Abstract
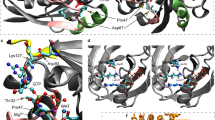
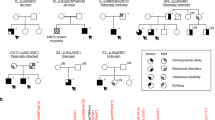
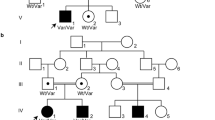
We recently identified de novo missense variants affecting the small GTPase ARF3 as the cause of a disorder characterized by developmental delay/intellectual disability, microcephaly, brain atrophy, epilepsy and minor skeletal defects. In vitro and in vivo analyses documented impaired Golgi integrity, vesicle trafficking, and brain and body axes development. Here, we report clinical features of five additional patients and the functional characterization of three novel ARF3 variants. Cell-based assays corroborate a deleterious, variant-specific effect on protein stability, GTP binding and Golgi morphology. Zebrafish models confirm the dominant behavior of the tested variants and their variable impact on development. ARF3 mutants significantly affected Golgi integrity in vivo as well as brain size, recapitulating patients’ microcephaly. These findings expand the ARF3-related Golgipathy mutational spectrum, strengthen previous observations linking variants with dominant negative behavior to a markedly severe phenotype, and underscore the specific vulnerability of the nervous system to ARF and Golgi dysfunction.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
The functional data generated in this study can be found in the supplementary material. The sequencing data are not publicly available due to due to privacy/ethical restrictions. The new variants identified in this work and their clinical association have been submitted to ClinVar (SCV006550956-SCV006550959).
References
Wilson C, Venditti R, Rega LR, Colanzi A, D’Angelo G, De Matteis MA. The Golgi apparatus: an organelle with multiple complex functions. Biochem J. 2011;433:1–9.
Wei JH, Seemann J. Mitotic division of the mammalian Golgi apparatus. Semin Cell Dev Biol. 2009;20:810–6.
Cohen LA, Donaldson JG. Analysis of Arf GTP-binding protein function in cells. Curr Protoc Cell Biol. 2010;Chapter 3:Unit 14.12.1–17.
Li T, Guo Y. ADP-Ribosylation Factor Family of Small GTP-Binding Proteins: Their Membrane Recruitment, Activation, Crosstalk and Functions. Front Cell Dev Biol. 2022;10:813353.
Ge X, Gong H, Dumas K, Litwin J, Phillips JJ, Waisfisz Q, et al. Missense-depleted regions in population exomes implicate ras superfamily nucleotide-binding protein alteration in patients with brain malformation. NPJ Genom Med. 2016;1:16036.
Fasano G, Muto V, Radio FC, Venditti M, Mosaddeghzadeh N, Coppola S, et al. Dominant ARF3 variants disrupt Golgi integrity and cause a neurodevelopmental disorder recapitulated in zebrafish. Nat Commun. 2022;13:6841.
Sakamoto M, Sasaki K, Sugie A, Nitta Y, Kimura T, Gürsoy S, et al. De novo ARF3 variants cause neurodevelopmental disorder with brain abnormality. Hum Mol Genet. 2021;31:69–81.
Rasika S, Passemard S, Verloes A, Gressens P, El Ghouzzi V. Golgipathies in Neurodevelopment: A New View of Old Defects. Dev Neurosci. 2018;40:396–416.
Sobreira N, Schiettecatte F, Valle D, Hamosh A. GeneMatcher: a matching tool for connecting investigators with an interest in the same gene. Hum Mutat. 2015;36:928–30.
Richards S, Aziz N, Bale S, Bick D, Das S, Gastier-Foster J, et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med. 2015;17:405–24.
Ratcliffe CDH, Sahgal P, Parachoniak CA, Ivaska J, Park M. Regulation of Cell Migration and β1 Integrin Trafficking by the Endosomal Adaptor GGA3. Traffic. 2016;17:670–88.
Manolea F, Chun J, Chen DW, Clarke I, Summerfeldt N, Dacks JB, et al. Arf3 is activated uniquely at the trans-Golgi network by brefeldin A-inhibited guanine nucleotide exchange factors. Mol Biol Cell. 2010;21:1836–49.
