Abstract
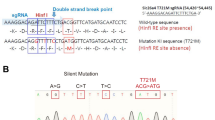
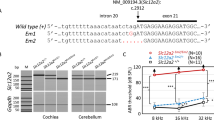
SLC26A4 encodes pendrin, a crucial anion exchanger essential for maintaining hearing function. Mutations in SLC26A4, including the prevalent c.919-2 A > G splice-site mutation among East Asian individuals, can disrupt inner ear electrolyte balance, leading to syndromic and non-syndromic hearing loss, such as Pendred syndrome and DFNB4. To explore potential therapeutic strategies, we utilized CRISPR/Cas9-mediated exon skipping to create a Slc26a4∆E8+E9/∆E8+E9 mouse model. We assessed pendrin expression in the inner ear and evaluated vestibular and auditory functions. The Slc26a4∆E8+E9/∆E8+E9 mice demonstrated reframed pendrin in the inner ear and normal vestibular functions, contrasting with severely abnormal vestibular functions observed in the Slc26a4 c.919-2 A > G splicing mutation mouse model. However, despite these molecular achievements, hearing function did not show the expected improvement, consistent with observed pathology, including cochlear hair cell loss and elevated hearing thresholds. Consequently, our findings highlight the necessity for alternative genetic editing strategies to address hearing loss caused by the SLC26A4 c.919-2 A > G mutation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 6 print issues and online access
$259.00 per year
only $43.17 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
All the data produced in this study have been either included in the published paper or are accessible through the lead contact upon request.
References
Korver AM, Smith RJ, Van Camp G, Schleiss MR, Bitner-Glindzicz MA, Lustig LR, et al. Congenital hearing loss. Nat Rev Dis Prim. 2017;3:16094.
Carpena NT, Lee MY. Genetic hearing loss and gene therapy. Genom Inf. 2018;16:e20.
Nicolson T. Navigating hereditary hearing loss: pathology of the inner ear. Front Cell Neurosci. 2021;15:660812.
Roesch S, Rasp G, Sarikas A, Dossena S. Genetic determinants of non-syndromic enlarged vestibular aqueduct: a review. Audio Res. 2021;11:423–42.
Amlal H, Petrovic S, Xu J, Wang Z, Sun X, Barone S, et al. Deletion of the anion exchanger Slc26a4 (pendrin) decreases apical Cl(-)/HCO3(-) exchanger activity and impairs bicarbonate secretion in kidney collecting duct. Am J Physiol Cell Physiol. 2010;299:C33–41.
Royaux IE, Belyantseva IA, Wu T, Kachar B, Everett LA, Marcus DC, et al. Localization and functional studies of pendrin in the mouse inner ear provide insight about the etiology of deafness in pendred syndrome. J Assoc Res Otolaryngol. 2003;4:394–404.
Campbell C, Cucci RA, Prasad S, Green GE, Edeal JB, Galer CE, et al. Pendred syndrome, DFNB4, and PDS/SLC26A4 identification of eight novel mutations and possible genotype-phenotype correlations. Hum Mutat. 2001;17:403–11.
Wu CC, Yeh TH, Chen PJ, Hsu CJ. Prevalent SLC26A4 mutations in patients with enlarged vestibular aqueduct and/or Mondini dysplasia: a unique spectrum of mutations in Taiwan, including a frequent founder mutation. Laryngoscope. 2005;115:1060–4.
Tsukamoto K, Suzuki H, Harada D, Namba A, Abe S, Usami S. Distribution and frequencies of PDS (SLC26A4) mutations in Pendred syndrome and nonsyndromic hearing loss associated with enlarged vestibular aqueduct: a unique spectrum of mutations in Japanese. Eur J Hum Genet. 2003;11:916–22.
Wu J, Cao Z, Su Y, Wang Y, Cai R, Chen J, et al. Molecular diagnose of a large hearing loss population from China by targeted genome sequencing. J Hum Genet. 2022;67:643–49.
Anwar S, Riazuddin S, Ahmed ZM, Tasneem S, Ateeq Ul J, Khan SY, et al. SLC26A4 mutation spectrum associated with DFNB4 deafness and Pendred’s syndrome in Pakistanis. J Hum Genet. 2009;54:266–70.
Del Castillo I, Morin M, Dominguez-Ruiz M, Moreno-Pelayo MA. Genetic etiology of non-syndromic hearing loss in Europe. Hum Genet. 2022;141:683–96.
Chen N, Tranebjaerg L, Rendtorff ND, Schrijver I. Mutation analysis of SLC26A4 for Pendred syndrome and nonsyndromic hearing loss by high-resolution melting. J Mol Diagn. 2011;13:416–26.
Dai P, Li Q, Huang D, Yuan Y, Kang D, Miller DT, et al. SLC26A4 c.919-2A>G varies among Chinese ethnic groups as a cause of hearing loss. Genet Med. 2008;10:586–92.
Lee HJ, Jung J, Shin JW, Song MH, Kim SH, Lee JH, et al. Correlation between genotype and phenotype in patients with bi-allelic SLC26A4 mutations. Clin Genet. 2014;86:270–5.
