Abstract
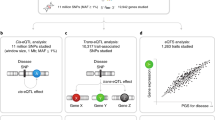
Insights into individual differences in gene expression and its heritability (h2) can help in understanding pathways from DNA to phenotype. We estimated the heritability of gene expression of 52,844 genes measured in whole blood in the largest twin RNA-Seq sample to date (1497 individuals including 459 monozygotic twin pairs and 150 dizygotic twin pairs) from classical twin modeling and identity-by-state-based approaches. We estimated for each gene h2total, composed of cis-heritability (h2cis, the variance explained by single nucleotide polymorphisms in the cis-window of the gene), and trans-heritability (h2res, the residual variance explained by all other genome-wide variants). Mean h2total was 0.26, which was significantly higher than heritability estimates earlier found in a microarray-based study using largely overlapping (>60%) RNA samples (mean h2 = 0.14, p = 6.15 × 10−258). Mean h2cis was 0.06 and strongly correlated with beta of the top cis expression quantitative loci (eQTL, ρ = 0.76, p < 10−308) and with estimates from earlier RNA-Seq-based studies. Mean h2res was 0.20 and correlated with the beta of the corresponding trans-eQTL (ρ = 0.04, p < 1.89 × 10−3) and was significantly higher for genes involved in cytokine-cytokine interactions (p = 4.22 × 10−15), many other immune system pathways, and genes identified in genome-wide association studies for various traits including behavioral disorders and cancer. This study provides a thorough characterization of cis- and trans-h2 estimates of gene expression, which is of value for interpretation of GWAS and gene expression studies.
Similar content being viewed by others
Log in or create a free account to read this content
Gain free access to this article, as well as selected content from this journal and more on nature.com
or
References
Leek JT, Storey JD. Capturing heterogeneity in gene expression studies by surrogate variable analysis. PLoS Genet. 2007;3:1724–35.
Bryois J, Buil A, Ferreira PG, Panousis NI, Brown AA, Viñuela A, et al. Time-dependent genetic effects on gene expression implicate aging processes. Genome Res. 2017;27:545–52.
Jansen R, Batista S, Brooks AI, Tischfield JA, Willemsen G, van Grootheest G, et al. Sex differences in the human peripheral blood transcriptome. BMC Genomics. 2014;15:33.
Meder B, Backes C, Haas J, Leidinger P, Stähler C, Großmann T, et al. Influence of the confounding factors age and sex on microRNA profiles from peripheral blood. Clin Chem. 2014;60:1200–8.
Tower J. Sex-specific gene expression and life span regulation. Trends Endocrinol Metab. 2017;28:735–47.
Lloyd-Jones LR, Holloway A, McRae A, Yang J, Small K, Zhao J, et al. The genetic architecture of gene expression in peripheral blood. Am J Hum Genet. 2017;100:228–37.
Wright FA, Sullivan PF, Brooks AI, Zou F, Sun W, Xia K, et al. Heritability and genomics of gene expression in peripheral blood. Nat Genet. 2014;46:430–7.
Vinuela A, Brown AA, Buil A, Tsai PC, Davies MN, Bell JT, et al. Age-dependent changes in mean and variance of gene expression across tissues in a twin cohort. Hum Mol Genet. 2018;27:732–41.
Vink JM, Jansen R, Brooks A, Willemsen G, van Grootheest G, de Geus E, et al. Differential gene expression patterns between smokers and non‐smokers: cause or consequence? Addiction Biol. 2017;22:550–60.
Willemsen G, de Geus EJ, Bartels M, van Beijsterveldt CE, Brooks AI. Estourgie-van Burg GF et al: The Netherlands Twin Register biobank: a resource for genetic epidemiological studies. Twin Res Hum Genet. 2010;13:231–45.
Willemsen G, Vink JM, Abdellaoui A, den Braber A, van Beek JH, Draisma HH, et al. The adult netherlands twin register: twenty-five years of survey and biological data collection. Twin Res Hum Genet. 2013;16:271–81.
Boomsma D, Busjahn A, Peltonen L. Classical twin studies and beyond. Nat Rev Genet. 2002;3:872–82.
