Abstract
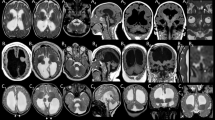
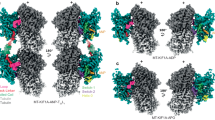
Variants in multiple tubulin genes have been implicated in neurodevelopmental disorders, including malformations of cortical development (MCD) and congenital fibrosis of the extraocular muscles (CFEOM). Distinct missense variants in the beta-tubulin encoding genes TUBB3 and TUBB2B cause MCD, CFEOM, or both, suggesting substitution-specific mechanisms. Variants in the alpha tubulin-encoding gene TUBA1A have been associated with MCD, but not with CFEOM. Using exome sequencing (ES) and genome sequencing (GS), we identified 3 unrelated probands with CFEOM who harbored novel heterozygous TUBA1A missense variants c.1216C>G, p.(His406Asp); c.467G>A, p.(Arg156His); and c.1193T>G, p.(Met398Arg). MRI revealed small oculomotor-innervated muscles and asymmetrical caudate heads and lateral ventricles with or without corpus callosal thinning. Two of the three probands had MCD. Mutated amino acid residues localize either to the longitudinal interface at which α and β tubulins heterodimerize (Met398, His406) or to the lateral interface at which tubulin protofilaments interact (Arg156), and His406 interacts with the motor domain of kinesin-1. This series of individuals supports TUBA1A variants as a cause of CFEOM and expands our knowledge of tubulinopathies.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Lasser M, Tiber J, Lowery LA. The role of the microtubule cytoskeleton in neurodevelopmental disorders. Front Cell Neurosci. 2018;12:165.
Lopata MA, Cleveland DW. In vivo microtubules are copolymers of available beta-tubulin isotypes: localization of each of six vertebrate beta-tubulin isotypes using polyclonal antibodies elicited by synthetic peptide antigens. J Cell Biol. 1987;105:1707–20.
Tilney LG, Bryan J, Bush DJ, Fujiwara K, Mooseker MS, Murphy DB, et al. Microtubules: evidence for 13 protofilaments. J Cell Biol. 1973;59:267–75.
Mitchison T, Kirschner M. Dynamic instability of microtubule growth. Nature. 1984;312:237–42.
Fulton C, Simpson P. Selective synthesis and utilization of flagellar tubulin. The multi-tubulin hypothesis. Cell Motil. 1976;3:987–1005.
Gonçalves FG, Freddi T de AL, Taranath A, Lakshmanan R, Goetti R, Feltrin FS, et al. Tubulinopathies. Topics Magn Reson Imaging. 2018;27:395–408.
Köhler S, Carmody L, Vasilevsky N, Jacobsen J, Danis D, Gourdine J, et al. Expansion of the Human Phenotype Ontology (HPO) knowledge base and resources. Nucleic Acids Res. 2019;47:D1018–27.
Engle EC, Goumnerov BC, McKeown CA, Schatz M, Johns DR, Porter JD, et al. Oculomotor nerve and muscle abnormalities in congenital fibrosis of the extraocular muscles. Ann Neurol. 1997;41:314–25.
Demer JL, Clark RA, Engle EC. Magnetic Resonance Imaging Evidence For Widespread Orbital Dysinnervation in Congenital Fibrosis of Extraocular Muscles Due to Mutations in KIF21A. Invest Ophth Vis Sci. 2005;46:530–9.
Lim K, Engle E, Demer J. Abnormalities of the oculomotor nerve in congenital fibrosis of the extraocular muscles and congenital oculomotor palsy. Invest Ophth Vis Sci. 2007;48:1601–6.
Online Mendelian Inheritance in Man, OMIM. Mc-Kusick-Nathans Institute of Genetic Medicine, Johns Hopkins University (Baltimore, MD), accessed December 10, 2020. https://omim.org/
Yamada K, Andrews C, Chan W-M, McKeown CA, Magli A, Berardinis Tde, et al. Heterozygous mutations of the kinesin KIF21A in congenital fibrosis of the extraocular muscles type 1 (CFEOM1). Nat Genet. 2003;35:318–21.
Tischfield M, Baris H, Wu C, Rudolph G, Maldergem L, He W, et al. Human TUBB3 mutations perturb microtubule dynamics, kinesin interactions, and axon guidance. Cell. 2010;140:74–87.
Chew S, Balasubramanian R, Chan W, Kang P, Andrews C, Webb B, et al. A novel syndrome caused by the E410K amino acid substitution in the neuronal β-tubulin isotype 3. Brain. 2013;136:522–35.
Cederquist GY, Luchniak A, Tischfield MA, Peeva M, Song Y, Menezes MP, et al. An inherited TUBB2B mutation alters a kinesin-binding site and causes polymicrogyria, CFEOM and axon dysinnervation. Hum Mol Genet. 2012;21:5484–99.
