Abstract
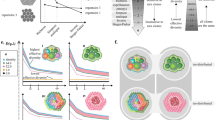
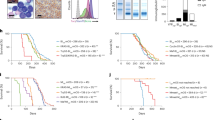
Myasthenia gravis (MG) is a T/B cell-driven autoimmune disease. The immunomodulatory mechanisms of the common immunosuppressant tacrolimus (TAC) on the immune repertoire are unclear. This study investigated TAC’s immunomodulatory effects via high-throughput sequencing of peripheral blood mononuclear cells from four MG patients pre- and post-four months of TAC monotherapy, revealing dynamic T-cell receptor (TCR) and B-cell receptor (BCR) repertoire remodeling. The immune repertoire of MG patients was characterized by a skewed usage of TRBV gene families compared to healthy controls, indicating an underlying immune dysfunction. Longitudinal analysis post-TAC therapy revealed potential downregulation of IGHV1-69 and IGHV3-43 gene frequencies (p < 0.05, FDR > 0.1), alongside non-significant trends toward shorter CDR3 lengths and reduced clonal diversity in both IGH and TRB (p > 0.05). The BCR repertoire underwent greater dynamic remodeling, while the TCR repertoire remained relatively stable, as evidenced by the persistence of dominant clones and overlapping TRB clones. BCR diversity and V or J gene usage in TCR and BCR showed potential associations with clinical severity. These data reveal skewed antigen recognition profiles in MG pathogenesis, with TAC orchestrating multimodal immunomodulation through peripheral immune repertoire reshaping.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 6 digital issues and online access to articles
$119.00 per year
only $19.83 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The data that support the findings of this study are available on request from the corresponding author.
References
Kaminski HJ, Sikorski P, Coronel SI, Kusner LL Myasthenia gravis: the future is here. J Clin Invest. 2024;134:e179742.
Uzawa A, Kuwabara S, Suzuki S, Imai T, Murai H, Ozawa Y, et al. Roles of cytokines and T cells in the pathogenesis of myasthenia gravis. Clin Exp Immunol. 2021;203:366–74.
Yi JS, Guptill JT, Stathopoulos P, Nowak RJ, O’Connor KC. B cells in the pathophysiology of myasthenia gravis. Muscle Nerve. 2018;57:172–84.
Fichtner ML, Jiang R, Bourke A, Nowak RJ, O’Connor KC. Autoimmune pathology in myasthenia gravis disease subtypes is governed by divergent mechanisms of immunopathology. Front Immunol. 2020;11:776.
Zhao CB, Zhang X, Zhang H, Hu XQ, Lu JH, Lu CZ, et al. Clinical efficacy and immunological impact of tacrolimus in Chinese patients with generalized myasthenia gravis. Int Immunopharmacol. 2011;11:519–24.
Fan Z, Li Z, Shen F, Zhang X, Lei L, Su S, et al. Favorable effects of tacrolimus monotherapy on myasthenia gravis patients. Front Neurol. 2020;11:594152.
Itani K, Nakamura M, Wate R, Kaneko S, Fujita K, Iida S, et al. Efficacy and safety of tacrolimus as long-term monotherapy for myasthenia gravis. Neuromuscul Disord. 2021;31:512–8.
Duan W, Peng Y, Jin W, Ouyang S, Yang H. Tacrolimus as single-agent immunotherapy and minimal manifestation status in nonthymoma myasthenia gravis. J Immunol Res. 2021;2021:9138548.
Fan Z, Lei L, Su S, Zhang S, Xie N, Li L, et al. Comparison between mono-tacrolimus and mono-glucocorticoid in the treatment of myasthenia gravis. Ann Clin Transl Neurol. 2023;10:589–98.
Sathasivam S. Steroids and immunosuppressant drugs in myasthenia gravis. Nat Clin Pr Neurol. 2008;4:317–27.
Wu H, Chen L, Zhou X, Wu Y, Yan Y, Zhu Y, et al. Effect of tacrolimus on soluble costimulatory molecules in patients with refractory myasthenia gravis. J Neuroimmunol. 2022;372:577955.
Wu H, Wang Z, Xi J, Liu J, Yan C, Song J, et al. Therapeutic and immunoregulatory effects of tacrolimus in patients with refractory generalized myasthenia gravis. Eur Neurol. 2020;83:500–7.
Bao J, Gao S, Weng Y, Zhu J, Ye H, Zhang X. Clinical efficacy of tacrolimus for treating myasthenia gravis and its influence on lymphocyte subsets. Rev Neurol. 2019;175:65–72.
De Bruyne R, Bogaert D, De Ruyck N, Lambrecht BN, Van Winckel M, Gevaert P, et al. Calcineurin inhibitors dampen humoral immunity by acting directly on naive B cells. Clin Exp Immunol. 2015;180:542–50.
