Abstract
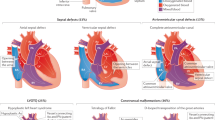
The etiology of congenital heart disease (CHD) is complex, comprising both genetic and environmental factors. Despite documented familial occurrences, the genetic etiology remains largely elusive. Trio exome sequencing identified a heterozygous FLT4 splice site variant in two families with respectively tetralogy of Fallot (TOF), and variable CHD comprising both the TOF spectrum and aortic coarctation. In the first family, Sanger sequencing on cDNA confirmed aberrant splicing for the c.985+1G > A variant. In the second family, transcriptome sequencing uncovered altered splicing for the c.1657+6T > C variant, despite normal targeted Sanger sequencing. In conclusion, our study establishes FLT4 splice site variants as a molecular cause of both left and right-sided isolated CHD, with incomplete penetrance. RNA-sequencing emerges as a valuable technique in unraveling the missing inheritability of CHD.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Data availability
Data sharing is not applicable to this article as no datasets were generated or analyzed during the current study. All variants have been submitted to the Clinvar database and can be accessed using the following accession numbers: SCV005407801 for NM_182925.5: c.1657+6T > C, and SCV005407803 for NM_182925.5: c.985+1G > A.
Change history
12 March 2025
The original online version of this article was revised: Author Tim Van Damme’s name was incorrectly written as Tim Vandamme.
01 April 2025
A Correction to this paper has been published: https://doi.org/10.1038/s41431-025-01831-y
References
Moons P, Sluysmans T, De Wolf D, Massin M, Suys B, Benatar A, et al. Congenital heart disease in 111 225 births in Belgium: birth prevalence, treatment and survival in the 21st century. Acta Paediatr. 2009;98:472–7. https://doi.org/10.1111/J.1651-2227.2008.01152.X
Jang MY, Patel PN, Pereira AC, Willcox JAL, Haghighi A, Tai AC, et al. Contribution of previously unrecognized RNA splice-altering variants to congenital heart disease. Circ Genom Precis Med. 2023;16:224. https://doi.org/10.1161/CIRCGEN.122.003924
Lambrechts D, Devriendt K, Driscoll DA, Goldmuntz E, Gewillig M, Vlietinck R, et al. Low expression VEGF haplotype increases the risk for tetralogy of Fallot: a family based association study. J Med Genet. 2005;42:519–22. https://doi.org/10.1136/JMG.2004.026443
Škorić-Milosavljević D, Lahrouchi N, Bosada FM, Dombrowsky G, Williams SG, Lesurf R, et al. Rare variants in KDR, encoding VEGF receptor 2, are associated with tetralogy of Fallot. Genet Med. 2021;23:1952–60. https://doi.org/10.1038/s41436-021-01212-y.
Kawasaki T, Kitsukawa T, Bekku Y, Matsuda Y, Sanbo M, Yagi T, et al. A requirement for neuropilin-1 in embryonic vessel formation. Development. 1999;126:4895–902. https://doi.org/10.1242/DEV.126.21.4895
Stalmans I, Lambrechts D, De Smet F, Jansen S, Wang J, Maity S, et al. VEGF: a modifier of the del22q11 (DiGeorge) syndrome? Nat Med. 2003;9:173–82. https://doi.org/10.1038/nm819.
Page DJ, Miossec MJ, Williams SG, Monaghan RM, Fotiou E, Cordell HJ, et al. Whole exome sequencing reveals the major genetic contributors to non-syndromic tetralogy of Fallot Europe PMC Funders Group. Circ Res. 2019;124:553–63. https://doi.org/10.1161/CIRCRESAHA.118.313250
Jin SC, Homsy J, Zaidi S, Lu Q, Morton S, Depalma SR, et al. Contribution of rare inherited and de novo variants in 2,871 congenital heart disease probands. Nat Genet. 2017. https://doi.org/10.1038/ng.3970.
Tabib A, Talebi T, Ghasemi S, Pourirahim M, Naderi N, Maleki M, et al. A novel stop-gain pathogenic variant in FLT4 and a nonsynonymous pathogenic variant in PTPN11 associated with congenital heart defects. Eur J Med Res. 2022;27:286. https://doi.org/10.1186/s40001-022-00920-8
Reuter MS, Jobling R, Chaturvedi RR, Manshaei R, Costain G, Heung T, et al. Haploinsufficiency of vascular endothelial growth factor related signaling genes is associated with tetralogy of Fallot. Genet Med. 2018;21:1001–7. https://doi.org/10.1038/s41436-018-0260-9.
