Abstract
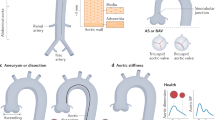
Thoracic aortic aneurysm with or without dissection (TAAD) can be broadly categorized as syndromic TAAD (sTAAD) and isolated TAAD (iTAAD). sTAAD and is highly correlated with genetics. However, although the incidence of iTAAD is much higher, its monogenic contribution is not yet clear. Here, we sequenced 15 known TAAD genes for 578 iTAAD cases from four cardiac centers in China and found that 10.6% patients with a pathogenic/likely pathogenic (P/LP) variant. Other 7.27% of patients carried variants of uncertain significance in these target genes. We further investigated the correlations among genetics, clinical features, and long-term outcomes. Genetic patients showed younger onset ages (P = 1.31E-13) and larger aortic diameter (P = 1.00E-6), with the youngest age in patients with FBN1 P/LP variants. Monogenic variants were also associated with more aortic segments involved (P = 0.043) and complicated with initial dissection (P = 4.50E-5), especially for genetic patients with non-FBN1 P/LP variants. MACEs occurred in 14.9% patients during follow-up of median 55 months. Genetic status (P = 0.001) and initial dissection (P = 3.00E-6) were two major risk factors for poor prognosis. Early onset age was associated with MACEs in non-genetic cases without initial dissection (P = 0.005). Our study revealed the monogenic contribution in known TAAD genes to iTAAD patients. The genotype–phenotype correlations may complement the risk stratification of iTAAD patients and identification of higher risk subgroups, as well as assist the development of tailored precision medicine in iTAAD.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Cury M, Zeidan F, Lobato AC. Aortic disease in the young: genetic aneurysm syndromes, connective tissue disorders, and familial aortic aneurysms and dissections. Int J Vasc Med. 2013;2013:267215.
Quintana RA, Taylor WR. Cellular mechanisms of aortic aneurysm formation. Circ Res. 2019;124:607–18.
Isselbacher EM, Lino Cardenas CL, Lindsay ME. Hereditary influence in thoracic aortic aneurysm and dissection. Circulation. 2016;133:2516–28.
Prakash SK, LeMaire SA, Guo DC, Russell L, Regalado ES, Golabbakhsh H, et al. Rare copy number variants disrupt genes regulating vascular smooth muscle cell adhesion and contractility in sporadic thoracic aortic aneurysms and dissections. Am J Hum Genet. 2010;87:743–56.
LeMaire SA, McDonald ML, Guo DC, Russell L, Miller CC 3rd, Johnson RJ, et al. Genome-wide association study identifies a susceptibility locus for thoracic aortic aneurysms and aortic dissections spanning FBN1 at 15q21.1. Nat Genet. 2011;43:996–1000.
Pyeritz RE. Heritable thoracic aortic disorders. Curr Opin Cardiol. 2014;29:97–102.
Blunder S, Messner B, Aschacher T, Zeller I, Turkcan A, Wiedemann D, et al. Characteristics of TAV- and BAV-associated thoracic aortic aneurysms-smooth muscle cell biology, expression profiling, and histological analyses. Atherosclerosis. 2012;220:355–61.
Norton E, Yang B. Managing thoracic aortic aneurysm in patients with bicuspid aortic valve based on aortic root-involvement. Front Physiol. 2017;8:397.
Teixido-Tura G, Franken R, Galuppo V, Gutierrez Garcia-Moreno L, Borregan M, Mulder BJ, et al. Heterogeneity of aortic disease severity in patients with Loeys-Dietz syndrome. Heart. 2016;102:626–32.
Franken R, den Hartog AW, Radonic T, Micha D, Maugeri A, van Dijk FS, et al. Beneficial outcome of losartan therapy depends on type of FBN1 mutation in Marfan syndrome. Circ Cardiovasc Genet. 2015;8:383–8.
Johnston KW, Rutherford RB, Tilson MD, Shah DM, Hollier L, Stanley JC. Suggested standards for reporting on arterial aneurysms. Subcommittee on Reporting Standards for Arterial Aneurysms, Ad Hoc Committee on Reporting Standards, Society for Vascular Surgery and North American Chapter, International Society for Cardiovascular Surgery. J Vasc Surg. 1991;13:452–8.
Campens L, Demulier L, De Groote K, Vandekerckhove K, De Wolf D, Roman MJ, et al. Reference values for echocardiographic assessment of the diameter of the aortic root and ascending aorta spanning all age categories. Am J Cardiol. 2014;114:914–20.
Xiong HY, Alipanahi B, Lee LJ, Bretschneider H, Merico D, Yuen RK, et al. RNA splicing. The human splicing code reveals new insights into the genetic determinants of disease. Science. 2015;347:1254806.
Jagadeesh KA, Wenger AM, Berger MJ, Guturu H, Stenson PD, Cooper DN, et al. M-CAP eliminates a majority of variants of uncertain significance in clinical exomes at high sensitivity. Nat Genet. 2016;48:1581–6.
Ioannidis NM, Rothstein JH, Pejaver V, Middha S, McDonnell SK, Baheti, et al. REVEL: an ensemble method for predicting the pathogenicity of rare missense variants. Am J Hum Genet. 2016;99:877–85.
