Abstract
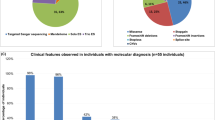
TBR1, a T-box transcription factor expressed in the cerebral cortex, regulates the expression of several candidate genes for autism spectrum disorders (ASD). Although TBR1 has been reported as a high-confidence risk gene for ASD and intellectual disability (ID) in functional and clinical reports since 2011, TBR1 has only recently been recorded as a human disease gene in the OMIM database. Currently, the neurodevelopmental disorders and structural brain anomalies associated with TBR1 variants are not well characterized. Through international data sharing, we collected data from 25 unreported individuals and compared them with data from the literature. We evaluated structural brain anomalies in seven individuals by analysis of MRI images, and compared these with anomalies observed in TBR1 mutant mice. The phenotype included ID in all individuals, associated to autistic traits in 76% of them. No recognizable facial phenotype could be identified. MRI analysis revealed a reduction of the anterior commissure and suggested new features including dysplastic hippocampus and subtle neocortical dysgenesis. This report supports the role of TBR1 in ID associated with autistic traits and suggests new structural brain malformations in humans. We hope this work will help geneticists to interpret TBR1 variants and diagnose ASD probands.
Similar content being viewed by others
Log in or create a free account to read this content
Gain free access to this article, as well as selected content from this journal and more on nature.com
or
References
Bulfone A, Smiga SM, Shimamura K, Peterson A, Puelles L, Rubenstein JLR. T-brain-1: a homolog of Brachyury whose expression defines molecularly distinct domains within the cerebral cortex. Neuron. 1995;15:63–78.
Hevner RF, Shi L, Justice N, Hsueh YP, Sheng M, Smiga S, et al. Tbr1 regulates differentiation of the preplate and layer 6. Neuron. 2001;29:353–66.
Méndez-Gómez HR1, Vergaño-Vera E, Abad JL, Bulfone A, Moratalla R, de Pablo F, et al. The T-box brain 1 (Tbr1) transcription factor inhibits astrocyte formation in the olfactory bulb and regulates neural stem cell fate. Mol Cell Neurosci. 2011;46:108–21.
Huang TN, Chuang HC, Chou WH, Chen CY, Wang HF, Chou SJ, et al. Tbr1 haploinsufficiency impairs amygdalar axonal projections and results in cognitive abnormality. Nat Neurosci. 2014;17:240–7.
Notwell JH, Heavner WE, Darbandi SF, Katzman S, McKenna WL, Ortiz-Londono CF, et al. TBR1 regulates autism risk genes in the developing neocortex. Genome Res. 2016;26:1013–22.
Hsueh YP, Wang TF, Yang FC, Sheng M. Nuclear translocation and transcription regulation by the membrane-associated guanylate kinase CASK/LIN-2. Nature. 2000;404:298–302.
Hevner RF, Hodge RD, Daza RA, Englund C. Transcription factors in glutamatergic neurogenesis: conserved programs in neocortex, cerebellum, and adult hippocampus. Neurosci Res. 2006;55:223–33.
Bedogni F, Hodge RD, Elsen GE, Nelson BR, Daza RA, Beyer RP, et al. Tbr1 regulates regional and laminar identity of postmitotic neurons in developing neocortex. Proc Natl Acad Sci USA. 2010;107:13129–34.
Chuang HC, Huang TN, Hsueh YP. Neuronal excitation upregulates Tbr1, a high-confidence risk gene of autism, mediating Grin2b expression in the adult brain. Front Cell Neurosci. 2014;8:280.
Han W, Kwan KY, Shim S, Lam MM, Shin Y, Xu X, et al. TBR1 directly represses Fezf2 to control the laminar origin and development of the corticospinal tract. Proc Natl Acad Sci USA. 2011;108:3041–6.
McKenna WL, Betancourt J, Larkin KA, Abrams B, Guo C, Rubenstein JL, et al. Tbr1 and Fezf2 regulate alternate corticofugal neuronal identities during neocortical development. J Neurosci. 2011;31:549–64.
Magri C, Piovani G, Pilotta A, Michele T, Buzi F, Barlati S. De novo deletion of chromosome 2q24.2 region in a mentally retarded boy with muscular hypotonia. Eur J Med Genet. 2011;54:361–4.