Whitman MC, Andrews C, Chan W-M, Tischfield MA, Stasheff SF, Brancati F, et al. Two unique TUBB3 mutations cause both CFEOM3 and malformations of cortical development. Am J Med Genet A. 2016;170A:297–305.
Poirier K, Keays D, Francis F, Saillour Y, Bahi N, Manouvrier S, et al. Large spectrum of lissencephaly and pachygyria phenotypes resulting from de novo missense mutations in tubulin alpha1A (TUBA1A). Hum Mutat. 2007;28:1055–64.
Jaglin X, Poirier K, Saillour Y, Buhler E, Tian G, Bahi-Buisson N, et al. Mutations in the beta-tubulin gene TUBB2B result in asymmetrical polymicrogyria. Nat Genet. 2009;41:746–52.
Poirier K, Saillour Y, Bahi-Buisson N, Jaglin X, Fallet-Bianco C, Nabbout R, et al. Mutations in the neuronal ß-tubulin subunit TUBB3 result in malformation of cortical development and neuronal migration defects. Hum Mol Genet. 2010;19:4462–73.
Latremoliere A, Cheng L, DeLisle M, Wu C, Chew S, Hutchinson EB, et al. Neuronal-specific TUBB3 is not required for normal neuronal function but is essential for timely axon regeneration. Cell Rep. 2018;24:1865–79.e9.
Bittermann E, Abdelhamed Z, Liegel R, Menke C, Timms A, Beier D, et al. Differential requirements of tubulin genes in mammalian forebrain development. Plos Genet. 2019;15:e1008243–e1008243.
Keays D, Tian G, Poirier K, Huang G-J, Siebold C, Cleak J, et al. Mutations in alpha-tubulin cause abnormal neuronal migration in mice and lissencephaly in humans. Cell. 2007;128:45–57.
Furuse T, Yamada I, Kushida T, Masuya H, Miura I, Kaneda H, et al. Behavioral and neuromorphological characterization of a novel Tuba1 mutant mouse. Behavioural Brain Res. 2012;227:167–74.
Hanson MG, Aiken J, Sietsema DV, Sept D, Bates EA, Niswander L, et al. Novel α-tubulin mutation disrupts neural development and tubulin proteostasis. Dev Biol. 2016;409:406–19.
Hebebrand M, Hüffmeier U, Trollmann R, Hehr U, Uebe S, Ekici A, et al. The mutational and phenotypic spectrum of TUBA1A-associated tubulinopathy. Orphanet J Rare Dis. 2019;14:38–38.
Nellhaus G. Head circumference from birth to eighteen years. Pediatrics. 1968;41:106–14.
Li H, Durbin R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics. 2009;25:1754–60.
McKenna A, Hanna M, Banks E, Sivachenko A, Cibulskis K, Kernytsky A, et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 2010;20:1297–303.
McLaren W, Gil L, Hunt SE, Riat HS, Ritchie GRS, Thormann A, et al. The ensembl variant effect predictor. Genome Biol. 2016;17:122.
Beaulieu CL, Majewski J, Schwartzentruber J, Samuels ME, Fernandez BA, Bernier FP, et al. FORGE Canada Consortium: outcomes of a 2-year national rare-disease gene-discovery project. Am J Hum Genetics. 2014;94:809–17.
Paila U, Chapman BA, Kirchner R, Quinlan AR. GEMINI: integrative exploration of genetic variation and genome annotations. Plos Comput Biol. 2013;9:e1003153.
Srour M, Schwartzentruber J, Hamdan F, Ospina L, Patry L, Labuda D, et al. Mutations in C5ORF42 cause Joubert syndrome in the French Canadian population. Am J Hum Genetics. 2012;90:693–700.
Lek M, Karczewski K, Minikel E, Samocha K, Banks E, Fennell T, et al. Analysis of protein-coding genetic variation in 60,706 humans. Nature. 2016;536:285–91.
Karczewski K, Francioli L, Tiao G, Cummings B, Alföldi J, Wang Q, et al. The mutational constraint spectrum quantified from variation in 141,456 humans. Nature. 2020;581:434–43.
Auton A, Abecasis GR, Altshuler DM, Durbin RM, Abecasis GR, Bentley DR, et al. A global reference for human genetic variation. Nature. 2015;526:68–74.
Landrum MJ, Lee JM, Benson M, Brown GR, Chao C, Chitipiralla S, et al. ClinVar: improving access to variant interpretations and supporting evidence. Nucleic Acids Res. 2018;46:D1062–7.
Zhang R, Alushin GM, Brown A, Nogales E. Mechanistic origin of microtubule dynamic instability and its modulation by EB proteins. Cell. 2015;162:849–59.
Gigant B, Wang W, Dreier B, Jiang Q, Pecqueur L, Plückthun A, et al. Structure of a kinesin–tubulin complex and implications for kinesin motility. Nat Struct Mol Biol. 2013;20:1001–7.
Sievers F, Wilm A, Dineen D, Gibson TJ, Karplus K, Li W, et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol Syst Biol. 2011;7:539–539.
