Abstract
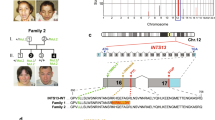
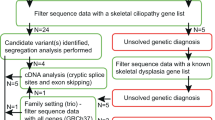
Whole exome sequencing (WES) is utilized in diagnostic odyssey cases to identify the underlying genetic cause associated with complex phenotypes. Recent publications suggest that WES reveals the genetic cause in ~25% of these cases and is most successful when applied to children with neurological disease. The residual 75% of cases remain genetically elusive until more information becomes available in the literature or functional studies are pursued. WES performed on three families with presumed ciliopathy diagnoses, including orofaciodigital (OFD) syndrome, fetal encephalocele, or Joubert-related disorder, identified compound heterozygous variants in C2CD3. Biallelic variants in C2CD3 have previously been associated with ciliopathies, including OFD syndrome type 14 (OFD14; MIM: 615948). As three of the six identified variants were predicted to affect splicing, exon-skipping analysis using either RNA sequencing or PCR-based methods were completed to determine the pathogenicity of these variants, and showed that each of the splicing variants led to a frameshifted protein product. Using these studies in combination with the 2015 ACMG guidelines, each of the six identified variants were classified as either pathogenic or likely pathogenic, and are therefore likely responsible for our patients’ phenotypes. Each of the families had a distinct clinical phenotype and severity of disease, extending from lethal to viable. These findings highlight that there is a broad phenotypic spectrum associated with C2CD3-mediated disease and not all patients present with the typical features of OFD14.
Similar content being viewed by others
Log in or create a free account to read this content
Gain free access to this article, as well as selected content from this journal and more on nature.com
or
References
Braun DA, Hildebrandt F: Ciliopathies. Cold Spring Harb Perspect Biol. 2017;9:a028191.
Bruel AL, Franco B, Duffourd Y, Thevenon J, Jego L, Lopez E, et al. Fifteen years of research on oral-facial-digital syndromes: from 1 to 16 causal genes. J Med Genet. 2017;54:371–80.
Waters AM, Beales PL. Ciliopathies: an expanding disease spectrum. Pediatr Nephrol. 2011;26:1039–56.
Hoover AN, Wynkoop A, Zeng H, Jia J, Niswander LA, Liu A. C2cd3 is required for cilia formation and Hedgehog signaling in mouse. Development. 2008;135:4049–58.
Thauvin-Robinet C, Lee JS, Lopez E, Herranz-Perez V, Shida T, Franco B, et al. The oral-facial-digital syndrome gene C2CD3 encodes a positive regulator of centriole elongation. Nat Genet. 2014;46:905–11.
Bachmann-Gagescu R, Dempsey JC, Phelps IG, O'Roak BJ, Knutzen DM, Rue TC, et al. Joubert syndrome: a model for untangling recessive disorders with extreme genetic heterogeneity. J Med Genet. 2015;52:514–22.
Srour M, Hamdan FF, McKnight D, Davis E, Mandel H, Schwartzentruber J, et al. Joubert syndrome in French Canadians and identification of mutations in CEP104. Am J Hum Genet. 2015;97:744–53.
Cortés CR, McInerney-Leo AM, Vogel I, Rondon Galeano MC, Leo PJ, Harris JE, et al. Mutations in human C2CD3 cause skeletal dysplasia and provide new insights into phenotypic and cellular consequences of altered C2CD3 function. Sci Rep. 2016;6:24083.
Asmann YW, Middha S, Hossain A, Baheti S, Li Y, Chai HS, et al. TREAT: a bioinformatics tool for variant annotations and visualizations in targeted and exome sequencing data. Bioinformatics. 2012;28:277–8.
Kalari KR, Nair AA, Bhavsar JD, O'Brien DR, Davila JI, Bockol MA, et al. MAP-RSeq: Mayo Analysis Pipeline for RNA sequencing. BMC Bioinforma. 2014;27:224.
