Abstract
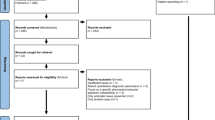
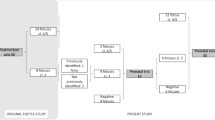
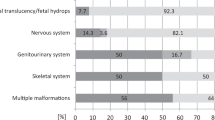
In 30–40% of fetuses with structural defects, the causal variant remains undiagnosed after karyotype, chromosomal microarray, and exome sequencing. This study presents the results of a reanalysis of unsolved prenatal ES (pES) cases and investigates how postnatal/postmortem phenotyping contributes to identifying relevant variants. pES data was prospectively reanalyzed for unsolved cases enrolled in the AnDDI-Prénatome cohort study. Postnatal/postmortem data were included with prenatal features using Human Phenotype Ontology terms up to 3 years after pES. The reanalysis involved updating bioinformatic processing and querying raw data using a GREP query. We reanalyzed 58/94 (62%) unsolved pES cases, including 8 variants of unknown significance. Data for clinical examination at birth was available for all live newborns, and postmortem examination was available in 12 terminated fetuses. Additional features were identified at birth in 27/58 cases (44%): 9 terminated fetuses, 2 stillbirths, and 16 live newborns. One diagnosis (SNAPC4) was obtained through a periodic query following recent associations with human disease, and without additional clinical data. Three additional VUS were identified through reanalysis with the addition of new clinical features, illustrating the limited contribution of updated postnatal/postmortem phenotyping in identifying relevant variants after negative pES. In conclusion, the benefit of prospective reanalysis of unsolved pES is limited, even over time. Postnatal genome sequencing may be a more appropriate option than reanalysis with postnatal/postmortem phenotyping to establish a causal diagnosis.
This is a preview of subscription content, access via your institution
Access options

Similar content being viewed by others
Data availability
The data that support the findings of this study are available from the corresponding author on reasonable request.
References
Liu L, Oza S, Hogan D, Perin J, Rudan I, Lawn JE, et al. Global, regional, and national causes of child mortality in 2000-13, with projections to inform post-2015 priorities: an updated systematic analysis. Lancet 2015;385:430–40.
Mellis R, Oprych K, Scotchman E, Hill M, Chitty LS. Diagnostic yield of exome sequencing for prenatal diagnosis of fetal structural anomalies: A systematic review and meta-analysis. Prenat Diagn. 2022;42:662–85.
Normand EA, Braxton A, Nassef S, Ward PA, Vetrini F, He W, et al. Clinical exome sequencing for fetuses with ultrasound abnormalities and a suspected Mendelian disorder. Genome Med. 2018;10:74.
Bruel AL, Vitobello A, Tran Mau-Them F, Nambot S, Sorlin A, Denommé-Pichon AS, et al. Next-generation sequencing approaches and challenges in the diagnosis of developmental anomalies and intellectual disability. Clin Genet 2020;98:433–44.
Bruel AL, Nambot S, Quéré V, Vitobello A, Thevenon J, Assoum M, et al. Increased diagnostic and new genes identification outcome using research reanalysis of singleton exome sequencing. Eur J Hum Genet. 2019;27:1519–31.
Nambot S, Thevenon J, Kuentz P, Duffourd Y, Tisserant E, Bruel AL, et al. Clinical whole-exome sequencing for the diagnosis of rare disorders with congenital anomalies and/or intellectual disability: substantial interest of prospective annual reanalysis. Genetics Med. 2018;20:645–54.
Sarmady M, Abou Tayoun A. Need for Automated Interactive Genomic Interpretation and Ongoing Reanalysis. JAMA Pediatr. 2018;172:1113–4.
Dai P, Honda A, Ewans L, McGaughran J, Burnett L, Law M, et al. Recommendations for next generation sequencing data reanalysis of unsolved cases with suspected Mendelian disorders: A systematic review and meta-analysis. Genet Med. 2022;24:1618–29.
Tran Mau-Them F, Delanne J, Denommé-Pichon AS, Safraou H, Bruel AL, Vitobello A, et al. Prenatal diagnosis by trio exome sequencing in fetuses with ultrasound anomalies: A powerful diagnostic tool. Front Genet. 2023;14:1099995.
Lefebvre M, Bruel AL, Tisserant E, Bourgon N, Duffourd Y, Collardeau-Frachon S, et al. Genotype-first in a cohort of 95 fetuses with multiple congenital abnormalities: when exome sequencing reveals unexpected fetal phenotype-genotype correlations. J Med Genet. 2021;58:400–13.
Tran Mau-Them F, Overs A, Bruel AL, Duquet R, Thareau M, Denommé-Pichon AS, et al. Combining globally search for a regular expression and print matching lines with bibliographic monitoring of genomic database improves diagnosis. Frontiers in Genetics [Internet]. 2023 [cité 13 janv 2024];14. Disponible sur: https://www.frontiersin.org/articles/10.3389/fgene.2023.1122985
Frost FG, Morimoto M, Sharma P, Ruaud L, Belnap N, Calame DG, et al. Bi-allelic SNAPC4 variants dysregulate global alternative splicing and lead to neuroregression and progressive spastic paraparesis. Am J Hum Genet. 2023;110:663–80.
