Abstract
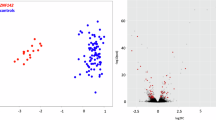
ZNF711 is one of eleven zinc-finger genes on the X chromosome that have been associated with X-linked intellectual disability. This association is confirmed by the clinical findings in 20 new cases in addition to 11 cases previously reported. No consistent growth aberrations, craniofacial dysmorphology, malformations or neurologic findings are associated with alterations in ZNF711. The intellectual disability is typically mild and coexisting autism occurs in half of the cases. Carrier females show no manifestations. A ZNF711-specific methylation signature has been identified which can assist in identifying new cases and in confirming the pathogenicity of variants in the gene.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The summarized, anonymized data for each subject in this study are described in the text. Publicly available DNA methylation datasets have been deposited in GEO, and include data referring to various developmental syndromes (e.g., Kabuki syndrome, Sotos syndrome, CHARGE syndrome, immunodeficiency-centromeric instability-facial anomalies (ICF) syndrome, Williams- Beuren syndrome, Chr7q11.23 duplication syndrome, BAFopathies, Down syndrome), a large cohort of unresolved subjects with developmental delay/intellectual disability and congenital abnormalities, and also several large cohorts of DNA methylation data from the general population. Data in the EpiSign Knowledge Database, including methylation data from this ZNF711 study cohort, are not available due to Research Ethics Board and institutional restrictions. EpiSignTM is proprietary software and is not publicly available.
Change history
01 December 2023
A Correction to this paper has been published: https://doi.org/10.1038/s41431-023-01499-2
References
Colleaux L, May M, Belougne J, Lepaslier D, Schwartz CE, Fontes M. Localisation of two candidate genes for mental retardation using a YAC physical map of the Xq21.1-21.2 subbands. J Med Genet. 1996;33:353–7.
May M, Colleaux L, Murgia A, Aylsworth A, Nussbaum R, Fontes M, et al. Molecular analysis of four males with mental retardation and deletions of Xq21 places the putative MR region in Xq21.1 between DXS233 and CHM. Hum Mol Genet. 1995;4:1465–6.
Tarpey PS, Smith R, Pleasance E, Whibley A, Edkins S, Hardy C, et al. A systematic, large-scale resequencing screen of X-chromosome coding exons in mental retardation. Nat Genet. 2009;41:535–43.
van der Werf IM, Dijck AV, Reyniers E, Helsmoortel C, Kumar AA, Kalscheuer VM, et al. Mutations in two large pedigrees highlight the role of ZNF711 in X-linked intellectual disability. Gene. 2017;605:92–8.
Delphin N, Hanein S, Taie LF, Zanlonghi X, Bonneau D, Moisan JP, et al. Intellectual disability associated with retinal dystrophy in the Xp11.3 deletion syndrome: ZNF674 on trial. Guilty or innocent? Eur J Hum Genet. 2012;20:352–6.
Piton A, Redin C, Mandel JL. XLID-causing mutations and associated genes challenged in light of data from large-scale human exome sequencing. Am J Hum Genet. 2013;93:368–83.
Neri G, Schwartz CE, Lubs HA, Stevenson RE. X-linked intellectual disability update 2017. Am J Med Genet A. 2018;176:1375–88.
Schwartz CE, Norris JW, Harr MH, Orrico A, Zackai EH. Mutations in ZFP92, a novel KRAB Zinc-finger protein, results in an X-linked intellectual disability and mitochondrial dysfunction disorder. American Society of Human Genetics Annual Meeting. 2018; 68: SB (abstract 54).
Hirata H, Nanda I, van Riesen A, McMichael G, Hu H, Hambrock M, et al. ZC4H2 mutations are associated with arthrogryposis multiplex cogenita and intellectual disability through impairment of central and peripheral synaptic plasticity. Am J Hum Genet. 2013;92:681–95.