Yang J, Lee SH, Goddard ME, Visscher PM. GCTA: a tool for genome-wide complex trait analysis. Am J Hum Genet. 2011;88:76–82.
Willemsen G, Vink JM, Abdellaoui A, den Braber A, van Beek JH, Draisma HH, et al. The Adult Netherlands Twin Register: twenty-five years of survey and biological data collection. Twin Res Hum Genet. 2013;16:271–81.
Genome of the Netherlands C. Whole-genome sequence variation, population structure and demographic history of the Dutch population. Nat Genet. 2014;46:818–25.
Zhernakova DV, Deelen P, Vermaat M, van Iterson M, van Galen M, Arindrarto W, et al. Identification of context-dependent expression quantitative trait loci in whole blood. Nat Genet. 2017;49:139–45.
Boomsma DI, Wijmenga C, Slagboom EP, Swertz MA, Karssen LC, Abdellaoui A, et al. The Genome of the Netherlands: design, and project goals. Eur J Hum Genet. 2014;22:221–7.
R Development Core Team. R: a language and environment for statistical computing. Vienna: R Foundation for Statistical Computing; 2014.
Neale MC, Hunter MD, Pritikin JN, Zahery M, Brick TR, Kirkpatrick RM, et al. OpenMx 2.0: extended structural equation and statistical modeling. Psychometrika. 2016;81:535–49.
Zaitlen N, Kraft P, Patterson N, Bogdan P, Gaurav, Samuela P, et al. Using extended genealogy to estimate components of heritability for 23 quantitative and dichotomous traits. PLoS Genet. 2013;9:e1003520.
Visscher PM, Hemani G, Vinkhuyzen AA, Chen GB, Lee SH, Wray NR, et al. Statistical power to detect genetic (co) variance of complex traits using SNP data in unrelated samples. PLoS Genet. 2014;10:e1004269.
Lek M, Karczewski KJ, Minikel EV, Samocha KE, Banks E, Fennell T, et al. Analysis of protein-coding genetic variation in 60,706 humans. Nature. 2016;536:285–91.
Smedley D, Haider S, Durinck S, Pandini L, Provero P, Allen J, et al. The BioMart community portal: an innovative alternative to large, centralized data repositories. Nucleic acids Res. 2015;43:W589–W598.
Kinsella RJ, Kähäri A, Haider S, Zamora J, Proctor G, Spudich G et al: Ensembl BioMarts: a hub for data retrieval across taxonomic space. Database 2011;2011:bar030.
MacArthur J, Bowler E, Cerezo M, Gil L, Hall P, Hastings E, et al. The new NHGRI-EBI Catalog of published genome-wide association studies (GWAS Catalog). Nucleic Acids Res. 2017;45:D896–901.
Battle A, Mostafavi S, Zhu X, Patash JB, Weissman MM, McCormick C, et al. Characterizing the genetic basis of transcriptome diversity through RNA-sequencing of 922 individuals. Genome Res. 2014;24:14–24.
Consortium GT, Laboratory DA, Coordinating Center -Analysis Working G, Enhancing GTEx (eGTEx) groups, NIH Common Fund, NIH/NCI, et al. Genetic effects on gene expression across human tissues. Nature. 2017;550:204–13.
Franić S, Dolan CV, Borsboom D, Boomsma DI. Structural equation modeling in genetics. In: Hoyle RH (ed) Handbook of structural equation modeling. New York: Guilford Press; 2012. pp 617–35.
Wheeler HE, Shah KP, Brenner J, Garcia T, Aquino-Michaels K, GTEX Consortium et al. Survey of the heritability and sparse architecture of gene expression traits across human tissues. PLoS Genet. 2016;12:e1006423.
Kanehisa M, Furumichi M, Tanabe M, Sato Y, Morishima K. KEGG: new perspectives on genomes, pathways, diseases and drugs. Nucleic Acids Res. 2016;45:D353–61.
Acknowledgements
We very warmly thank all participants in the study. This study makes use of data in the Netherlands Twin Register with prof. D.I. Boomsma as principle investigator.