Min YL, Li H, Rodriguez-Caycedo C, Mireault AA, Huang J, Shelton JM, et al. CRISPR-Cas9 corrects Duchenne muscular dystrophy exon 44 deletion mutations in mice and human cells. Sci Adv. 2019;5:eaav4324.
Feng P, Xu Z, Chen J, Liu M, Zhao Y, Wang D, et al. Rescue of mis-splicing of a common SLC26A4 mutant associated with sensorineural hearing loss by antisense oligonucleotides. Mol Ther Nucleic acids. 2022;28:280–92.
Hwang J, Yokota T. Recent advancements in exon-skipping therapies using antisense oligonucleotides and genome editing for the treatment of various muscular dystrophies. Expert Rev Mol Med. 2019;21:e5.
Li X, Sanneman JD, Harbidge DG, Zhou F, Ito T, Nelson R, et al. SLC26A4 targeted to the endolymphatic sac rescues hearing and balance in Slc26a4 mutant mice. PLoS Genet. 2013;9:e1003641.
Choi BY, Kim HM, Ito T, Lee KY, Li X, Monahan K, et al. Mouse model of enlarged vestibular aqueducts defines temporal requirement of Slc26a4 expression for hearing acquisition. J Clin Invest. 2011;121:4516–25.
Rao SR, Olechnowicz SWZ, Kratschmer P, Jepson JEC, Edwards CM, Edwards JR. Small animal video tracking for activity and path analysis using a novel open-source multi-platform application (AnimApp). Sci Rep. 2019;9:12343.
Lu YC, Wu CC, Shen WS, Yang TH, Yeh TH, Chen PJ, et al. Establishment of a knock-in mouse model with the SLC26A4 c.919-2A>G mutation and characterization of its pathology. PLoS One. 2011;6:e22150.
Choi HJ, Lee HJ, Choi JY, Jeon IH, Noh B, Devkota S, et al. DNAJC14 ameliorates inner ear degeneration in the DFNB4 mouse model. Mol Ther Methods Clin Dev. 2020;17:188–97.
Baek M, Baker D. Deep learning and protein structure modeling. Nat Methods. 2022;19:13–14.
Ge J, Elferich J, Dehghani-Ghahnaviyeh S, Zhao Z, Meadows M, von Gersdorff H, et al. Molecular mechanism of prestin electromotive signal amplification. Cell. 2021;184:4669–4679.e4613.
Griffith AJ, Wangemann P. Hearing loss associated with enlargement of the vestibular aqueduct: mechanistic insights from clinical phenotypes, genotypes, and mouse models. Hear Res. 2011;281:11–17.
Ito T, Muskett J, Chattaraj P, Choi BY, Lee KY, Zalewski CK, et al. SLC26A4 mutation testing for hearing loss associated with enlargement of the vestibular aqueduct. World J Otorhinolaryngol. 2013;3:26–34.
Liu Q, Zhang X, Huang H, Chen Y, Wang F, Hao A, et al. Asymmetric pendrin homodimer reveals its molecular mechanism as anion exchanger. Nat Commun. 2023;14:3012.
Dehghani-Ghahnaviyeh S, Zhao Z, Tajkhorshid E. Lipid-mediated prestin organization in outer hair cell membranes and its implications in sound amplification. Nat Commun. 2022;13:6877.
Whitfield TT. Development of the inner ear. Curr Opin Genet Dev. 2015;32:112–8.
Acknowledgements
We thank the National Laboratory Animal Center (NLAC), NARLabs, Taiwan, for technical support in contract breeding and testing services. We are also grateful to Dr. Yu-Chi Chou in the RNA Technology Platform and Gene Manipulation Core Facility (RNAi core) of the National Core Facility for Biopharmaceuticals at Academia Sinica in Taiwan for providing CRISPR reagents and related services. This work was supported by grants from the National Science and Technology Council (NSTC 111-2314-B-A49A-504-MY3), the Taipei Veterans General Hospital (V110C-123, V112C-141), the National Health Research Institutes (NHRI-EX112-11005NI), and the Taipei Veterans General Hospital-National Taiwan University Hospital Joint Research Program (VN111-16, VN112-08).
Author information
Authors and Affiliations
Contributions
YFC and CCW contributed to the study design. CYH and YFC contributed to the mouse model design. PTW, PYH, and YCC contributed to mouse breeding and genotyping. YHT and PTW contributed to fluorescence and H&E staining. YFC and PTW contributed to vestibular function. JSL and YCC contributed to protein structure modeling. CYH, YHT, YFC, and YFC contributed to the writing of the manuscript. All the authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Huang, CY., Tsai, YH., Cheng, YF. et al. CRISPR/Cas9-mediated exon skipping to restore premature translation termination in a DFNB4 mouse model. Gene Ther 31, 531–540 (2024). https://doi.org/10.1038/s41434-024-00483-9
Received:
Revised:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41434-024-00483-9