May M, Hwang KS, Miles J, Williams C, Niranjan T, Kahler SG, et al. ZC4H2, an XLID gene, is required for the generation of a specific subset of CNS interneurons. Hum Mol Genet. 2015;17:4848–61.
Raymond FL, Tarpey PS, Edkins S, Tofts C, O’Meara S, Teague J, et al. Mutations in ZDHHC9, which encodes a palmitoyltransferase of NRAS and HRAS, cause X-linked mental retardation associated with a marfanoid habitus. Am J Hum Genet. 2007;80:982–7.
Baker K, Astle DE, Scerif G, Barnes J, Smith J, Moffat G, et al. Epilepsy, cognitive deficits and neuroanatomy in males with ZDHHC9 mutations. Ann Clin Trans Neurol. 2015;5:559–69.
Philips AK, Sirén A, Avela K, Somer M, Peippo M, Ahvenainen M, et al. X-exome sequencing in Finnish families with intellectual disability-four novel mutations and two novel syndromic phenotypes. Orphanet J Rare Dis. 2014;9:49.
Louie RJ, Tan QK, Gilner JB, Rogers RC, Younge N, Wechsler SB, et al. Novel pathogenic variants in FOXP3 in fetuses with echogenic bowel and skin desquamation identified by ultrasound. Am J Med Genet A. 2017;173:1219–25.
Larcher L, Norris JW, Lejuene EL, Buratti J, Mignot C, Garel C, et al. The complete loss of function of the SMS gene results in a severe form of Snyder-Robinson syndrome. Eur J Med Genet. 2020;63:103777.
Aref-Eshghi E, Kerkhof J, Pedro VP, France GD, Barat-Houari M, et al. Evaluation of DNA methylation episignatures for diagnosis and phenotype correlations in 42 Mendelian neurodevelopmental disorders. Am J Hum Genet. 2020;106:356–70.
Ho DE, Imai K, King G, Stuart EA. MatchIt: nonparametric preprocessing for parametric causal inference. J Stat Softw. 2011;42:8.
Ritchie ME, Phipson B, Wu D, Hu Y, Law CW, Shi W, et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 2015;43:e47.
Houseman EA, Accomando WP, Koestler DC, Christensen BC, Marsit CJ, Nelson HH, et al. DNA methylation arrays as surrogate measures of cell mixture distribution. BMC Bioinform. 2012;13:86.
Yntema HG, van den Helm B, Knoers NV, Smits AP, van Roosmalen T, Smeets DF, et al. X-linked mental retardation: Evidence for a recent mutation in a five-generation family (MRX65) linked to the pericentromeric region. Am J Med Genet. 1999;85:305–8.
Lossi AM, Laugier-Anfossi F, Depetris D, Gecz J, Gedeon A, Kooy F, et al. Abnormal expression of the KLF8 (ZNF741) gene in a female patient with an X;autosome translocation t(X;21)(p11.2;q22.3) and non-syndromic mental retardation. J Med Genet. 2002;39:113–7.
Cho G, Bhat SS, Gao J, Collins JS, Rogers RC, Simensen RJ, et al. Evidence that SIZN1 is a candidate X-linked mental retardation gene. Am J Med Genet A. 2008;146A:2644–50.
Steinback P, Horstmann W, Scholz W. Tandem duplication dup(X)(q13q22) in a male proband inherited from the mother showing mosaicism of X-Inactivation. Hum Genet. 1980;54:309–13.
Gabbett MT, Peters GB, Carmichael JM, Darmanian AP, Collins FA. Prader-Willi syndrome phenocopy due to duplication of Xq21.1-q21.31, with array CGH of the critical region. Clin Genet. 2008;73:353–9.
Martínez F, Roselló M, Mayo S, Monfort S, Oltra S, Orellana C. Duplication at Xq13.3-q21.1 with syndromic intellectual disability, a probably role for the ATRX gene. Am J Med Genet A. 2014;164A:918–23.