Funding
This work was performed within the framework of the BBMRI - NL Consortium, a research infrastructure financed by the Dutch government (NWO, nos. 184.021.007 and 184.033.111). Genotyping was made possible by grants from NWO/SPI 56-464-14192, Genetic Association Information Network (GAIN) of the Foundation for the National Institutes of Health, Rutgers University Cell and DNA Repository (NIMH U24 MH068457-06), the Avera Institute, Sioux Falls (USA) and the National Institutes of Health (NIH R01 HD042157-01A1, MH081802, Grand Opportunity grants 1RC2 MH089951 and 1RC2 MH089995) and European Research Council (ERC-230374). DIB acknowledges her KNAW Academy Professor Award (PAH/6635).
BIOS Consortium
Bastiaan T. Heijmans12, Peter A. C. ’t Hoen13, Joyce van Meurs14, Aaron Isaacs15, Rick Jansen16, Lude Franke17, Dorret I. Boomsma18, René Pool18, Jenny van Dongen18, Jouke J. Hottenga18, Marleen M. J. van Greevenbroek19, Coen D. A. Stehouwer19, Carla J. H. van der Kallen19, Casper G. Schalkwijk19, Cisca Wijmenga17, Lude Franke17, Sasha Zhernakova17, Ettje F. Tigchelaar17, P. Eline Slagboom12, Marian Beekman12, Joris Deelen12, Diana van Heemst20, Jan H. Veldink21, Leonard H. van den Berg21, Cornelia M. van Duijn15, Bert A. Hofman22, Aaron Isaacs15, André G. Uitterlinden14, Joyce van Meurs14, P. Mila Jhamai14, Michael Verbiest14, H. Eka D. Suchiman12, Marijn Verkerk14, Ruud van der Breggen12, Jeroen van Rooij14, Nico Lakenberg12, Hailiang Mei23, Maarten van Iterson12, Michiel van Galen13, Jan Bot24, Dasha V. Zhernakova17, Rick Jansen16, Peter van’t Hof23, Patrick Deelen17, Irene Nooren24, Peter A. C. ’t Hoen13, Bastiaan T. Heijmans12, Matthijs Moed12, Lude Franke17, Martijn Vermaat14, Dasha V. Zhernakova17, René Luijk12, Marc Jan Bonder17, Maarten van Iterson12, Patrick Deelen17, Freerk van Dijk25, Michiel van Galen13, Wibowo Arindrarto23, Szymon M. Kielbasa26, Morris A. Swertz25, Erik. W van Zwet26, Rick Jansen16, Peter-Bram ’t Hoen13, Bastiaan T. Heijmans12
Author information
Authors and Affiliations
Consortia
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
The NTR study was approved by the Central Ethics Committee on Research Involving Human Subjects of the VU University Medical Center, Amsterdam (institutional review board [IRB] number IRB-2991 under Federal wide Assurance 3703; IRB/institute code NTR 03-180). All participants provided written informed consent.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Members of the BIOS Consortium are listed at the end of the paper.
Supplementary information
Rights and permissions
About this article
Cite this article
Ouwens, K.G., Jansen, R., Nivard, M.G. et al. A characterization of cis- and trans-heritability of RNA-Seq-based gene expression. Eur J Hum Genet 28, 253–263 (2020). https://doi.org/10.1038/s41431-019-0511-5
Received:
Revised:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41431-019-0511-5
This article is cited by
-
Deciphering the coordinated roles of the host genome, duodenal mucosal genes, and microbiota in regulating complex traits in chickens
Microbiome (2025)
-
Transcripts with high distal heritability mediate genetic effects on complex metabolic traits
Nature Communications (2025)
-
Diallel panel reveals a significant impact of low-frequency genetic variants on gene expression variation in yeast
Molecular Systems Biology (2024)
-
Genetic analysis of the blood transcriptome of young healthy pigs to improve disease resilience
Genetics Selection Evolution (2023)
-
Metabolomic epidemiology offers insights into disease aetiology
Nature Metabolism (2023)