Abstract
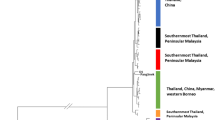
It is important to have clearly delineated taxonomic units informed by eco-evolutionary processes to ensure effective implementation of conservation efforts. For a group as threatened as Asian pangolins, a timely understanding and recognition of cryptic species is especially critical to inform policy and management decisions. Recent genomic investigations into the group have revealed genomic distinctions between currently recognised species and the putative Manis cf. mysteria, but only incorporate limited representation of the new lineage and its sister taxa, M. javanica and M. culionensis, which may obscure evolutionary inferences within the group and overlook important genetic variation at species boundaries. In this study, we broadly sample seizure materials to incorporate as holistic a representation of each lineage as possible so as to further verify genomic distinctions between the lineages, including new samples of M. cf. mysteria and M. culionensis, the latter of which was only genomically represented by one museum sample previously. We find that while M. cf. mysteria, M. javanica and M. culionensis do form distinct clades with deep divergences, new samples of M. cf. mysteria demonstrate mitonuclear discordance and admixture with M. javanica, illuminating more evolutionary complexity between the three lineages than previously reported. We also find much higher variation in individual heterozygosity within M. cf. mysteria than its sister taxa. Our findings highlight gaps in our understanding of the contemporary evolutionary dynamics of Southeast Asian pangolins, but also demonstrate the barriers these gaps create for practical and timely implementations of conservation effort trafficked taxa at large.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
Raw Illumina sequences along with associated metadata of each sample and assembled mitogenomes are deposited in the NCBI Sequence Read Archive and GenBank under BioProject PRJNA1289811.
References
Allio R, Schomaker-Bastos A, Romiguier J, Prosdocimi F, Nabholz B, Delsuc F (2020) MitoFinder: efficient automated large-scale extraction of mitogenomic data in target enrichment phylogenomics. Mol Ecol Resour 20:892–905
Alpers DL, Van Vuuren BJ, Arctander P, Robinson TJ (2004) Population genetics of the roan antelope (Hippotragus equinus) with suggestions for conservation. Mol Ecol 13:1771–1784
Archer LJ, Papworth SK, Apale CM, Corona DB, Gacilos JT, Amada RL et al (2020) Scaling up local ecological knowledge to prioritise areas for protection: determining Philippine pangolin distribution, status and threats. Glob Ecol Conserv 24: e01395
Bickford D, Lohman DJ, Sodhi NS, Ng PK, Meier R, Winker K et al (2007) Cryptic species as a window on diversity and conservation. Trends Ecol Evol 22:148–155
de Bruyn M, Stelbrink B, Morley RJ, Hall R, Carvalho GR, Cannon CH et al (2014) Borneo and Indochina are major evolutionary hotspots for Southeast Asian biodiversity. Syst Biol 63:879–901
Cao P, Dai Q, Deng C, Zhao X, Qin S, Yang J et al (2021) Genome-wide signatures of mammalian skin covering evolution. Sci China Life Sci 64:1765–1780
Cascini M, Doyle CA, Mulcahy A, McMaster ES, Dimon R, Hogbin PM et al (2025) The impact of taxonomic confusion on conservation resources – Why population genomics should inform threatened species determination. Biol Conserv 306: 111113
Challender DWS, Waterman C, Baillie JEM (2014) Scaling up pangolin conservation: IUCN SSC Pangolin Specialist Group Action Plan. Zoological Society of London, London, UK
