Abstract
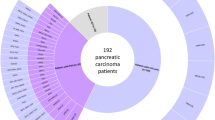
Pancreatic ductal adenocarcinoma occur in the context of a suspected or proven genetic predisposition in 5–10% of cases. While universal germline multigene panel testing is currently recommended by NCCN and ASCO, this approach was previously limited to patients with personal and/or family criteria suggestive of hereditary predisposition. We report the results of this « selective » approach applied in our institution from January 2018 to June 2023. Germline testing of a panel of 13 « clinically actionable » genes (APC, ATM, BRCA1, BRCA2, CDKN2A, MLH1, MSH2, MSH6, PALB2, PMS2, RAD51C, RAD51D, STK11) was performed in 496 patients with pancreatic ductal adenocarcinoma and suspected genetic predisposition based on the validation of prespecified clinical criteria. A germline pathogenic/likely pathogenic variant of one of these genes was identified in 49 patients corresponding to 9.9% of the study population. ATM and BRCA2 were the two most frequently implicated genes (18 and 16 cases, respectively) and the prevalence of pathogenic/likely pathogenic variants of these genes was significantly higher than in gnomAD controls. The overall contribution of core and non-core genes of the Homologous Recombination DNA repair system was 83.7% while the contribution of the Mismatch Repair system was 10.2%. An exploratory approach consisting of unmasking the results of the NGS analysis of 123 « research » genes involved in the carcinogenesis was applied to the 447 patients tested negative for the different genes of our diagnostic panel. This approach failed to identify other susceptibility genes to pancreatic adenocarcinoma.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
Data availability
The datasets generated and/or analysed during the current study are available from the corresponding author on reasonable request.
References
Goggins M, Overbeek KA, Brand R, Syngal S, Del Chiaro M, Bartsch DK, et al. Management of patients with increased risk for familial pancreatic cancer: updated recommendations from the international Cancer of the Pancreas Screening (CAPS) consortium. Gut. 2020;69:7–17.
Leman R, Parfait B, Vidaud D, Girodon E, Pacot L, Le Gac G, et al. SPiP: splicing Prediction Pipeline, a machine learning tool for massive detection of exonic and intronic variant effects on mRNA splicing. Hum Mutat. 2022;43:2308–23.
Jaganathan K, Panagiotopoulou SK, McRae JF, Darbandi SF, Knowles D, Li YI, et al. Predicting splicing from primary sequence with deep learning. Cell. 2019;176:535–48.
Richards S, Aziz N, Bale S, Bick D, Das S, Gastier-Foster J, et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med. 2015;17:405–24.
Conroy T, Pfeiffer P, Vilgrain V, Lamarca A, Seufferlein T, O’Reilly EM, et al. ESMO Guidelines Committee. Pancreatic cancer: ESMO Clinical Practice Guideline for diagnosis, treatment and follow-up. Ann Oncol. 2023;34:987–1002.
Daly MB, Pal T, Maxwell KN, Churpek J, Kohlmann W, AlHilli Z, et al. NCCN guidelines insights: genetic/familial high-risk assessment: breast, ovarian, and pancreatic, version 2.2024. J Natl Compr Canc Netw. 2023;21:1000–10.
Frebourg T, Bajalica Lagercrantz S, Oliveira C, Magenheim R, Evans DG. European Reference Network GENTURIS. Guidelines for the Li-Fraumeni and heritable TP53-related cancer syndromes. Eur J Hum Genet. 2020;28:1379–86.
Uson PLS Jr, Samadder NJ, Riegert-Johnson D, Boardman L, Borad MJ, Ahn D, et al. Clinical impact of pathogenic germline variants in pancreatic cancer: results from a multicenter, prospective, universal genetic testing study. Clin Transl Gastroenterol. 2021;12:e00414.
Hu C, Hart SN, Polley EC, Gnanaolivu R, Shimelis H, Lee KY, et al. Association between inherited germline mutations in cancer predisposition genes and risk of pancreatic cancer. JAMA. 2018;319:2401–9.