Palencia-Campos A, Ullah A, Nevado J, Yildirim R, Unal E, Ciorraga M, et al. GLI1 inactivation is associated with developmental phenotypes overlapping with Ellis-van Creveld syndrome. Hum Mol Genet. 2017;26:4556–71.
Palencia-Campos A, Martínez-Fernández ML, Altunoglu U, Soto-Bielicka P, Torres A, Marín P, et al. Heterozygous pathogenic variants in GLI1 are a common finding in isolated postaxial polydactyly A/B. Hum Mutat. 2020;41:265–76.
Guimier A, Gabriel GC, Bajolle F, Tsang M, Liu H, Noll A, et al. MMP21 is mutated in human heterotaxy and is required for normal left-right asymmetry in vertebrates. Nat Genet. 2015;47:1260–3.
Rossi AC, Prefumo F. Correlation between fetal autopsy and prenatal diagnosis by ultrasound: A systematic review. Eur J Obstet Gynecol Reprod Biol. 2017;210:201–6.
Vimercati A, Grasso S, Abruzzese M, Chincoli A, de Gennaro A, Miccolis A, et al. Correlation between ultrasound diagnosis and autopsy findings of fetal malformations. J Prenat Med. 2012;6:13–7.
Godbole K, Bhide V, Nerune S, Kulkarni A, Moghe M, Kanade A. Role of fetal autopsy as a complementary tool to prenatal ultrasound. J Matern Fetal Neonatal Med. 2014;27:1688–92.
Bourgon N, Garde A, Bruel AL, Lefebvre M, Mau-Them FT, Moutton S, et al. Same performance of exome sequencing before and after fetal autopsy for congenital abnormalities: toward a paradigm shift in prenatal diagnosis? Eur J Hum Genet. 2022;16:1–9.
Schubert C, Milverton J, Goodall S, Merlin T. A systematic review to assess the utility of genomic autopsy using exome or genome sequencing in cases of congenital anomalies and perinatal death. Genet Med. 2024;26:101159.
Westphal DS, Leszinski GS, Rieger-Fackeldey E, Graf E, Weirich G, Meitinger T, et al. Lessons from exome sequencing in prenatally diagnosed heart defects: A basis for prenatal testing. Clin Genet. 2019;95:582–9.
Aarabi M, Sniezek O, Jiang H, Saller DN, Bellissimo D, Yatsenko SA, et al. Importance of complete phenotyping in prenatal whole exome sequencing. Hum Genet. 2018;137:175–81.
Aggarwal S, Vineeth VS, Das Bhowmik A, Tandon A, Kulkarni A, Narayanan DL, et al. Exome sequencing for perinatal phenotypes: The significance of deep phenotyping. Prenat Diagn 2020;40:260–73.
Rinaldi B, Race V, Corveleyn A, Van Hoof E, Bauters M, Van Den Bogaert K, et al. Next-generation sequencing in prenatal setting: Some examples of unexpected variant association. Eur J Med Genet. 2020;63:103875.
Vora NL, Norton ME. Prenatal exome and genome sequencing for fetal structural abnormalities. Am J Obstet Gynecol. 2023;228:140–9.
Mone F, Abu Subieh H, Doyle S, Hamilton S, Mcmullan DJ, Allen S, et al. Evolving fetal phenotypes and clinical impact of progressive prenatal exome sequencing pathways: cohort study. Ultrasound Obstet Gynecol. 2022;59:723–30.
Acknowledgements
We are grateful to the patients and their families for their participation in this study. We thank the Centre de Calcul de l’Univeristé de Bourgogne (CCuB) for technical support and management of the informatics platform, and the GeneMatcher platform for data sharing. Several authors of this publication are members of the European Reference Network for Developmental Anomalies and Intellectual Disability (ERN-ITHACA). We thank Suzanne Rankin from the Dijon-Bourgogne University Hospital for editing the English language of the manuscript.
Funding
This work was supported by grants from the Dijon University Hospital, the Burgundy-Franche Comté regional council, the French Ministry of Health (national AnDDI-rares network), and the European Union through the FEDER programs (PERSONALISE). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript. This study was funded by the national network AnDDI-rares (AnDDI-Prénatome) (NCT03964441)
Author information
Authors and Affiliations
Contributions
Conception: CTR, CB, NB. Data acquisition: AG, MF, JD, CR, TR, SN, SM, CQ, CC, A-CB, A-MG, CR, AP, PB, SO, ES, OB, AG, AG, CA, GM, MF, BI, MV, CF, GV, FP, MN, CW, MJ, CD, AZ, MG, PSV, KC, PB, ES, LF. Data interpretation: YD, HS, A-SD-P, AV, CP, MF, ALB, FTMT, NB. Drafted or revised the manuscript: NB, CT. Approved the final version: NB
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical approval
All fetuses were initially included in this AnDDI-Prénatome study (NCT03964441), approved by institutional review board and ethics committee. Informed written consent was obtained from all subjects and participating family members.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Thauvin-Robinet, C., Garde, A., Favier, M. et al. Reanalysis of unsolved prenatal exome sequencing for structural defects: diagnostic yield and contribution of postnatal/postmortem features. Eur J Hum Genet 33, 675–682 (2025). https://doi.org/10.1038/s41431-025-01823-y
Received:
Revised:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41431-025-01823-y
This article is cited by
-
Editorial Déjà Vu: This time, it’s actually ‘what’s new in EJHG in May 2025?’
European Journal of Human Genetics (2025)