Challender DWS, ‘t Sas-Rolfes M, Ades GW, Chin JS, Sun NCM, Chong J et al (2019a) Evaluating the feasibility of pangolin farming and its potential conservation impact. Glob Ecol Conserv 20: e00714
Challender DWS, Willcox DHA, Panjang E, Lim N, Nash H, Heinrich S et al (2019b) Manis javanica. The IUCN Red List of Threatened Species 2019: e.T12763A123584856. https://doi.org/10.2305/IUCN.UK.2019-3.RLTS.T12763A123584856.en. Accessed 08 Sept 2022
Challender DWS, Wu S, Kaspal P, Khatiwada A, Ghose A, Sun NCM et al. (2019c) Manis pentadactyla (errata version published in 2020). The IUCN Red List of Threatened Species 2019: e.T12764A168392151. https://doi.org/10.2305/IUCN.UK.2019-3.RLTS.T12764A168392151.en. Accessed 19 October 2022
Chattopadhyay B, Garg KM, Mendenhall IH, Rheindt FE (2019) Historic DNA reveals Anthropocene threat to a tropical urban fruit bat. Curr Biol 29:R1299–R1300
Chen S, Zhou Y, Chen Y, Gu J (2018) fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34:i884–i890
Coffroth MA, Lasker HR, Diamond ME, Bruenn JA, Bermingham E (1992) DNA fingerprints of a gorgonian coral: a method for detecting clonal structure in a vegetative species. Mar Biol 114:317–325
Cruickshank TE, Hahn MW (2014) Reanalysis suggests that genomic islands of speciation are due to reduced diversity, not reduced gene flow. Mol Ecol 23:3133–3157
Danecek P, Bonfield JK, Liddle J, Marshall J, Ohan V, Pollard MO et al (2021) Twelve years of SAMtools and BCFtools. GigaScience 10: giab008
Davis HR, Das I, Leaché AD, Karin BR, Brennan IG, Jackman TR et al (2021) Genetically diverse yet morphologically conserved: Hidden diversity revealed among Bornean geckos (Gekkonidae: Cyrtodactylus). J Zool Syst Evol Res 59:1113–1135
Donath A, Jühling F, Al-Arab M, Bernhart SH, Reinhardt F, Stadler PF et al (2019) Improved annotation of protein-coding genes boundaries in metazoan mitochondrial genomes. Nucleic Acids Res 47:10543–10552
Dierckxsens N, Mardulyn P, Smits G (2016) NOVOPlasty: de novo assembly of organelle genomes from whole genome data. Nucleic Acids Res 45: e18
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620
Ewart KM, Lightson AL, Sitam FT, Rovie-Ryan J, Nguyen SG, Morgan KI et al (2021) DNA analyses of large pangolin scale seizures: Species identification validation and case studies. Forensic Sci Int: Animals Environ 1: 100014
Funk WC, McKay JK, Hohenlohe PA, Allendorf FW (2012) Harnessing genomics for delineating conservation units. Trends Ecol Evol 27:489–496
Garcia-Erill G, Albrechtsen A (2020) Evaluation of model fit of inferred admixture proportions. Mol Ecol Resour 20:936–949
Gaubert P, Antunes A, Meng H, Miao L, Peigné S, Justy F et al (2018) The complete phylogeny of pangolins: scaling up resources for the molecular tracing of the most trafficked mammals on earth. J Hered 109:347–359
Gu TT, Wu H, Yang F, Gaubert P, Heighton SP, Fu Y et al (2023) Genomic analysis reveals a cryptic pangolin species. Proc Natl Acad Sci USA 120: e2304096120
Guo B, Fang B, Shikano T, Momigliano P, Wang C, Kravchenko A et al (2019) A phylogenomic perspective on diversity, hybridization and evolutionary affinities in the stickleback genus Pungitius. Mol Ecol 28:4046–4064
Hanghøj K, Moltke I, Andersen PA, Manica A, Korneliussen TS (2019) Fast and accurate relatedness estimation from high-throughput sequencing data in the presence of inbreeding. GigaScience 8: giz034
Harris DJ, Froufe E (2005) Taxonomic inflation: species concept or historical geopolitical bias. Trends Ecol Evol 20:6–7
Heighton SP, Allio R, Murienne J, Salmona J, Meng H, Scornavacca C et al (2023) Pangolin genomes offer key insights and resources for the world’s most trafficked wild mammals. Mol Biol Evol 40: msad190