Gardiner A, Kidd J, Elias MC, Young K, Mabey B, Taherian N, et al. Pancreatic ductal carcinoma risk associated with hereditary cancer-risk genes. J Natl Cancer Inst. 2022;114:996–1002.
LaDuca H, Polley EC, Yussuf A, Hoang L, Gutierrez S, Hart SN, et al. A clinical guide to hereditary cancer panel testing: evaluation of gene-specific cancer associations and sensitivity of genetic testing criteria in a cohort of 165,000 high-risk patients. Genet Med. 2020;22:407–15.
Kindler HL, Hammel P, Reni M, Van Cutsem E, Macarulla T, Hall MJ, et al. Overall survival results from the POLO trial: a phase III study of active maintenance olaparib versus placebo for germline BRCA-mutated metastatic pancreatic cancer. J Clin Oncol. 2022;40:3929–39.
Park W, O’Connor CA, Bandlamudi C, Forman D, Chou JF, Umeda S, et al. Clinico-genomic characterization of ATM and HRD in pancreas cancer: application for practice. Clin Cancer Res. 2022;28:4782–92.
Golan T, O’Kane GM, Denroche RE, Raitses-Gurevich M, Grant RC, Holter S, et al. Genomaic features and classification of homologous recombination deficient pancreatic ductal adenocarcinoma. Gastroenterology. 2021;160: 2119–32.
Lesueur F, Easton DF, Renault AL, Tavtigian SV, Bernstein JL, Kote-Jarai Z, et al. First international workshop of the ATM and cancer risk group (4-5 December 2019). Fam Cancer. 2022;21:211–27.
Sargen MR, Helgadottir H, Yang XR, Harland M, Hatton JN, Jones K, et al. Impact of transcript (p16/p14ARF) alteration on cancer risk in CDKN2A germline pathogenic variant carriers. JNCI Cancer Spectr. 2022;6(6):pkac074.
Tateo V, Marchese PV, Mollica V, Massari F, Kurzrock R, Adashek JJ. Agnostic approvals in oncology: getting the right drug to the right patient with the right genomics. Pharm (Basel). 2023;16:614.
Looney AM, Nawaz K, Webster RM. Tumor-agnostic therapies. Nat Rev Drug Discov. 2020;19:383–4.
Marchetti P, Botticelli A, Ascierto AP, Curigliano G, Giannarelli D. Agnostic evaluation of ipilimumab and nivolumab association: a metanalysis. J Transl Med. 2020;18:446.
Fortuno C, Lee K, Olivier M, Pesaran T, Mai PL, de, et al. ClinGen TP53 variant curation expert panel. Specifications of the ACMG/AMP variant interpretation guidelines for germline TP53 variants. Hum Mutat. 2021;42:223–36.
Kratz CP, Freycon C, Maxwell KN, Nichols KE, Schiffman JD, Evans DG, et al. Analysis of the Li-fraumeni spectrum based on an international germline TP53 variant data set: an international agency for research on cancer TP53 database analysis. JAMA Oncol. 2021;7:1800–5.
Chittenden A, Haraldsdottir S, Ukaegbu C, Underhill-Blazey M, Gaonkar S, Uno H, et al. Implementing systematic genetic counseling and multigene germline testing for individuals with pancreatic cancer. JCO Oncol Pr. 2021;17:e236–e247.
Esplin ED, Nielsen SM, Bristow SL, Garber JE, Hampel H, Rana HQ, et al. Universal germline genetic testing for hereditary cancer syndromes in patients with solid tumor cancer. JCO Precis Oncol. 2022;6:e2100516.
Mandelker D, Zhang L, Kemel Y, Stadler ZK, Joseph V, Zehir A, et al. Mutation detection in patients with advanced cancer by universal sequencing of cancer-related genes in tumor and normal DNA vs guideline-based germline testing. JAMA. 2017;318:825–35.
