Abstract
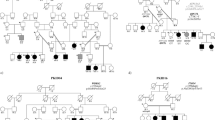
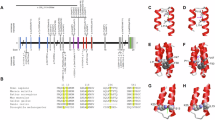
Intellectual disability (ID) represents an extremely heterogeneous group of disorders, characterized by significant limitations in intellectual function and adaptive behavior. Among the monogenic causes, autosomal recessive genes (ARID) are responsible for more than 50% of ID. Here, we report a novel in-frame homozygous deletion variant [c.730_753del; p.(Ala244_Gly251del)] in SOX4 (sex-determining region Y-related high-mobility group box 4), segregating with moderate to severe ID, hypotonia, and developmental delay in a Pakistani family. Our identified variant p.(Ala244_Gly251del) is predicted to remove evolutionarily conserved residues from the interdomain region and may destabilize the protein secondary structure. SOX4 belongs to group C of the SOX transcription regulating family known to be involved in early embryo development. Single-cell RNA data analysis of developing telencephalon revealed highly overlapping expression of SOX4 with SOX11 and DCX, known neurogenesis regulators. Our study expands the mutational landscape of SOX4 and the repertoire of the known genetic causes of ARID.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Data availability
The variant has been submitted to ClinVar database (accession number: SCV001571323).
References
Laudet V, Stehelin D, Clevers H. Ancestry and diversity of the HMG box superfamily. Nucleic Acids Res. 1993;21:2493–501.
Koopman P, Schepers G, Brenner S, Venkatesh B. Origin and diversity of the SOX transcription factor gene family: genome-wide analysis in Fugu rubripes. Gene. 2004;328:177–86.
Zawerton A, Yao B, Yeager JP, Pippucci T, Haseeb A, Smith JD, et al. De novo SOX4 variants cause a neurodevelopmental disease associated with mild dysmorphism. Am J Hum Genet. 2019;104:246–59.
Wilson M, Koopman P. Matching SOX: partner proteins and co-factors of the SOX family of transcriptional regulators. Curr Opin Genet Dev. 2002;12:441–6.
Richard EM, Santos-Cortez RLP, Faridi R, Rehman AU, Lee K, Shahzad M, et al. Global genetic insight contributed by consanguineous Pakistani families segregating hearing loss. Hum Mutat. 2019;40:53–72.
Riazuddin S, Hussain M, Razzaq A, Iqbal Z, Shahzad M, Polla D, et al. Exome sequencing of Pakistani consanguineous families identifies 30 novel candidate genes for recessive intellectual disability. Mol Psychiatry. 2017;22:1604–14.
Chen C, Lee GA, Pourmorady A, Sock E, Donoghue MJ. Orchestration of neuronal differentiation and progenitor pool expansion in the developing cortex by SoxC genes. J Neurosci. 2015;35:10629–42.
Hargrave M, Wright E, Kun J, Emery J, Cooper L, Koopman P. Expression of the Sox11 gene in mouse embryos suggests roles in neuronal maturation and epithelio‐mesenchymal induction. Developmental Dyn. 1997;210:79–86.
Ohshima T. Neuronal migration and protein kinases. Front Neurosci. 2015;8:458.
Lv X, Ren S-Q, Zhang X-J, Shen Z, Ghosh T, Xianyu A, et al. TBR2 coordinates neurogenesis expansion and precise microcircuit organization via Protocadherin 19 in the mammalian cortex. Nat Commun. 2019;10:1–15.
Cong Y, So V, Tijssen MA, Verbeek DS, Reggiori F, Mauthe M. WDR45, one gene associated with multiple neurodevelopmental disorders. Autophagy. 2021;12:1–16.
Hoser M, Potzner MR, Koch JM, Bösl MR, Wegner M, Sock E. Sox12 deletion in the mouse reveals nonreciprocal redundancy with the related Sox4 and Sox11 transcription factors. Mol Cell Biol. 2008;28:4675–87.
Dy P, Penzo-Mendez A, Wang H, Pedraza CE, Macklin WB, Lefebvre V. The three SoxC proteins—Sox4, Sox11 and Sox12—exhibit overlapping expression patterns and molecular properties. Nucleic Acids Res. 2008;36:3101–17.
Murakami A, Ishida S, Thurlow J, Revest J-M, Dickson C. SOX6 binds CtBP2 to repress transcription from the Fgf-3 promoter. Nucleic Acids Res. 2001;29:3347–55.
Marshall OJ, Harley VR. Identification of an interaction between SOX9 and HSP70. FEBS Lett. 2001;496:75–80.
Acknowledgements
We thank the PKMR225 family members for participating in this study. We would like to thank Muhammad Asad Usmani for his guidance and technical assistance.
Funding
This work was supported by National Institute of Neurological Disorders and Stroke, National Institutes of Health grant R01NS107428 to Saima Riazuddin.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval
This study was approved by the Institutional Review Boards (IRB) at the University of Maryland Baltimore, USA, and the Centre of Excellence in Molecular Biology (CEMB), University of The Punjab, Lahore, Pakistan.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Ghaffar, A., Rasheed, F., Rashid, M. et al. Biallelic in-frame deletion of SOX4 is associated with developmental delay, hypotonia and intellectual disability. Eur J Hum Genet 30, 243–247 (2022). https://doi.org/10.1038/s41431-021-00968-w
Received:
Revised:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41431-021-00968-w
This article is cited by
-
Mechanisms of Fat Mass and Obesity-Associated Protein in Regulating Cognitive Impairment Induced by Sevoflurane Anesthesia in Neonatal Rats
Neurochemical Research (2025)
-
A new system for variant classification?
European Journal of Human Genetics (2022)