Volpicelli-Daley LA, Li Y, Zhang CJ, Kahn RA. Isoform-selective Effects of the Depletion of ADP-Ribosylation Factors 1–5 on Membrane Traffic. Mol Biol Cell. 2005;16:4495–508.
Zhang CJ, Rosenwald AG, Willingham MC, Skuntz S, Clark J, Kahn RA. Expression of a dominant allele of human ARF1 inhibits membrane traffic in vivo. J Cell Biol. 1994;124:289–300.
Peri F, Nüsslein-Volhard C. Live Imaging of Neuronal Degradation by Microglia Reveals a Role for v0-ATPase a1 in Phagosomal Fusion In Vivo. Cell. 2008;133:916–27.
Dos Santos Henrique S, França MJ, Silva Junior RC, Santos MLSF, do Valle DA. Neurodevelopmental disorder associated with gene ARF3: A case report. Am J Med Genet A. 2024;194:e63658.
Donaldson JG, Jackson CL. ARF family G proteins and their regulators: roles in membrane transport, development and disease. Nat Rev Mol Cell Biol. 2011;12:362–75.
Sheen VL, Ganesh VS, Topcu M, Sebire G, Bodell A, Hill RS, et al. Mutations in ARFGEF2 implicate vesicle trafficking in neural progenitor proliferation and migration in the human cerebral cortex. Nat Genet. 2004;36:69–76.
Macken WL, Godwin A, Wheway G, Stals K, Nazlamova L, Ellard S, et al. Biallelic variants in COPB1 cause a novel, severe intellectual disability syndrome with cataracts and variable microcephaly. Genome Med. 2021;25:34.13.
Brault JB, Bardin S, Lampic M, Carpentieri JA, Coquand L, Penisson M, et al. RAB6 and dynein drive post-Golgi apical transport to prevent neuronal progenitor delamination. EMBO Rep. 2022;23:e54605.
Acknowledgements
We thank the families participating in the study. One patient included in this study was diagnosed thanks to Defidiag (NCT04154891), a project supported by The French Ministry of Health in the framework of the French initiative for genomic medicine (https://pfmg2025.fr/) and by French government funding from the Agence Nationale de la Recherche under the “Investissements d’avenir” program (ANR- 10-IAHU-01).
Funding
The work was supported by “Fondazione Telethon ETS” (grant number GMR23T1168, to AL), Italian Ministry of Health (5x1000, to AL; Current Research Funds, PNRR-MR1-2022-12376811 and RF-2021-12374963, to MT) and Istituto Superiore di Sanità (Bando Ricerca Indipendente ISS20-2e15b898baf0, to SC).
Author information
Authors and Affiliations
Contributions
VM, SC, EZ: functional analyses (in vitro); GF, CP: functional analyses (in vivo); FCR: clinical and molecular data assessment and data curation; MC: structural analyses; CS-B, CB, SEC, BG, AdSM, DK, ES, CC, TH, A-SH, LvE, EEB, UH, AS: clinical and molecular data collection; MT, AL: study coordination, funding acquisition, writing.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical approval
All subjects were referred for diagnostic genetic testing. Clinical data and DNA samples were collected, stored, and used in accordance with the ethical standards of the Declaration of Helsinki, following written informed consent from participating families. The study was approved by the local Institutional Ethical Committee of the Ospedale Pediatrico Bambino Gesù, Rome (ref. RF-2021-12374963 and PNRR-MR1-2022-12376811). Written informed consents were obtained for publication of individual pictures and clinical details. All animal experiments were conducted according to ARRIVE guidelines (https://arriveguidelines.org/) upon approval of the Italian Ministry of Health (Direzione Generale della Salute Animale - Ufficio 6 EX DGSAF, ref.n° 652/2024-PR).
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Muto, V., Fasano, G., Radio, F.C. et al. Newly identified ARF3 variants strengthen the causal link between Golgi fragmentation and brain malformations. Eur J Hum Genet 34, 438–443 (2026). https://doi.org/10.1038/s41431-025-02001-w
Received:
Revised:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41431-025-02001-w