Hendry AP, Lohmann LG, Conti E, Cracraft J, Crandall KA, Faith DP et al (2010) Evolutionary biology in biodiversity science, conservation, and policy: a call to action. Evolution 64:1517–1528
Hohenlohe PA, Funk WC, Rajora OP (2020) Population genomics for wildlife conservation and management. Mol Ecol 30:62–82
Hu JY, Hao ZQ, Frantz L, Wu SF, Chen W, Jiang YF et al (2020a) Genomic consequences of population decline in critically endangered pangolins and their demographic histories. Natl Sci Rev 7:798–814
Hu J, Roos C, Lv X, Kuang W, Yu L (2020b) Molecular genetics supports a potential fifth Asian pangolin species (Mammalia, Pholidota, Manis). Zool Sci 37:538–543
Hughes LJ, Morton O, Scheffers BR, Edwards DP (2023) The ecological drivers and consequences of wildlife trade. Biol Rev 98:775–791
Jacobs RL, Baker BW (2018) The species dilemma and its potential impact on enforcing wildlife trade laws. Evol Anthropol 21:261–266
Johnson RN, Wilson-Wilde L, Linacre A (2014) Current and future directions of DNA in wildlife forensic science. Forensic Sci Int Genet 10:1–11
Jouganous J, Long W, Ragsdale AP, Gravel S (2017) Inferring the joint demographic history of multiple populations: beyond the diffusion approximation. Genetics 206:1549–1567
Kalyaanamoorthy S, Minh BQ, Wong TK, Von Haeseler A, Jermiin LS (2017) ModelFinder: fast model selection for accurate phylogenetic estimates. Nat Methods 14:587–589
Katoh K, Misawa K, Kuma KI, Miyata T (2002) MAFFT: a novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res 30:3059–3066
Kinnison MT, Hairston NG (2007) Eco-evolutionary conservation biology: contemporary evolution and the dynamics of persistence. Funct Ecol 21:444–454
Korneliussen TS, Albrechtsen A, Nielsen R (2014) ANGSD: analysis of next generation sequencing data. BMC Bioinform 15:1–13.
Kousathanas A, Leuenberger C, Link V, Sell C, Burger J, Wegmann D (2017) Inferring Heterozygosity from Ancient and Low Coverage Genomes. Genetics 205:317–332
Leigh JW, Bryant D (2015) POPART: full-feature software for haplotype network construction. Methods Ecol Evol 6:1110–1116
Li B, Li H, Shi M, Wang Q, Li H, Guo C et al (2026) Population genomics reveals deep diversification in Malayan pangolins. Mol Biol Evol 3:msag016
Li H (2011) Improving SNP discovery by base alignment quality. Bioinformatics 27:1157–1158
Li H (2013) Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. arXiv. https://doi.org/10.48550/arXiv.1303.3997.
Lou RN, Therkildsen NO (2021) Batch effects in population genomic studies with low-coverage whole genome sequencing data: causes, detection and mitigation. Mol Ecol Resour 22:1678–1692
Mace GM (2004) The role of taxonomy in species conservation. Phil Trans R Soc B 359:711–719
Mace GM, Purvis A (2007) Evolutionary biology and practical conservation: bridging a widening gap. Mol Ecol 17:9–19
Meisner J, Albrechtsen A (2018) Inferring population structure and admixture proportions in low-depth NGS data. Genetics 210:719–731
Momigliano P, Florin AB, Merilä J (2021) Biases in demographic modeling affect our understanding of recent divergence. Mol Biol Evol 38:2967–2985
Morin DJ, Challender DWS, Ichu IG, Ingram DJ, Nash HC, Panaino W et al (2020) Developing robust ecological monitoring methodologies for pangolin conservation. In: Challender DWS, Nash HC, Waterman C (eds) Pangolins: science, society and conservation. Academic Press, pp 545–558
Nash, Wirdateti HC, Low GW, Choo SW, Chong JL, Semiadi G et al. (2018) Conservation genomics reveals possible illegal trade routes and admixture across pangolin lineages in Southeast Asia. Conserv Genet 19:1083–1095
NCBI (2025) BLAST: Basic Local Alignment Search Tool. https://blast.ncbi.nlm.nih.gov.