Neale BM, Kou Y, Liu L, Ma’ayan A, Samocha E, Sabo A, et al. Patterns and rates of exonic de novo mutations in autism spectrum disorders. Nature. 2012;485:242–5.
O’Roak BJ, Vives L, Girirajan S, Karakoc E, Krumm N, Coe BP, et al. Sporadic autism exomes reveal a highly interconnected protein network of de novo mutations. Nature. 2012;485:246–50.
O’Roak BJ, Vives L, Fu W, Egertson JD, Stanaway IB, Phelps IG, et al. Multiplex targeted sequencing identifies recurrently mutated genes in autism spectrum disorders. Science. 2012;338:1619–22.
O’Roak BJ, Stessman HA, Boyle EA, Witherspoon KT, Martin B, Lee C, et al. Recurrent de novo mutations implicate novel genes underlying simplex autism risk. Nat Commun. 2014;5:5595.
Gilissen C, Hehir-Kwa JY, Thung DT, Witherspoon KT, Martin B, Lee C, et al. Genome sequencing identifies major causes of severe intellectual disability. Nature. 2014;511:344–7.
Hamdan FF, Srour M, Capo-Chichi JM, Daoud H, Nassif C, Patry L, et al. De novo mutations in moderate or severe intellectual disability. PLoS Genet. 2014;10:e1004772.
Traylor RN, Dobyns WB, Rosenfeld JA, Wheeler P, Spence JE, Bandholz AM, et al. Investigation of TBR1 hemizygosity: four individuals with 2q24 microdeletions. Mol Syndromol. 2012;3:102–12.
Palumbo O, Fichera M, Palumbo P, Rizzo R, Mazzolla E, Cocuzza DM, et al. TBR1 is the candidate gene for intellectual disability in patients with a 2q24.2 interstitial deletion. Am J Med Genet A. 2014;164A:828–33.
McDermott JH, Study DDD, Clayton-Smith J, Briggs TA. The TBR1-related autistic-spectrum-disorder phenotype and its clinical spectrum. Eur J Med Genet. 2017;17:30427–5.
Vegas N, Cavallin M, Kleefstra T, de Boer L, Philbert M, Maillard C, et al. Mutations in TBR1 gene leads to cortical malformations and intellectual disability. Eur J Med Genet. 2018;61:759–764
Deriziotis P, O’Roak BJ, Graham SA, Estruch SB, Dimitropoulou D, Bernier RA, et al. De novo TBR1 mutations in sporadic autism disrupt protein functions. Nat Commun. 2014;5:4954.
Den Hoed J, Sollis E, Venselaar H, Estruch SB, Deriziotis P, Fisher SE. Functional characterization of TBR1 variants in neurodevelopmental disorder. Sci Rep. 2018;8:14279.
Sobreira N, Schiettecatte F, Valle D, Hamosh A. GeneMatcher: a matching tool for connecting investigators with an interest in the same gene. Hum Mutat. 2015;36:928–30.
Tanaka AJ, Cho MT, Millan F, Juusola J, Retterer K, Joshi C, et al. Mutations in SPATA5 are associated with microcephaly, intellectual disability, seizures, and hearing loss. Am J Hum Genet. 2015;97:457–64.
Thevenon J, Duffourd Y, Masurel-Paulet A, Lefebvre M, Feillet F, El Chehadeh-Djebbar S, et al. Diagnostic odyssey in severe neurodevelopmental disorders: toward clinical whole-exome sequencing as a first-line diagnostic test. Clin Genet. 2016;89:700–7.
Yang Y, Muzny DM, Reid JG, Bainbridge MN, Willis A, Ward PA, et al. Clinical whole-exome sequencing for the diagnosis of mendelian disorders. N. Engl J Med. 2013;369:1502–11.
Tarailo-Graovac M, Shyr C, Ross CJ, Horvath GA, Salvarinova R, Ye XC, et al. Exome Sequencing and the Management of Neurometabolic Disorders. N. Engl J Med. 2016;374:2246–55.
Ruggeri G, Timms AE, Cheng C, Weiss A, Kollros P, Chapman T, et al. Bi-allelic mutations of CCDC88C are a rare cause of severe congenital hydrocephalus. Am J Med Genet A. 2018;176:676–81.
Bowling KM, Thompson ML, Amaral MD, Finnila CR, Hiatt SM, Engel KL, et al. Genomic diagnosis for children with intellectual disability and/or developmental delay. Genome Med. 2017;9:43.