Nielsen SCA, Boyd SD. Human adaptive immune receptor repertoire analysis-past, present, and future. Immunol Rev. 2018;284:9–23.
Minervina A, Pogorelyy M, Mamedov I. T-cell receptor and B-cell receptor repertoire profiling in adaptive immunity. Transpl Int. 2019;32:1111–23.
Xu BY, Giscombe R, Söderlund A, Troye-Blomberg M, Pirskanen R, Lefvert AK. Abnormal T cell receptor V gene usage in myasthenia gravis: prevalence and characterization of expanded T cell populations. Clin Exp Immunol. 1998;113:456–64.
Fozza C, Barraqueddu F, Corda G, Contini S, Virdis P, Dore F, et al. Study of the T-cell receptor repertoire by CDR3 spectratyping. J Immunol Methods. 2017;440:1–11.
Hou XL, Wang L, Ding YL, Xie Q, Diao HY. Current status and recent advances of next generation sequencing techniques in immunological repertoire. Genes Immun. 2016;17:153–64.
Kanai T, Uzawa A, Kawaguchi N, Himuro K, Oda F, Ozawa Y, et al. Adequate tacrolimus concentration for myasthenia gravis treatment. Eur J Neurol. 2017;24:270–5.
Zhong H, Ruan Z, Yan C, Lv Z, Zheng X, Goh LY, et al. Short-term outcome prediction for myasthenia gravis: an explainable machine learning model. Ther Adv Neurol Disord. 2023;16:17562864231154976.
Chen S, Zhou Y, Chen Y, Gu J. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics. 2018;34:i884–90.
Bolotin DA, Poslavsky S, Mitrophanov I, Shugay M, Mamedov IZ, Putintseva EV, et al. MiXCR: software for comprehensive adaptive immunity profiling. Nat Methods. 2015;12:380–1.
Lefranc MP. IMGT, the international immunogenetics information system. Cold Spring Harb Protoc. 2011;2011:595–603.
Huang H, Wang C, Rubelt F, Scriba TJ, Davis MM. Analyzing the mycobacterium tuberculosis immune response by T-cell receptor clustering with GLIPH2 and genome-wide antigen screening. Nat Biotechnol. 2020;38:1194–202.
Vander Heiden JA, Stathopoulos P, Zhou JQ, Chen L, Gilbert TJ, Bolen CR, et al. Dysregulation of B cell repertoire formation in myasthenia gravis patients revealed through deep sequencing. J Immunol. 2017;198:1460–73.
Miao Y, Shi Z, Zhang W, Zhu L, Tang S, Chen H, et al. Immune repertoire profiling reveals its clinical application potential and triggers for neuromyelitis optica spectrum disorders. Neurol Neuroimmunol Neuroinflamm. 2023;10:e200134.
Stewart JJ, Lee CY, Ibrahim S, Watts P, Shlomchik M, Weigert M, et al. A Shannon entropy analysis of immunoglobulin and T cell receptor. Mol Immunol. 1997;34:1067–82.
Huang T, Pi C, Xu X, Feng Y, Zhang J, Gu H, et al. Effect of BAFF blockade on the B cell receptor repertoire and transcriptome in a mouse model of systemic lupus erythematosus. Front Immunol. 2023;14:1307392.
van Bladel DAG, van den Brand M, Rijntjes J, Pamidimarri Naga S, Haacke D, Luijks J, et al. Clonality assessment and detection of clonal diversity in classic Hodgkin lymphoma by next-generation sequencing of immunoglobulin gene rearrangements. Mod Pathol. 2022;35:757–66.
Grunewald J, Ahlberg R, Lefvert AK, DerSimonian H, Wigzell H, Janson CH. Abnormal T-cell expansion and V-gene usage in myasthenia gravis patients. Scand J Immunol. 1991;34:161–8.
Gigliotti D, Lefvert AK, Jeddi-Tehrani M, Esin S, Hodara V, Pirskanen R, et al. Overexpression of select T cell receptor V beta gene families within CD4+ and CD8+T cell subsets of myasthenia gravis patients: a role for superantigen(s)? Mol Med. 1996;2:452–9.
Lu C, Pi X, Xu W, Qing P, Tang H, Li Y, et al. Clinical significance of T cell receptor repertoire in primary Sjogren’s syndrome. EBioMedicine. 2022;84:104252.
Lee Y, Kim SW, Lee E, Shin HY, Kim M, Lee CY, et al. Stereotypic T cell receptor clonotypes in the thymus and peripheral blood of Myasthenia gravis patients. Heliyon. 2024;10:e26663.
Arvidsson G, Czarnewski P, Johansson A, Raine A, Imgenberg-Kreuz J, Nordlund J, et al. Multimodal single-cell sequencing of B cells in primary Sjögren’s syndrome. Arthritis Rheumatol. 2024;76:255–67.
