Abstract
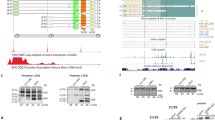
RNA-binding proteins play a key role in post-transcriptional events, such as mRNA splicing, transport, stability, translation and decay. Dysregulation of RNA life can have dramatic consequences. CELF RNA-binding proteins appear to be essential during embryo development. In this study, we identified 15 patients with heterozygous missense or loss-of-function variants in the CELF4 gene by exome or genome sequencing. All variants affecting the N-terminus of the protein are essential and sufficient for the RNA-binding and splicing activity or RRM domains. Most patients presented with neurodevelopmental disorders including global developmental delay/intellectual disability (11/14), seizures (9/15) and overweight/obesity (10/14) that began in childhood. Clinical features are similar to the reported celf4-mouse mutant phenotype. This study highlights the essential role of CELF4 in development and its involvement as a novel etiology of neurodevelopmental disorders with obesity.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Data availability
Additional data are available from the corresponding author upon reasonable request.
References
Prashad S, Gopal PP. RNA-binding proteins in neurological development and disease. RNA Biol. 2021;18:972–87.
Parra AS, Johnston CA. Emerging roles of RNA-binding proteins in neurodevelopment. J Dev Biol. 2022;10:23.
Zhou H, Mangelsdorf M, Liu J, Zhu L, Wu JY. RNA-binding proteins in neurological diseases. Sci China Life Sci. 2014;57:432–44.
Gebauer F, Schwarzl T, Valcárcel J, Hentze MW. RNA-binding proteins in human genetic disease. Nat Rev Genet. 2021;22:185–98.
Gerstberger S, Hafner M, Ascano M, Tuschl T. Evolutionary conservation and expression of human RNA-binding proteins and their role in human genetic disease. Adv Exp Med Biol. 2014;825:1–55.
Forman TE, Dennison BJC, Fantauzzo KA. The role of RNA-binding proteins in vertebrate neural crest and craniofacial development. J Dev Biol. 2021;9:34.
Nussbacher JK, Tabet R, Yeo GW, Lagier-Tourenne C. Disruption of RNA metabolism in neurological diseases and emerging therapeutic interventions. Neuron. 2019;102:294–320.
Wagnon JL, Briese M, Sun W, Mahaffey CL, Curk T, Rot G, et al. CELF4 regulates translation and local abundance of a vast set of mRNAs, including genes associated with regulation of synaptic function. PLoS Genet. 2012;8:e1003067.
Brinegar AE, Cooper TA. Roles for RNA-binding proteins in development and disease. Brain Res. 2016;1647:1–8.
Dasgupta T, Ladd AN. The importance of CELF control: molecular and biological roles of the CUG-BP, Elav-like family of RNA-binding proteins. Wiley Interdiscip Rev RNA. 2012;3:104–21.
Ladd AN, Charlet N, Cooper TA. The CELF family of RNA binding proteins is implicated in cell-specific and developmentally regulated alternative splicing. Mol Cell Biol. 2001;21:1285–96.
Salamon I, Park Y, Miškić T, Kopić J, Matteson P, Page NF, et al. Celf4 controls mRNA translation underlying synaptic development in the prenatal mammalian neocortex. Nat Commun. 2023;14:6025.
MacPherson MJ, Erickson SL, Kopp D, Wen P, Aghanoori MR, Kedia S, et al. Nucleocytoplasmic transport of the RNA-binding protein CELF2 regulates neural stem cell fates. Cell Rep. 2021;35:109226.
Itai T, Hamanaka K, Sasaki K, Wagner M, Kotzaeridou U, Brösse I, et al. De novo variants in CELF2 that disrupt the nuclear localization signal cause developmental and epileptic encephalopathy. Hum Mutat. 2021;42:66–76.
Dougherty JD, Maloney SE, Wozniak DF, Rieger MA, Sonnenblick L, Coppola G, et al. The disruption of Celf6, a gene identified by translational profiling of serotonergic neurons, results in autism-related behaviors. J Neurosci J Soc Neurosci. 2013;33:2732–53.
Wagnon JL, Mahaffey CL, Sun W, Yang Y, Chao HT, Frankel WN. Etiology of a genetically complex seizure disorder in Celf4 mutant mice. Genes Brain Behav. 2011;10:765–77.
Sun W, Wagnon JL, Mahaffey CL, Briese M, Ule J, Frankel WN. Aberrant sodium channel activity in the complex seizure disorder of Celf4 mutant mice. J Physiol. 2013;591:241–55.