Samadder NJ, Riegert-Johnson D, Boardman L, Rhodes D, Wick M, Okuno S, et al. Comparison of universal genetic testing vs guideline-directed targeted testing for patients with hereditary cancer syndrome. JAMA Oncol. 2021;7:230–7.
Jones JC, Golafshar MA, Coston TW, Rao R, Wysokinska E, Johnson E, et al. Universal genetic testing vs. guideline-directed testing for hereditary cancer syndromes among traditionally underrepresented patients in a community oncology program. Cureus. 2023;15:e37428.
Blanco Abad C, Gomila Pons P, Campos Ramírez S, Álvarez Alejandro M, Torres Ramón MI, et al. Hereditary pancreatic cancer: advances in genetic testing, early detection strategies, and personalized management. J Clin Med. 2025;14:367.
Subbiah V, Kurzrock R. Universal germline and tumor genomic testing needed to win the war against cancer: genomics is the diagnosis. J Clin Oncol. 2023;41:3100–3.
Lowery MA, Wong W, Jordan EJ, Lee JW, Kemel Y, Vijai J, et al. Prospective evaluation of germline alterations in patients with exocrine pancreatic neoplasms. J Natl Cancer Inst. 2018;110:1067–74.
Llach J, Luzko I, Earl J, Barreto E, Rodríguez-Garrote M, Lleixà M, et al. Should we offer universal germline genetic testing to all patients with pancreatic cancer? a multicenter study. Cancers (Basel). 2024;16:3779.
Tempero MA. NCCN guidelines updates: pancreatic cancer. J Natl Compr Canc Netw. 2019;17:603–5.
McDonald HG, Kennedy A, Solomon AL, Williams CM, Reagan AM, Cassim E, et al. Development of a novel protocol for germline testing in pancreatic cancer. Ann Surg Oncol. 2024;31:7705–12.
Sanoba S, Wadlow R. Universal genetic testing for pancreatic cancer : closing the gap on a missed opportunity. JCO Oncol Pract. 2025;OP2500154 https://doi.org/10.1200/OP-25-00154.
Slavin TP, Banks KC, Chudova D, Oxnard GR, Odegaard JI, Nagy RJ, et al. Identification of incidental germline mutations in patients with advanced solid tumors who underwent cell-free circulating tumor DNA sequencing. J Clin Oncol. 2018;36:3459–65.
Kuzbari Z, Bandlamudi C, Loveday C, Garrett A, Mehine M, George A, et al. Germline-focused analysis of tumour-detected variants in 49,264 cancer patients: ESMO precision medicine working group recommendations. Ann Oncol. 2023;34:215–27.
Hoffman TL, Kershberg H, Goff J, Holmquist KJ, Haque R, Alvarado M. Next-generation universal hereditary cancer screening: implementation of an automated hereditary cancer screening program for patients with advanced cancer undergoing tumor sequencing in a large HMO. Fam Cancer. 2023;22:225–35.
Gentiluomo M, Canzian F, Nicolini A, Gemignani F, Landi S, Campa D. Germline genetic variability in pancreatic cancer risk and prognosis. Semin Cancer Biol. 2022;79:105–31.
Klein AP, Wolpin BM, Risch HA, Stolzenberg-Solomon RZ, Mocci E, Zhang M, et al. Genome-wide meta-analysis identifies five new susceptibility loci for pancreatic cancer. Nat Commun. 2018;9:556.
Funding
This study did not benefit from financial support.
Author information
Authors and Affiliations
Contributions
All authors were involved in drafting the manuscript, have read and approved the final paper and agree to be accountable to all aspects of this work
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical approval
All patients included in this study provided written informed consent for MGPT as well as for additional exploratory research analyses. In this context, approval by an ethics committee was not required.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Buecher, B., Warcoin, M., Rolland, E. et al. Results of a multigene panel testing approach targeting patients with suspected genetic predisposition to pancreatic ductal adenocarcinoma. Eur J Hum Genet (2026). https://doi.org/10.1038/s41431-026-02020-1
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41431-026-02020-1