Pham MC, Masi G, Patzina R, Obaid AH, Oxendine SR, Oh S, et al. Individual myasthenia gravis autoantibody clones can efficiently mediate multiple mechanisms of pathology. Acta Neuropathol. 2023;146:319–36.
Masi G, Chen K, Bayer AC, Bayarri-Olmos R, Pham MC, Wallace A, et al. IgA autoantibodies demonstrate a novel mechanism of MuSK myasthenia gravis pathology. Brain. 2025;10:awaf410.
Zimmermann M, Rose N, Lindner JM, Kim H, Gonçalves AR, Callegari I, et al. Antigen extraction and B cell activation enable identification of rare membrane antigen specific human B cells. Front Immunol. 2019;10:829.
Liu S, Hou XL, Sui WG, Lu QJ, Hu YL, Dai Y. Direct measurement of B-cell receptor repertoire’s composition and variation in systemic lupus erythematosus. Genes Immun. 2017;18:22–7.
Bashford-Rogers RJM, Bergamaschi L, McKinney EF, Pombal DC, Mescia F, Lee JC, et al. Analysis of the B cell receptor repertoire in six immune-mediated diseases. Nature. 2019;574:122–6.
Wang Q, Feng D, Jia S, Lu Q, Zhao M. B-cell receptor repertoire: recent advances in autoimmune diseases. Clin Rev Allergy Immunol. 2024;66:76–98.
Wu M, Pan W, Jia C, He Z, Zhao M, Tang C, et al. Systemic lupus erythematosus patients contain B-cell receptor repertoires sensitive to immunosuppressive drugs. Eur J Immunol. 2022;52:669–80.
van Lieverloo GGA, Al-Soudi A, Wieske L, Klarenbeek PL, Anang DC, Adrichem ME, et al. B-cell and T-cell receptor repertoire in chronic inflammatory demyelinating polyneuropathy, a prospective cohort study. J Peripher Nerv Syst. 2023;28:69–78.
Jiang R, Fichtner ML, Hoehn KB, Pham MC, Stathopoulos P, Nowak RJ, et al. Single-cell repertoire tracing identifies Rituximab-resistant B cells during myasthenia gravis relapses. JCI Insight. 2020;5:e136471.
Fichtner ML, Hoehn KB, Ford EE, Mane-Damas M, Oh S, Waters P, et al. Reemergence of pathogenic, autoantibody-producing B cell clones in myasthenia gravis following B cell depletion therapy. Acta Neuropathol Commun. 2022;10:154.
Nagafuchi Y, Ota M, Hatano H, Inoue M, Kobayashi S, Okubo M, et al. Control of naive and effector CD4 T cell receptor repertoires by rheumatoid-arthritis-risk HLA alleles. J Autoimmun. 2022;133:102907.
Sorosina M, Santoro S, Ferrè L, Mascia E, Clarelli F, Giordano A, et al. Risk HLA variants affect the T-cell repertoire in multiple sclerosis. Neurol Neuroimmunol Neuroinflamm. 2023;10:e200093.
Loh TJ, Lim JJ, Jones CM, Dao HT, Tran MT, Baker DG, et al. The molecular basis underlying T cell specificity towards citrullinated epitopes presented by HLA-DR4. Nat Commun. 2024;15:6201.
Galindo-Feria AS, Sharma RK, Dubnovitsky A, Gerstner C, Kozhukh G, Van Vollenhoven A et al. Autoreactive T cells identified in patients with anti-Jo1+ antisynthetase syndrome recognise a new epitope on histidyl t-RNA synthetase. Ann Rheum Dis. 2025;85:360–9.
Jiang R, Hoehn KB, Lee CS, Pham MC, Homer RJ, Detterbeck FC, et al. Thymus-derived B cell clones persist in the circulation after thymectomy in myasthenia gravis. Proc Natl Acad Sci USA. 2020;117:30649–60.
Acknowledgements
This study was supported by the National Natural Science Foundation of China (Grand No. 82371413 to Huan Yang).
Author information
Authors and Affiliations
Contributions
Huan Yang: conception, study design, funding acquisition, writing-review and editing; Ting He: writing-original draft, formal analysis, and visualization; Liu Wang: formal analysis, software, and visualization; Kangzhi Chen: data collection and software; Qian Zhou: data curation and investigation; Liqun Xu and Zhaohui Luo: writing-review and editing and supervision. All authors contributed to the article and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
He, T., Wang, L., Chen, K. et al. Longitudinal omics reveals immune repertoire remodeling in myasthenia gravis patients post-tacrolimus therapy. Genes Immun (2026). https://doi.org/10.1038/s41435-026-00395-1
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41435-026-00395-1