Richards S, Aziz N, Bale S, Bick D, Das S, Gastier-Foster J, et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med. 2015;17:405–24.
Franken R, Groenink M, de Waard V, Feenstra HM, Scholte AJ, van den Berg MP, et al. Genotype impacts survival in Marfan syndrome. Eur Heart J. 2016;37:3285–90.
Akinrinade O, Ollila L, Vattulainen S, Tallila J, Gentile M, Salmenpera P, et al. Genetics and genotype-phenotype correlations in Finnish patients with dilated cardiomyopathy. Eur Heart J. 2015;36:2327–37.
Haas J, Frese KS, Peil B, Kloos W, Keller A, Nietsch R, et al. Atlas of the clinical genetics of human dilated cardiomyopathy. Eur Heart J. 2015;36:1123–35a.
Verhagen JMA, Kempers M, Cozijnsen L, Bouma BJ, Duijnhouwer AL, Post JG, et al. Expert consensus recommendations on the cardiogenetic care for patients with thoracic aortic disease and their first-degree relatives. Int J Cardiol. 2018;258:243–8.
Mizuguchi T, Collod-Beroud G, Akiyama T, Abifadel M, Harada N, Morisaki T, et al. Heterozygous TGFBR2 mutations in Marfan syndrome. Nat Genet. 2004;36:855–60.
Ki CS, Jin DK, Chang SH, Kim JE, Kim JW, Park BK, et al. Identification of a novel TGFBR2 gene mutation in a Korean patient with Loeys-Dietz aortic aneurysm syndrome; no mutation in TGFBR2 gene in 30 patients with classic Marfan’s syndrome. Clin Genet. 2005;68:561–3.
Yetman AT, Beroukhim RS, Ivy DD, Manchester D. Importance of the clinical recognition of Loeys-Dietz syndrome in the neonatal period. Pediatrics. 2007;119:e1199–202.
Watanabe Y, Sakai H, Nishimura A, Miyake N, Saitsu H, Mizuguchi T, et al. Paternal somatic mosaicism of a TGFBR2 mutation transmitting to an affected son with Loeys-Dietz syndrome. Am J Med Genet A. 2008;146A:3070–4.
Loeys BL, Chen J, Neptune ER, Judge DP, Podowski M, Holm T, et al. A syndrome of altered cardiovascular, craniofacial, neurocognitive and skeletal development caused by mutations in TGFBR1 or TGFBR2. Nat Genet. 2005;37:275–81.
Pannu H, Fadulu VT, Chang J, Lafont A, Hasham SN, Sparks E, et al. Mutations in transforming growth factor-beta receptor type II cause familial thoracic aortic aneurysms and dissections. Circulation. 2005;112:513–20.
Colzani E, Liljegren A, Johansson AL, Adolfsson J, Hellborg H, Hall PF, et al. Prognosis of patients with breast cancer: causes of death and effects of time since diagnosis, age, and tumor characteristics. J Clin Oncol. 2011;29:4014–21.
Anders CK, Hsu DS, Broadwater G, Acharya CR, Foekens JA, Zhang Y, et al. Young age at diagnosis correlates with worse prognosis and defines a subset of breast cancers with shared patterns of gene expression. J Clin Oncol. 2008;26:3324–30.
Micha D, Guo DC, Hilhorst-Hofstee Y, van Kooten F, Atmaja D, Overwater E, et al. SMAD2 mutations are associated with arterial aneurysms and dissections. Hum Mutat. 2015;36:1145–9.
Kuang SQ, Medina-Martinez O, Guo DC, Gong L, Regalado ES, Reynolds CL, et al. FOXE3 mutations predispose to thoracic aortic aneurysms and dissections. J Clin Investig. 2016;126:948–61.
Lee VS, Halabi CM, Hoffman EP, Carmichael N, Leshchiner I, Lian CG, et al. Loss of function mutation in LOX causes thoracic aortic aneurysm and dissection in humans. Proc Natl Acad Sci USA. 2016;113:8759–64.
Li Y, Gao S, Han Y, Song L, Kong Y, Jiao Y, et al: Variants of focal adhesion scaffold genes cause thoracic aortic aneurysm. Circ Res. 2021;128:8–23.
Acknowledgements
We thank all the patients for their participation in this study.
Funding
This study was supported by grants from the National Key Research and Development Program of China (2016YFC0903000), the National Science Foundation of China (81800218 and 81930014), the Key Laboratory of Remodeling-Related Cardiovascular Diseases, Ministry of Education of China (PXM2014-014226-000012), International Cooperation Project from the Ministry of Science and Technology of China (2015DFA31070), and the Beijing Collaborative Innovative Research Center for Cardiovascular Diseases.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Li, Y., Kong, Y., Duan, W. et al. Evaluating the monogenic contribution and genotype–phenotype correlation in patients with isolated thoracic aortic aneurysm. Eur J Hum Genet 29, 1129–1138 (2021). https://doi.org/10.1038/s41431-021-00857-2
Received:
Revised:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41431-021-00857-2