Abstract
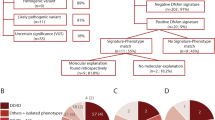
DNA methylation (DNAm) episignature analysis is an emerging tool for diagnosing individuals with neurodevelopmental disorders, congenital anomalies, and growth disorders. We evaluated its clinical utility as a first-tier test in 62 individuals without prior molecular testing. DNAm arrays identified a diagnosis in 30.6% (19/62) of cases. The positivity rate was highest for Fragile X syndrome (100%, 5/5), followed by syndromic disorders (44%, 8/18) and imprinting disorders (25%, 6/24), including Silver-Russell, Beckwith-Wiedemann, and Prader-Willi syndromes. No diagnoses were made in 15 individuals with non-syndromic neurodevelopmental disorders. Alternative diagnoses were identified in 4.8% (3/62) of cases. These findings suggest that DNAm arrays can serve as an effective first-tier diagnostic tool, particularly for syndromic and imprinting disorders, with potential to improve diagnostic efficiency and reduce reliance on sequential genetic testing. While these findings support the use of DNAm arrays as an effective first-tier tool in selected populations, larger, unselected cohort studies are needed to validate its generalizability.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
Data availability
Data that support the findings of this study are available on request from the corresponding author with reasonable patient privacy restrictions.
References
Moeschler JB, Shevell M. Committee on Genetics. Comprehensive evaluation of the child with intellectual disability or global developmental delays. Pediatrics. 2014;134:e903–18.
Levy MA, McConkey H, Kerkhof J, Barat-Houari M, Bargiacchi S, Biamino E, et al. Novel diagnostic DNA methylation episignatures expand and refine the epigenetic landscapes of Mendelian disorders. HGG Adv. 2021;3:100075.
Aref-Eshghi E, Kerkhof J, Pedro VP, France GD, Barat-Houari M, Ruiz-Pallares N, et al. Evaluation of DNA methylation episignatures for diagnosis and phenotype correlations in 42 Mendelian neurodevelopmental disorders. Am J Hum Genet. 2021;108:1161–3.
Sadikovic B, Levy MA, Kerkhof J, Aref-Eshghi E, Schenkel L, Stuart A, et al. Clinical epigenomics: genome-wide DNA methylation analysis for the diagnosis of Mendelian disorders. Genet Med. 2021;23:1065–74.
Aref-Eshghi E, Rodenhiser DI, Schenkel LC, Lin H, Skinner C, Ainsworth P, et al. Genomic DNA methylation signatures enable concurrent diagnosis and clinical genetic variant classification in neurodevelopmental syndromes. Am J Hum Genet. 2018;102:156–74.
Sadikovic B, Levy MA, Aref-Eshghi E. Functional annotation of genomic variation: DNA methylation episignatures in neurodevelopmental Mendelian disorders. Hum Mol Genet. 2020;29:R27–R32.
Schenkel LC, Schwartz C, Skinner C, Rodenhiser DI, Ainsworth PJ, Pare G, et al. Clinical validation of fragile X syndrome screening by DNA methylation array. J Mol Diagn. 2016;18:834–41.
Aref-Eshghi E, Kerkhof J, Pedro VP, Groupe DI France, Barat-Houari M, Ruiz-Pallares N, et al. Evaluation of DNA methylation episignatures for diagnosis and phenotype correlations in 42 Mendelian neurodevelopmental disorders. Am J Hum Genet. 2020;106:356–70.
EpiSign. EpiSign conditions list. London (ON): EpiSign; 2024. https://episign.com/wp-content/uploads/2024/06/EpiSign-Conditions-List.pdf
Kerkhof J, Squeo GM, McConkey H, Levy MA, Piemontese MR, Castori M, et al. DNA methylation episignature testing improves molecular diagnosis of Mendelian chromatinopathies. Genet Med. 2022;24:51–60.
Fontana L, Bedeschi MF, Maitz S, Cereda A, Faré C, Motta S, et al. Characterization of multi-locus imprinting disturbances and underlying genetic defects in patients with chromosome 11p15.5 related imprinting disorders. Epigenetics. 2018;13:897–909.
Krzyzewska IM, Alders M, Maas SM, Bliek J, Venema A, Henneman P, et al. Genome-wide methylation profiling of Beckwith-Wiedemann syndrome patients without molecular confirmation after routine diagnostics. Clin Epigenetics. 2019;11:53.
Kerkhof J, Rastin C, Levy MA, Relator R, McConkey H, Demain L, et al. Diagnostic utility and reporting recommendations for clinical DNA methylation episignature testing in genetically undiagnosed rare diseases. Genet Med. 2024;26:101075.
Acknowledgements
We thank all the individuals and their families for their participation in the study. Tinatin Tkemaladze also wishes to express her gratitude to Sofia Douzgou Houge for facilitating the connection with Siddhant Banka, the lead author of the study.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: NIHR Manchester Biomedical Research Centre (NIHR203308) and the MRC Epigenomics of Rare Diseases Node (MR/Y008170/1) grants.
Author information
Authors and Affiliations
Contributions
Conceptualization: T.T., S.B.; Data Curation: T.T., C.C., S.B.; Formal Analysis: T.T., K.B., C.C.; Investigation: T.T., C.C., L.D., S.B.; Methodology: B.S., S.B.; Project Administration: T.T., S.J., S.H., S.B.; Resources: T.T., S.B.; Software: C.C., B.S., M.L., J.K., D.G.; Writing-original draft: T.T., C.C., E.K., K.B., E.A., S.B.; Writing-review & editing: T.T., C.C., K.B., E.K., E.A., L.D., S.J., S.H., B.S., M.L., J.K., D.G., S.B.
Corresponding author
Ethics declarations
Competing interests
B.S. is a shareholder in EpiSign Inc.
Ethical approval
The study was approved by local Institutional Review Board. Written and oral consent from all individuals’ legal representatives was obtained. Consent was obtained to publish identifying photos.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Tkemladze, T., Campbell, C., Bregvadze, K. et al. Evaluating DNA methylation episignatures as a first-tier diagnostic test in individuals with suspected genetic disorders. Eur J Hum Genet 34, 296–299 (2026). https://doi.org/10.1038/s41431-025-01939-1
Received:
Revised:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41431-025-01939-1
This article is cited by
-
To sign or not to sign: Is this still the question?
European Journal of Human Genetics (2025)