Waterhouse AM, Procter JB, Martin D, Clamp M, Barton GJ. Jalview Version 2—a multiple sequence alignment editor and analysis workbench. Bioinformatics. 2009;25:1189–91.
Shoshany T, Robson C, Hunter D. Anomalous superior oblique muscles and tendons in congenital fibrosis of the extraocular muscles. J Am Assoc Pediatr Ophthalmol Strabismus. 2019;23:325.e1–325.e6.
Pollard KS, Hubisz MJ, Rosenbloom KR, Siepel A. Detection of nonneutral substitution rates on mammalian phylogenies. Genome Res. 2010;20:110–21.
Rentzsch P, Witten D, Cooper G, Shendure J, Kircher M. CADD: predicting the deleteriousness of variants throughout the human genome. Nucleic Acids Res. 2019;47:D886–94.
Richards S, Aziz N, Bale S, Bick D, Das S, Gastier-Foster J, et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med. 2015;17:405–23.
Samocha K, Robinson E, Sanders S, Stevens C, Sabo A, McGrath L, et al. A framework for the interpretation of de novo mutation in human disease. Nat Genet. 2014;46:944–50.
Nogales E, Whittaker M, Milligan RA, Downing KH. High-resolution model of the microtubule. Cell. 1999;96:79–88.
Löwe J, Li H, Downing KH, Nogales E. Refined structure of αβ-tubulin at 3.5 Å resolution. J Mol Biol. 2001;313:1045–57.
Romaniello R, Zucca C, Arrigoni F, Bonanni P, Panzeri E, Bassi MT, et al. Epilepsy in tubulinopathy: personal series and literature review. Cells. 2019;8:669.
Aiken J, Buscaglia G, Bates E, Moore J. The α-tubulin gene TUBA1A in brain development: a key ingredient in the neuronal isotype blend. J Dev Biol. 2017;5:8.
Romaniello R, Arrigoni F, Panzeri E, Poretti A, Micalizzi A, Citterio A, et al. Tubulin-related cerebellar dysplasia: definition of a distinct pattern of cerebellar malformation. Eur Radio. 2017;27:5080–92.
Acknowledgements
We thank the participants and their family members, Dr. Monkol Lek and Mr. Ben Weisburd for their efforts in reprocessing and upload of sequence data for analysis at the Broad Institute, and Dr. Dan Doherty of the University of Washington for connecting the Bostonian and Canadian researchers.
Funding
This project was supported by NEI R01EY027421 and NHLBI X01HL132377 (ECE), the Broad Institute of MIT and Harvard Center for Mendelian Genomics (NHGRI/NEI/NHLBI UM1HG008900), the Ocular Genomics Institute Genomics Core (Massachusetts Eye and Ear Infirmary/Harvard Medical School, NEI 2P30EY014104), NHGRI R01HG009141, the Care4Rare Canada Consortium funded by Genome Canada and the Ontario Genomics Institute (OGI-147), the Canadian Institutes of Health Research, Ontario Research Fund, Genome Alberta, Genome British Columbia, Genome Quebec, and Children’s Hospital of Eastern Ontario Foundation. JJ was supported by T32GM007748-42, 5T32NS007473-19, and 5T32EY007145-16. MW was supported by NEI 5K08EY027850 and BCH Ophthalmology Foundation Faculty Discovery Award. MW, SM, and DH receive research support from Children’s Hospital Ophthalmology Foundation, Inc., Boston, MA. KMB is a Tier 1 Canada Research Chair in Rare Disease Precision Health. ECE is a Howard Hughes Medical Institute Investigator.
Author information
Authors and Affiliations
Consortia
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Members of the Care4Rare Canada Consortium are listed in the Supplementary Information.
Supplementary information
Rights and permissions
About this article
Cite this article
Jurgens, J.A., Barry, B.J., Lemire, G. et al. Novel variants in TUBA1A cause congenital fibrosis of the extraocular muscles with or without malformations of cortical brain development. Eur J Hum Genet 29, 816–826 (2021). https://doi.org/10.1038/s41431-020-00804-7
Received:
Revised:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41431-020-00804-7
This article is cited by
-
Clinical and MRI differences in congenital fibrosis of extraocular muscles patients with KIF21A and TUBB3 variants
Japanese Journal of Ophthalmology (2025)
-
A cell type-aware framework for nominating non-coding variants in Mendelian regulatory disorders
Nature Communications (2024)
-
Comprehensive whole-genome sequence analyses provide insights into the genomic architecture of cerebral palsy
Nature Genetics (2024)
-
Phenotype, genotype, and management of congenital fibrosis of extraocular muscles type 1 in 16 Chinese families
Graefe's Archive for Clinical and Experimental Ophthalmology (2023)
-
Clinical and genetic characteristics of Chinese patients with congenital cranial dysinnervation disorders
Orphanet Journal of Rare Diseases (2022)