Linhares ND, Valadares ER, da Costa SS, Arantes RR, de Oliveira LR, Rosenberg C, et al. Inherited Xq13.2-q21.31 duplication in a boy with recurrent seizures and pubertal gynecomastia: clinical, chromosomal and aCGH characterization. Meta Gene. 2016;9:185–90.
Schenkel LC, Rodenhiser DI, Ainsworth PJ, Paré G, Sadikovic B. Clinical implications of the epigenome. Crit Rev Clin Lab Sci. 2016;53:147–65.
Aref-Eshghi E, Bend EG, Hood RL, Schenkel LC, Carere DA, Chakrabarti R, et al. BAFopathies’ DNA methylation epi-signatures demonstrate diagnostic utility and functional continuum of Coffin-Siris and Nicolaides-Baraitser syndromes. Nat Comm. 2018;9:4885.
Schenkel LC, Aref-Eshghi E, Skinner C, Ainsworth P, Lin H, Paré G, et al. Peripheral blood epi-signature of Claes-Jensen syndrome enables sensitive and specific identification of patients and healthy carriers with pathogenic mutations in KDM5C. Clin Epigenet. 2018;10:21.
Kernohan KD, Schenkel LC, Huang L, Smith A, Pare G, Ainsworth P, et al. Identification of a methylation profile for DNMT1-associated autosomal dominant cerebellar ataxia, deafness, and narcolepsy. Clin Epigenet. 2016;8:91.
Radio FC, Pang K, Ciolfi A, Levy MA, Hernández-García A, Pedace L, et al. SPEN haploinsufficiency causes a neurodevelopmental disorder overlapping proximal 1p36 deletion syndrome with an episignature of X chromosomes in females. Am J Hum Genet. 2021;108:502–16.
Sadikovic B, Aref-Eshghi E, Levy MA, Rodenhiser D. DNA methylation signatures in mendelian developmental disorders as a diagnostic bridge between genotype and phenotype. Epigenomics. 2019;11:563–75.
Sadikovic B, Levy MA, Kerkhof J, Aref-Eshghi E, Schenkel L, Stuart A, et al. Clinical epigenomics: genome-wide DNA methylation analysis for the diagnosis of Mendelian disorders. Genet Med. 2021;23:1065–74.
Tejada MI, Ibarluzea N. Non-syndromic X linked intellectual disability: Current knowledge in light of the recent advances in molecular and functional studies. Clin Genet. 2020;97:677–87.
Acknowledgements
We thank members of the families for participating in this research. Dedicated to the memory of Ethan Francis Schwartz, 1996–1998. Funding for this research was provided, in part, by NIH grant HD-26202 to CES, Curekids New Zealand to SPR, the Wellcome Trust Sanger Institute, the Greenwood Genetic Center Foundation, the South Carolina Department of Disabilities and Special Needs, and the London Health Sciences Molecular Diagnostics Development Fund and Genome Canada Genomic Applications Partnership Program Grant awarded to BS (Beyond Genomics: Assessing the Improvement in Diagnosis of Rare Diseases using Clinical Epigenomics in Canada, EpiSign-CAN).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval
The research protocol and consent were reviewed and approved by the Self Regional Healthcare Institutional Review Board (IRB).
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original online version of this article was revised: The Data availability statement was incorrect.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wang, J., Foroutan, A., Richardson, E. et al. Clinical findings and a DNA methylation signature in kindreds with alterations in ZNF711. Eur J Hum Genet 30, 420–427 (2022). https://doi.org/10.1038/s41431-021-01018-1
Received:
Revised:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41431-021-01018-1
This article is cited by
-
Combined single-cell profiling of chromatin–transcriptome and splicing across brain cell types, regions and disease state
Nature Biotechnology (2025)
-
No April fools in clinical genomics
European Journal of Human Genetics (2022)
-
ZNF711 puts a spell on DNA
European Journal of Human Genetics (2022)