Nei M, Li WH (1979) Mathematical model for studying genetic variation in terms of restriction endonucleases. Proc Natl Acad Sci USA 76:5269–5273
Nguyen LT, Schmidt HA, Von Haeseler A, Minh BQ (2015) IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol Biol Evol 32:268–274
Niemiller ML, Graening GO, Fenolio DB, Godwin JC, Cooley JR, Pearson WD et al (2013) Doomed before they are described? The need for conservation assessments of cryptic species complexes using an amblyopsid cavefish (Amblyopsidae: Typhlichthys) as a case study. Biodivers Conserv 22:1799–1820
Nijman V, Zhang MX, Shepherd CR (2016) Pangolin trade in the Mong La wildlife market and the role of Myanmar in the smuggling of pangolins into China. Glob Ecol Conserv 5:118–126
Novikova PY, Hohmann N, Nizhynska V, Tsuchimatsu T, Ali J, Muir G et al (2016) Sequencing of the genus Arabidopsis identifies a complex history of nonbifurcating speciation and abundant trans-specific polymorphism. Nat Genet 48:1077–1082
Pesole G, Gissi C, De Chirico A, Saccone C (1999) Nucleotide substitution rate of mammalian mitochondrial genomes. J Mol Evol 48:427–434
Prüfer K, Munch K, Hellmann I, Akagi K, Miller JR, Walenz B et al. (2012) The bonobo genome compared with the chimpanzee and human genomes. Nature 486:527–531.
R Core Team (2021) R: a language and environment for statistical computing. https://www.R-project.org/.
Rambaut A (2018) FigTree v1.4.4. https://github.com/rambaut/figtree.
Rasmussen MS, Garcia-Erill G, Korneliussen TS, Wiuf C, Albrechtsen A (2022) Estimation of site frequency spectra from low-coverage sequencing data using stochastic EM reduces overfitting, runtime, and memory usage. Genetics 222: iyac148
Rheindt FE, Wu MY, Movin N, Jønsson KA (2022) Cryptic species-level diversity in Dark-throated Oriole Oriolus xanthonotus. Bull Br Ornithol Club 142:254–267
Rojas M (1992) The species problem and conservation: what are we protecting? Conserv Biol 6:170–178
Roos MC, Kessler PJ, Robbert Gradstein S, Baas P (2004) Species diversity and endemism of five major Malesian islands: diversity–area relationships. J Biogeogr 31:1893–1908
Roux C, Fraisse C, Romiguier J, Anciaux Y, Galtier N, Bierne N (2016) Shedding light on the grey zone of speciation along a continuum of genomic divergence. PLoS Biol 14:e2000234
Runemark A, Vallejo-Marin M, Meier JI (2019) Eukaryote hybrid genomes. PLoS Genet 15:e1008404
Schoppe S, Katsis L, Lagrada L (2019) Manis culionensis. The IUCN Red List of Threatened Species 2019: e.T136497A123586862. https://doi.org/10.2305/IUCN.UK.2019-3.RLTS.T136497A123586862.en. Accessed 08 Sept 2022
Schoppe S, Katsis LK, Alvarado D, Acosta-Lagrada L (2020) Philippine pangolin Manis culionensis (de Elera 1915). In: Challender DWS, Nash HC, Waterman C (eds) Pangolins: science, society and conservation. Academic Press, pp 109–122
Sitam FT, Salgado-Lynn M, Denel A, Panjang E, McEwing R, Lightson A et al. (2023) Phylogeography of the Sunda pangolin, Manis javanica: implications for taxonomy, conservation management and wildlife forensics. Ecol Evol 13: e10373
Skotte L, Korneliussen TS, Albrechtsen A (2013) Estimating individual admixture proportions from next generation sequencing data. Genetics 195:693–702
Soraggi S, Wiuf C, Albrechtsen A (2018) Powerful inference with the D-statistic on low-coverage whole-genome data. G3: Genes Genomes Genet 8:551–566
Steiner CC, Putnam AS, Hoeck PE, Ryder OA (2013) Conservation genomics of threatened animal species. Annu Rev Anim Biosci 1:261–281
Tinsman JC, Gruppi C, Bossu CM, Prigge TL, Harrigan RJ, Zaunbrecher V et al. (2023) Genomic analyses reveal poaching hotspots and illegal trade in pangolins from Africa to Asia. Science 382:1282–1286
Toews DP, Brelsford A (2012) The biogeography of mitochondrial and nuclear discordance in animals. Mol Ecol 21:3907–3930