Boyle MP, Bernard A, Thompson CL, Ng L, Boe A, Mortrud M, et al. Cell-type-specific consequences of Reelin deficiency in the mouse neocortex, hippocampus, and amygdala. J Comp Neurol. 2011;519:2061–89.
Hodge RD, Kowalczyk TD, Wolf SA, Encinas JM, Rippey C, Enikolopov G, et al. Intermediate progenitors in adult hippocampal neurogenesis: Tbr2 expression and coordinate regulation of neuronal output. J Neurosci. 2008;28:3707–17.
Hodge RD, Garcia AJ, Elsen GE, Nelson BR, Mussar KE, Reiner SL, et al. Tbr2 expression in Cajal-Retzius cells and intermediate neuronal progenitors is required for morphogenesis of the dentate gyrus. J Neurosci. 2013;33:4165–80.
Ha S, Tripathi PP, Mihalas AB, Hevner RF, Beier DR. C-terminal region truncation of RELN disrupts an interaction with VLDLR, causing abnormal development of the cerebral cortex and hippocampus. J Neurosci. 2017;37:960–71.
Burrage LC, Eble TN, Hixson PM, Roney EK, Cheung SW, Franco LM. A mosaic 2q24.2 deletion narrows the critical region to a 0.4 Mb interval that includes TBR1, TANK, and PSMD14. Am J Med Genet A. 2013;161A:841–4.
Karoglan A, Schanze D, Bär C, Muschke P, Zenker M, Schanze I. A 2q24.2 microdeletion containing TANK as novel candidate gene for intellectual disability. Am J Med Genet A. 2019;179:832–6.
Nosten-Bertrand M, Kappeler C, Dinocourt C, Denis C, Germain J, Phan Dinh Tuy F, et al. Epilepsy in Dcx knockout mice associated with discrete lamination defects and enhanced excitability in the hippocampus. PLoS ONE. 2008;3:e2473.
Acknowledgements
This study makes use of data shared through the PhenomeCentral repository. Funding for PhenomeCentral was provided by Genome Canada and Canadian Institute of Health Research (CIHR). Research reported in this publication was supported by the French Ministry of Health (PHRC national 2008/2008-A00515-50), the Regional Council of Burgundy/Dijon University hospital (Plan d’Actions Régionales pour l’Innovation PARI 2012, AOI 2013), the European Union through the PO FEDER-FSE Bourgogne 2014/2020 programs (FEDER 2015), the National Institute of Neurological Disorders and Stroke (NINDS) under award number K08NS092898 and Jordan’s Guardian Angels (to GM), by NIH Grants R01 NS092339 and R01 NS085081 (to RFH), and by NHGRI grant UM1HG007301 to GMC. The CAUSES Study is funded by Mining for Miracles, British Columbia Children’s Hospital Foundation and Genome British Columbia. Several authors of this publication are members of the European Reference Network for Developmental Anomalies and Intellectual Disability (ERN-ITHACA). Proofreading of the manuscript was provided by Suzanne Rankin from the Dijon University Hospital.
CAUSES Study
Shelin Adam, Christèle du Souich, Alison M. Elliott, Anna Lehman, Jill Mwenifumbo, Tanya N. Nelson, Clara van Karnebeek, Jan M. Friedman.
Author information
Authors and Affiliations
Consortia
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Nambot, S., Faivre, L., Mirzaa, G. et al. De novo TBR1 variants cause a neurocognitive phenotype with ID and autistic traits: report of 25 new individuals and review of the literature. Eur J Hum Genet 28, 770–782 (2020). https://doi.org/10.1038/s41431-020-0571-6
Received:
Revised:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41431-020-0571-6
This article is cited by
-
Shared and divergent alteration of whole-brain connectivity and sensory deficits in multiple autism mouse models
Molecular Psychiatry (2025)
-
Possible roles of deep cortical neurons and oligodendrocytes in the neural basis of human sociality
Anatomical Science International (2024)
-
Dietary zinc supplementation rescues fear-based learning and synaptic function in the Tbr1+/− mouse model of autism spectrum disorders
Molecular Autism (2022)
-
Tbr1 Misexpression Alters Neuronal Development in the Cerebral Cortex
Molecular Neurobiology (2022)
-
Reply to Hsueh YP et al.
European Journal of Human Genetics (2020)