Kim D, Pertea G, Trapnell C, Pimentel H, Kelley R, Salzberg SL. Tophat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 2013;14: R36.
Langmead B. Aligning short sequencing reads with bowtie. Cur Protoc Bioinforma. 2010;32:11.17.11–11.17.14.
Anders S, Pyl PT, Huber W. Htseq - a python framework to work with high-throughput sequencing data. Bioinformatics. 2015;31:166–9.
Quinlan AR. Bedtools: the swiss-army tool for genome feature analysis. Curr Protoc Bioinforma. 2014;47:1–34.
R C Team. R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. URL https://www.R-project.org/; 2015.
Bates D, Maechler M, Bolker B, Walker S. Fitting linear mixed-effects models using. lme4 J Stat Softw. 2015;67:1–48.
Adzhubei IA, Schmidt S, Peshkin L, Ramensky VE, Gerasimova A, Bork P, et al. A method and server for predicting damaging missense mutations. Nat Methods. 2010;7:248–9.
Desmet FO, Hamroun D, Lalande M, Collod-Beroud G, Claustres M, Beroud C. Human Splicing Finder: an online bioinformatics tool to predict splicing signals. Nucleic Acids Res. 2009;37:e67.
Kumar P, Henikoff S, Ng PC. Predicting the effects of coding non-synonymous variants on protein function using the SIFT algorithm. Nat Protoc. 2009;4:1073–81.
Pertea M, Lin X, Salzberg SL. GeneSplicer: a new computational method for splice site prediction. Nucleic Acids Res. 2001;29:1185–90.
Reese MG, Eeckman FH, Kulp D, Haussler D. Improved splice site detection in genie. J Comput Biol. 1997;4:311–23.
Schwarz JM, Rödelsperger C, Schuelke M, Seelow D. MutationTaster evaluates disease-causing potential of sequence alterations. Nat Methods. 2010;7:575–6.
Tavtigian SV, Deffenbaugh AM, Yin L, Judkins T, Scholl T, Samollow PB, et al. Comprehensive statistical study of 452 BRCA1 missense substitutions with classification of eight recurrent substitutions as neutral. J Med Genet. 2006;43:295–305.
Yeo G, Burge CB. Maximum entropy modeling of short sequence motifs with applications to RNA splicing signals. J Comput Biol. 2004;11:377–94.
Zhang MQ. Statistical features of human exons and their flanking regions. Hum Mol Genet. 1998;7:919–32.
Zohn IE, Anderson KV, Niswander LA. Using genomewide mutagenesis screens to identify the genes required for neural tube closure in the mouse. Birth Defects Res A Clin Mol Teratol. 2005;73:583–90.
Chang CF, Schock EN, O’Hare EA, Dodgson J, Cheng HH, Muir WM, et al. The cellular and molecular etiology of the craniofacial defects in the avian ciliopathic mutant talpid2. Development. 2014;141:3003–12.
Schock EN, Chang CF, Struve JN, Chang YT, Chang J, Delany ME, et al. Using the avian mutant talpid2 as a disease model for understanding the oral-facial phenotypes of oral-facial-digital syndrome. Dis Model Mech. 2015;8:855–66.
Acknowledgements
Mayo Clinic Center for Individualized Medicine Functional Validation Program, DK059597 (PCH).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Boczek, N.J., Hopp, K., Benoit, L. et al. Characterization of three ciliopathy pedigrees expands the phenotype associated with biallelic C2CD3 variants. Eur J Hum Genet 26, 1797–1809 (2018). https://doi.org/10.1038/s41431-018-0222-3
Received:
Revised:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41431-018-0222-3
This article is cited by
-
Syndromic ciliopathy: a taiwanese single-center study
BMC Medical Genomics (2024)
-
NGS and phenotypic ontology-based approaches increase the diagnostic yield in syndromic retinal diseases
Human Genetics (2021)
-
CEP120 interacts with C2CD3 and Talpid3 and is required for centriole appendage assembly and ciliogenesis
Scientific Reports (2019)