Tolley KA, Telford NS, Taft JM, Bates MF, Conradie W, Makhubo BG, Alexander GJ (2022) Taxonomic inflation due to inadequate sampling: are girdled lizards (Cordylus minor species complex) from the Great Karoo one and the same? Biol J Linn Soc 135:1–24
Van der Auwera GA, O’Connor BD (2020) Genomics in the cloud: using Docker, GATK, and WDL in Terra. O’Reilly Media, USA
Volleth M, Müller S, Heller KG, Trifonov V, Liehr T, Yong HS et al. (2022) Cytogenetic analyses detect cryptic diversity in Megaderma spasma from Malaysia. Acta Chiropt 23:271–284
Wang J (2018) Effects of sampling close relatives on some elementary population genetics analyses. Mol Ecol Resour 18:41–54
Wangmo LK, Ghosh A, Dolker S, Joshi BD, Sharma LK, Thakur M (2025a) Indo-Burmese pangolin (Manis indoburmanica): a novel phylogenetic species of pangolin evolved in Asia. Mamm Biol 105:1–8
Wangmo LK, Dolker S, Ghosh A, Joshi BD, Sharma LK, Thakur M (2025b) Reply to Zjlstra (2025) – Status of Indo-Burmese pangolin (Manis indoburmanica). Mamm Biol 105:704–707
Waples RK, Albrechtsen A, Moltke I (2018) Allele frequency-free inference of close familial relationships from genotypes or low-depth sequencing data. Mol Ecol 28:35–48
Waples RS, Anderson EC (2017) Purging putative siblings from population genetic data sets: a cautionary view. Mol Ecol 26:1211–1224
Wasser SK, Torkelson A, Winters M, Horeaux Y, Tucker S, Otiende MY et al. (2018) Combating transnational organized crime by linking multiple large ivory seizures to the same dealer. Sci Adv 4: eaat0625
Worthington BM, Wong PYH, Kumaree KK, Prigge TL, Ng KH, Liao Y et al. (2024) Serological evidence of Sarbecovirus exposure in seized Sunda pangolins points to complex origins of infection. BMC Biol 22: 274
Zhang H, Miller MP, Yang F, Chan HK, Gaubert P, Ades G et al. (2015) Molecular tracing of confiscated pangolin scales for conservation and illegal trade monitoring in Southeast Asia. Glob Ecol Conserv 4:414–422
Zijlstra JS (2025) Manis aurita Hodgson 1837 as a valid species of pangolin: a comment to Wangmo et al. (2025). Mamm Biol 105:699–701
Acknowledgements
This work was supported by Research Impact Fund (R7021-20), University Grants Committee, Hong Kong Special Administrative Region (HKSAR). Seized pangolin tissues were donated for use in this study by AFCD and KFBG in Hong Kong, and DENR in the Philippines. Bioinformatic analyses were performed using research computing facilities offered by Information Technology Services, The University of Hong Kong (HKU). We would also like to acknowledge the valuable input from Dr. Simon Sin, Dr. Nina Therkildsen, and Dr. Lidane Noronha regarding the manuscript.
Author information
Authors and Affiliations
Contributions
PYHW, YC, PM and TCB designed the research. PYHW, TLP, HZ, LRJ, GA, MRMD, IKCF and TCB facilitated and secured the donation of confiscated specimens. PYHW, TLP, AK, NIF, LKPN and SY processed specimens and collected DNA from samples. PYHW, YC, TLP and PM performed the research and conducted the data analysis. TTYL, PM and TCB advised on study design and bioinformatic analyses. PYHW, YC and TCB wrote the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Research ethics statement
All necessary CITES and local governmental permits were acquired for sampling.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Associate editor: Paul Sunnucks.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wong, P.YH., Chen, Y., Prigge, TL. et al. Seizure samples reveal complex evolutionary dynamics among Southeast Asian pangolins. Heredity 135, 249–258 (2026). https://doi.org/10.1038/s41437-026-00826-9
Received:
Revised:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41437-026-00826-9