Reuter MS, Chaturvedi RR, Jobling RK, Pellecchia G, Hamdan O, Sung WWL, et al. Clinical genetic risk variants inform a functional protein interaction network for tetralogy of Fallot. Circ Genom Precis Med. 2021;14:E003410. https://doi.org/10.1161/CIRCGEN.121.003410/FORMAT/EPUB
Karczewski KJ, Francioli LC, Tiao G, Cummings BB, Alföldi J, Wang Q.Genome Aggregation Database Consortium et al. The mutational constraint spectrum quantified from variation in 141,456 humans. Nature. 2020;581:19. https://doi.org/10.1038/s41586-020-2308-7
Richards S, Aziz N, Bale S, Bick D, Das S, et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med. https://doi.org/10.1038/gim.2015.30.
den Dunnen JT, Dalgleish R, Maglott DR, Hart RK, Greenblatt MS, Mcgowan-Jordan J, et al. HGVS recommendations for the description of sequence variants: 2016 update. Hum Mutat. 2016;37:564–9. https://doi.org/10.1002/HUMU.22981
Martin FJ, Amode MR, Aneja A, Austine-Orimoloye O, Azov AG, Barnes I, et al. Ensembl 2023. Nucleic Acids Res. 2023;51:D933–41. https://doi.org/10.1093/NAR/GKAC958
Ttir H, Robinson JT, Mesirov JP. Integrative genomics viewer (IGV): high-performance genomics data visualization and exploration. Brief Bioinform. https://doi.org/10.1093/bib/bbs017.
Paysan-Lafosse T, Blum M, Chuguransky S, Grego T, Azaro Pinto BL´, Salazar GA, et al. InterPro in 2022. Nucleic Acids Res. 2023;51. https://doi.org/10.1093/nar/gkac993.
Gordon K, Spiden SL, Connell FC, Brice G, Cottrell S, Short J, et al. FLT4/VEGFR3 and Milroy disease: novel mutations, a review of published variants and database update. Hum Mutat. 2012. https://doi.org/10.1002/humu.22223.
Monaghan RM, Naylor RW, Flatman D, Kasher PR, Williams SG, Keavney BD. FLT4 causes developmental disorders of the cardiovascular and lymphovascular systems via pleiotropic molecular mechanisms. Cardiovasc Res. 2024. https://doi.org/10.1093/cvr/cvae104.
Fontana F, Haack T, Reichenbach M, Knaus P, Puceat M, Abdelilah-Seyfried S. Antagonistic activities of Vegfr3/Flt4 and notch1b fine-tune mechanosensitive signaling during zebrafish cardiac valvulogenesis. Cell Rep. 2020;32. https://doi.org/10.1016/J.CELREP.2020.107883.
Truty R, Ouyang K, Rojahn S, Garcia S, Colavin A, Hamlington B, et al. Spectrum of splicing variants in disease genes and the ability of RNA analysis to reduce uncertainty in clinical interpretation. Am J Hum Genet. 2021;108:696. https://doi.org/10.1016/J.AJHG.2021.03.006
Wai HA, Lord J, Lyon M, Gunning A, Kelly H, Cibin P, et al. Blood RNA analysis can increase clinical diagnostic rate and resolve variants of uncertain significance. Genet Med. 2020;22. https://doi.org/10.1038/s41436.
Acknowledgements
The authors thank the families for their kind availability in sharing the findings within the scientific community.
Funding
This project was supported by a grant “Scientific research on heart diseases (2023)” from the Philanthropic Center Pelicano to BC and by a Research Grant of the Research Foundation—Flanders (G035620N) to BC. BC is a senior clinical investigator of the Research Foundation—Flanders.
Author information
Authors and Affiliations
Contributions
Conceptualization, MV, TV, SV, and BC; methodology, ED, LD, BM, SS, PC, and SV; validation, ED, LD, SS, PC, and BC; formal analysis, MV, ED, and LD; investigation, MV, ED, and TV; resources, LM, KD, KV, JP, ER, TV, and BC; data curation, ED, LD, BM, and SV; writing—original draft preparation, MV, and BC; writing—review and editing, all authors; visualization, MV, ED, and LD; supervision, BC; project administration, BC; funding acquisition, BC. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical approval
This study was conducted in accordance with the 1984 Declaration of Helsinki and its subsequent revisions. The legal guardians of the individuals involved in this study provided written informed consent for the disclosure of case details.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original online version of this article was revised: Author Tim Van Damme’s name was incorrectly written as Tim Vandamme.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Verlee, M., D’haenens, E., De Cock, L. et al. RNA-sequencing unveils FLT4 splice site variants in variable congenital heart disease. Eur J Hum Genet 33, 1085–1089 (2025). https://doi.org/10.1038/s41431-025-01788-y
Received:
Revised:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41431-025-01788-y