Yang Y, Mahaffey CL, Bérubé N, Maddatu TP, Cox GA, Frankel WN. Complex seizure disorder caused by Brunol4 deficiency in mice. PLoS Genet. 2007;3:e124.
Shen Y, Zhang C, Xiao K, Liu D, Xie G. CELF4 regulates spine formation and depression-like behaviors of mice. Biochem Biophys Res Commun. 2022;605:39–44.
Peng S, Cai X, Chen J, Sun J, Lai B, Chang M, et al. The role of CELF family in neurodevelopment and neurodevelopmental disorders. Neurobiol Dis. 2024;197:106525.
Barone R, Fichera M, De Grandi M, Battaglia M, Lo Faro V, Mattina T, et al. Familial 18q12.2 deletion supports the role of RNA-binding protein CELF4 in autism spectrum disorders. Am J Med Genet A. 2017;173:1649–55.
Chen CP, Hsieh CH, Chern SR, Wu PS, Chen SW, Lai ST, et al. Prenatal diagnosis and molecular cytogenetic characterization of an interstitial deletion of 18q12.1-q12.3 encompassing DTNA, CELF4 and SETBP1. Taiwan J Obstet Gynecol. 2017;56:847–51.
Chen CP, Huang MC, Chen YY, Chern SR, Wu PS, Chen YT, et al. Prenatal diagnosis of de novo interstitial deletions involving 5q23.1-q23.3 and 18q12.1-q12.3 by array CGH using uncultured amniocytes in a pregnancy with fetal interrupted aortic arch and atrial septal defect. Gene. 2013;531:496–501.
Halgren C, Bache I, Bak M, Myatt MW, Anderson CM, Brøndum-Nielsen K, et al. Haploinsufficiency of CELF4 at 18q12.2 is associated with developmental and behavioral disorders, seizures, eye manifestations, and obesity. Eur J Hum Genet. 2012;20:1315–9.
Gilling M, Lauritsen MB, Møller M, Henriksen KF, Vicente A, Oliveira G, et al. A 3.2 Mb deletion on 18q12 in a patient with childhood autism and high-grade myopia. Eur J Hum Genet. 2008;16:312–9.
Acknowledgements
We are grateful to the patients and their families for their participation in this study. We thank the Centre de Calcul de l’Univeristé de Bourgogne (CCuB) for the technical support and management of the informatics platform. This study makes use of data generated by the French SequoiA platform (https://laboratoire-seqoia.fr/); and DECIPHER, ClinVAR and denovo-db community. A full list of centers who contributed to the generation of the data are available at https://deciphergenomics.org/about/stats, https://www.ncbi.nlm.nih.gov/clinvar/ and https://denovo-db.gs.washington.edu/denovo-db/. We thank Mohnish Suri, Jennifer Howe and Stephanie Einsele-Scholz for the data sharing. Patients cited in this study were gathered using GeneMatcher platform (https://genematcher.org/statistics/).
Funding
This work was supported by grants from the Dijon University Hospital, the Burgundy-Franche Comté regional council (Plan d’Actions Régional pour l’Innovation PARI and the European Union through the FEDER programs 2014/2020). The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript. This study is funded by Physicians’ Services Incorporated (PSI) Foundation Health Research Grant to Dr. Saadet Mercimek-Andrews. We would like to thank the PSI Foundation for their funding support (Patient 15).
Author information
Authors and Affiliations
Contributions
A-LB, CT-R and AV contributed to the writing of the manuscript. A-LB, CP, YD, FL and FB analyzed and/or interpreted exome data. ATV, GLG, BG-D, XLG, SR, MR, KNL, AB, MS, FG, LF, CT-R, VR, AG, FL, NB, RA, MI, X-RY, ER-O, MP-B, JCH, JC, AJ, RDK, SM-A, AA, ENW, IMW, ET, CG contributed to the phenotyping of patients. All authors discussed the results and contributed to the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests. ENW and IMW are employees of GeneDx, LLC.
Ethics approval and consent to participate
Parents signed consent forms for genetic analyses, participation to this study and for images and clinical data publication. Patient 15: this study is approved by the Institutional Research Ethics Boards (REB#100032244) and (ARISE# Pro00106741).
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Bruel, AL., Vulto-vanSilfhout, A.T., Bilan, F. et al. Heterozygous CELF4 variants in the N-term region crucial for the RNA-binding activity lead to neurodevelopmental disorder and obesity. Eur J Hum Genet 33, 852–859 (2025). https://doi.org/10.1038/s41431-025-01809-w
Received:
Revised:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41431-025-01809-w
This article is cited by
-
Summer reading 2025 in EJHG
European Journal of Human Genetics (2025)