencing identifies a de novo truncating mutation in TCF4 in non-syndromic intellectual disability. Clin Genet. 2013;83:198–200.
Kalscheuer VM, Feenstra I, Van Ravenswaaij-Arts CM, et al. Disruption of the TCF4 gene in a girl with mental retardation but without the classical Pitt-Hopkins syndrome. Am J Med Genet A. 2008;146A:2053–9.
Kharbanda M, Kannike K, Lampe A, Berg J, Timmusk T, Sepp M. Partial deletion of TCF4 in three generation family with non-syndromic intellectual disability, without features of Pitt-Hopkins syndrome. Eur J Med Genet. 2016;59:310–4.
de Ligt J, Willemsen MH, van Bon BW, et al. Diagnostic exome sequencing in persons with severe intellectual disability. N Engl J Med. 2012;367:1921–9.
Grozeva D, Carss K, Spasic-Boskovic O, et al. Targeted next-generation sequencing analysis of 1000 individuals with intellectual disability. Hum Mutat. 2015;36:1197–204.
Hamdan FF, Srour M, Capo-Chichi JM, et al. De novo mutations in moderate or severe intellectual disability. PLoS Genet. 2014;10:e1004772.
Rauch A, Wieczorek D, Graf E, et al. Range of genetic mutations associated with severe non-syndromic sporadic intellectual disability: an exome sequencing study. Lancet. 2012;380:1674–82.
Tan CA, Topper S, Del Gaudio D et al. Characterization of patients referred for non-specific intellectual disability testing: the importance of autosomal genes for diagnosis. Clin Genet. 2016;89:478–483.
Geoffroy V, Pizot C, Redin C, et al. VaRank: a simple and powerful tool for ranking genetic variants. PeerJ. 2015;3:e796.
Biancalana V, Beldjord C, Taillandier A, et al. Five years of molecular diagnosis of Fragile X syndrome (1997–2001): a collaborative study reporting 95% of the activity in France. Am J Med Genet A. 2004;129A:218–24.
Hagerman PJ. The fragile X prevalence paradox. J Med Genet. 2008;45:498–9.
Hoyer J, Ekici AB, Endele S, et al. Haploinsufficiency of ARID1B, a member of the SWI/SNF-a chromatin-remodeling complex, is a frequent cause of intellectual disability. Am J Hum Genet. 2012;90:565–72.
Deciphering Developmental Disorders S. Prevalence and architecture of de novo mutations in developmental disorders. Nature. 2017;542:433–8.
de Winter CF, Baas M, Bijlsma EK, van Heukelingen J, Routledge S, Hennekam RC. Phenotype and natural history in 101 individuals with Pitt-Hopkins syndrome through an internet questionnaire system. Orphanet J Rare Dis. 2016;11:37.
Zweier C, Sticht H, Bijlsma EK, et al. Further delineation of Pitt-Hopkins syndrome: phenotypic and genotypic description of 16 novel patients. J Med Genet. 2008;45:738–44.
Marangi G, Zollino M. Pitt-Hopkins syndrome and differential diagnosis: a molecular and clinical challenge. J Pediatr Genet. 2015;4:168–76.
Sepp M, Pruunsild P, Timmusk T. Pitt-Hopkins syndrome-associated mutations in TCF4 lead to variable impairment of the transcription factor function ranging from hypomorphic to dominant-negative effects. Hum Mol Genet. 2012;21:2873–88.
Rosenfeld JA, Leppig K, Ballif BC, et al. Genotype-phenotype analysis of TCF4 mutations causing Pitt-Hopkins syndrome shows increased seizure activity with missense mutations. Genet Med. 2009;11:797–805.
Giurgea I, Missirian C, Cacciagli P, et al. TCF4 deletions in Pitt-Hopkins Syndrome. Hum Mutat. 2008;29:E242–51.
Kousoulidou L, Tanteles G, Moutafi M, Sismani C, Patsalis PC, Anastasiadou V. 263.4 kb deletion within the TCF4 gene consistent with Pitt-Hopkins syndrome, inherited from a mosaic parent with normal phenotype. Eur J Med Genet. 2013;56:314–8.
Rossi M, Labalme A, Cordier MP, et al. Mosaic 18q21.2 deletions including the TCF4 gene: a clinical report. Am J Med Genet A. 2012;158A:3174–81.
Steinbusch CV, van Roozendaal KE, Tserpelis D, et al. Somatic mosaicism in a mother of two children with Pitt-Hopkins syndrome. Clin Genet. 2013;83:73–77.
Schluth-Bolard C, Labalme A, Cordier MP, et al. Breakpoint mapping by next generation sequencing reveals causative gene disruption in patients carrying apparently balanced chromosome rearrangements with intellectual deficiency and/or congenital malformations. J Med Genet. 2013;50:144–50.
Maduro V, Pusey BN, Cherukuri PF, et al. Complex translocation disrupting TCF4 and altering TCF4 isoform expression segregates as mild autosomal dominant intellectual disability. Orphanet J Rare Dis. 2016;11:62.
de Pontual L, Mathieu Y, Golzio C, et al. Mutational, functional, and expression studies of the TCF4 gene in Pitt-Hopkins syndrome. Hum Mutat. 2009;30:669–76.
Peippo MM, Simola KO, Valanne LK, et al. Pitt-Hopkins syndrome in two patients and further definition of the phenotype. Clin Dysmorphol. 2006;15:47–54.
Acknowledgements
We thank the families for their participation to the study. We also thank the Fondation Jerome Lejeune and the Agence de la Biomédecine for their financial support. They want also to thank all the people from the Strasbourg Hospital molecular diagnostic lab, from the IGBMC and CNG/CEA sequencing platform from UMR_S 1112 (Bernard Jost, Stéphanie Le Gras, Mathieu Jung, Céline Keime, Jean Muller, Véronique Geoffroy) and from the Mondor Hospital molecular diagnostic lab (Thierry Gaillon) for their technical and bioinformatical supports.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Mary, L., Piton, A., Schaefer, E. et al. Disease-causing variants in TCF4 are a frequent cause of intellectual disability: lessons from large-scale sequencing approaches in diagnosis. Eur J Hum Genet 26, 996–1006 (2018). https://doi.org/10.1038/s41431-018-0096-4
Received:
Revised:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41431-018-0096-4
This article is cited by
-
An attempt to explain what intrinsically disordered TCF4 does in its spare time when PTHS-related mutations prevent it from doing its job
Cell Communication and Signaling (2025)
-
Skeletal muscle vulnerability in a child with Pitt-Hopkins syndrome
Skeletal Muscle (2024)
-
Repurposing the Dihydropyridine Calcium Channel Inhibitor Nicardipine as a Nav1.8 Inhibitor In Vivo for Pitt Hopkins Syndrome
Pharmaceutical Research (2020)