Abstract
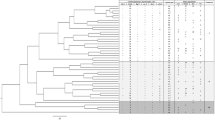
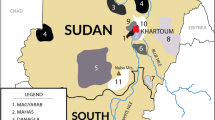
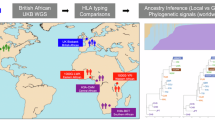
Sudan, a northeastern African country, is characterized by high levels of cultural, linguistic, and genetic diversity, which is believed to be affected by continuous migration from neighboring countries. Consistent with such demographic effect, genome-wide SNP data revealed a shared ancestral component among Sudanese Afro-Asiatic speaking groups and non-African populations, mainly from West Asia. Although this component is shared among all Afro-Asiatic speaking groups, the extent of this sharing in Semitic groups, such as Sudanese Arab, is still unknown. Using genotypes of six polymorphic human leukocyte antigen (HLA) genes (i.e., HLA-A, -C, -B, -DRB1, -DQB1, and -DPB1), we examined the genetic structure of eight East African ethnic groups with origins in Sudan, South Sudan, and Ethiopia. We identified informative HLA alleles using principal component analysis, which revealed that the two Semitic groups (Gaalien and Shokrya) constituted a distinct cluster from the other Afro-Asiatic speaking groups in this study. The HLA alleles that distinguished Semitic Arabs co-exist in the same extended HLA haplotype, and those alleles are in strong linkage disequilibrium. Interestingly, we find the four-locus haplotype “C*12:02-B*52:01-DRB1*15:02-DQB1*06:01” exclusively in non-African populations and it is widely spread across Asia. The identification of this haplotype suggests a gene flow from Asia, and likely these haplotypes were brought to Africa through back migration from the Near East. These findings will be of interest to biomedical and anthropological studies that examine the demographic history of northeast Africa.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Elhassan N, Gebremeskel EI, Elnour MA, Isabirye D, Okello J, Hussien A, et al. The episode of genetic drift defining the migration of humans out of Africa as derived from a large East African population size. PLoS One. 2014;9:e97674.
Tishkoff SA, Reed FA, Friedlaender FR, Ehret C, Ranciaro A, Froment A, et al. The genetic structure and history of Africans and African Americans. Science. 2009;324:1035–44.
Hassan HY, Underhill PA, Cavalli‐Sforza LL, Ibrahim ME. Y‐chromosome variation among Sudanese: restricted gene flow, concordance with language, geography, and history. Am J Phys Anthropol. 2008;137:316–23.
Hodgson JA, Mulligan CJ, Al-Meeri A, Raaum RL. Early Back-to-Africa migration into the Horn of Africa. PLoS Genet. 2014;10:e1004393.
Dobon B, Hassan HY, Laayouni H, Luisi P, Ricano-Ponce I, Zhernakova A, et al. The genetics of East African populations: a Nilo-Saharan component in the African genetic landscape. Sci Rep. 2015;5:9996.
Rodriguez-Flores JL, Fakhro K, Agosto-Perez F, Ramstetter MD, Arbiza L, Vincent TL, et al. Indigenous Arabs are descendants of the earliest split from ancient Eurasian populations. Genome Res. 2016;26:151–62.
Hollfelder N, Schlebusch CM, Günther T, Babiker H, Hassan HY, Jakobsson M. Northeast African genomic variation shaped by the continuity of indigenous groups and Eurasian migrations. PLoS Genet. 2017;13:e1006976.
Greenberg JH. The languages of Africa. 1st ed. Bloomington, Indiana, US: Indiana University; 1963.
Eberhard DM, Simons GF, Fennig CD. Ethnologue: languages of Africa and Europe. 22nd ed. Summer Institute of Linguistics, Academic Publications; 2019.
Babiker HM, Schlebusch CM, Hassan HY, Jakobsson M. Genetic variation and population structure of Sudanese populations as indicated by 15 Identifiler sequence-tagged repeat (STR) loci. Investigative Genet. 2011;2:12–12.
Hassan, HY. Genetic patterns of Y-chromosome and mitochondrial DNA variation, with implications to the peopling of the Sudan. PhD thesis, University of Khartoum; 2009.
Robinson J, Halliwell JA, Hayhurst JD, Flicek P, Parham P, Marsh SGE. The IPD and IMGT/HLA database: allele variant databases. Nucleic Acids Res. 2015;43:D423–31.
Radwan J, Babik W, Kaufman J, Lenz TL, Winternitz J. Advances in the evolutionary understanding of MHC polymorphism. Trends Genet. 2020;36:298–311.
Sanchez-Mazas A, Meyer D. The relevance of HLA sequencing in population genetics studies. J Immunol Res. 2014;2014:1–12.
Hernández-Frederick CJ, Cereb N, Giani AS, Ruppel J, Maraszek A, Pingel J, et al. Detection of 549 new HLA alleles in potential stem cell donors from the United States, Poland and Germany: detection of 549 new HLA alleles. HLA. 2016;87:31–5.
Rizvi SAH, Naqvi SA, Hussain Z, Hashmi A, Akhtar F, Hussain M, et al. Renal transplantation in developing countries. Kidney Int. 2003;63:S96–100.
Magzoub MM, Stephens HA, Sachs JA, Biro PA, Cutbush S, Wu Z, et al. HLA-DP polymorphism in Sudanese controls and patients with insulin-dependent diabetes mellitus. Tissue Antigens. 1992;40:64–8.
Dafalla AM, McCloskey DJ, Alemam AA, Ibrahim AA, Babikir AM, Gasmelseed N, et al. HLA polymorphism in Sudanese renal donors. Saudi J kidney Dis Transplant. 2011;22:834–40.
Hosomichi K, Jinam TA, Mitsunaga S, Nakaoka H, Inoue I. Phase-defined complete sequencing of the HLA genes by next-generation sequencing. BMC Genomics. 2013;14:355.
González-Galarza FF, Takeshita LY, Santos EJ, Kempson F, Maia MHT, Silva ALSda, et al. Allele frequency net 2015 update: new features for HLA epitopes, KIR and disease and HLA adverse drug reaction associations. Nucleic Acids Res. 2015;43:D784–8.
Ahmadloo S, Nakaoka H, Hayano T, Hosomichi K, You H, Utsuno E, et al. Rapid and cost-effective high-throughput sequencing for identification of germline mutations of BRCA1 and BRCA2. J Hum Genet. 2017;62:561–7.
Lancaster AK, Single RM, Solberg OD, Nelson MP, Thomson G. PyPop update—a software pipeline for large-scale multilocus population genomics. Tissue Antigens. 2007;69:192–7.
Excoffier L, Lischer HE. Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Mol Ecol Resour. 2010;10:564–7.
Guo SW, Thompson EA. Performing the exact test of Hardy-Weinberg proportion for multiple alleles. Biometrics. 1992;48:361–72.
Saitou N, Nei M. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evolution. 1987;4:406–25.
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S. MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evolution. 2013;30:2725–9.
Ewens WJ. The sampling theory of selectively neutral alleles. Theor Popul Biol. 1972;3:87–112.
Salamon H, Klitz W, Easteal S, Gao X, Erlich HA, Fernandez-Vina M, et al. Evolution of HLA class II molecules: allelic and amino acid site variability across populations. Genetics. 1999;152:393–400.
Slatkin M. An exact test for neutrality based on the Ewens sampling distribution. Genetical Res. 1994;64:71–4.
Excoffier L, Slatkin M. Maximum-likelihood estimation of molecular haplotype frequencies in a diploid population. Mol Biol Evol. 1995;12:921–7.
R Core Team. R: a language and environment for statistical computing. 2013. Available from: http://www.R-project.org.
Vina MAF, Hollenbach JA, Lyke KE, Sztein MB, Maiers M, Klitz W, et al. Tracking human migrations by the analysis of the distribution of HLA alleles, lineages and haplotypes in closed and open populations. Philos Trans Biol Sci. 2012;367:820–9.
Solberg OD, Mack SJ, Lancaster AK, Single RM, Tsai Y. Balancing selection and heterogeneity across the classical human leukocyte antigen loci: a meta-analytic review of 497 population studies. Hum Immunol. 2008;69:443–64.
Bryc K, Velez C, Karafet T, Moreno-Estrada A, Reynolds A, Auton A, et al. Genome-wide patterns of population structure and admixture among Hispanic/Latino populations. Proc Natl Acad Sci USA. 2010;107(Supplement_2):8954–61.
Tishkoff SA, Williams SM. Genetic analysis of African populations: human evolution and complex disease. Nat Rev Genet. 2002;3:611–21.
Buhler S, Sanchez-Mazas A. HLA DNA sequence variation among human populations: molecular signatures of demographic and selective events. PLoS One. 2011;6:e14643.
Černý V, Kulichová I, Poloni ES, Nunes JM, Pereira L, Mayor A, et al. Genetic history of the African Sahelian populations. HLA. 2018;91:153–66.
MacMichael HA.Tribes North Cent Kordofan. 1914;46:378.
Metz HC. Library of Congress Federal Research Division, Thomas Leiper Kane Collection. Sudan: a country study. 4th ed. Washington, DC: Federal Research Division, Library of Congress; 1992 (Area handbook series).
Di Giacomo F, Luca F, Popa LO, Akar N, Anagnou N, Banyko J, et al. Y chromosomal haplogroup J as a signature of the post-neolithic colonization of Europe. Hum Genet. 2004;115:357–71.
Sanchez-Mazas A, Djoulah S, Busson M, Le Monnier de Gouville I, Poirier JC, Dehay C, et al. A linkage disequilibrium map of the MHC region based on the analysis of 14 loci haplotypes in 50 French families. Eur J Hum Genet. 2000;8:33–41.
Abi-Rached L, Jobin MJ, Kulkarni S, McWhinnie A, Dalva K, Gragert L, et al. The shaping of modern human immune systems by multiregional admixture with archaic humans. Science. 2011;334:89–94.
Yasukochi Y, Yasukochi Y, Ohashi J, Ohashi J. Elucidating the origin of HLA-B73 allelic lineage: did modern humans benefit by archaic introgression? Immunogenetics. 2017;69:63–7.
Acknowledgements
The authors would like to thank all participants who generously donated their DNA samples. MGN was supported by a Spinoza grant of the Netherlands Organization for scientific research.
Author information
Authors and Affiliations
Contributions
II, NH, and AW conceived and designed the study. II and NMG provided materials and supervision. AW performed the experiments with contribution from HHY, and JM. HHY collected participants data with contribution from TH. AW, HN, performed HLA data analysis and interpretation with contribution from HHY, HK, and AMH. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Aamer, W., Hassan, H.Y., Nakaoka, H. et al. Analysis of HLA gene polymorphisms in East Africans reveals evidence of gene flow in two Semitic populations from Sudan. Eur J Hum Genet 29, 1259–1271 (2021). https://doi.org/10.1038/s41431-021-00845-6
Received:
Revised:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41431-021-00845-6