Abstract
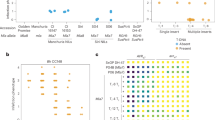
Stripe rust, caused by Puccinia striiformis f. sp. tritici, is a major threat to global wheat production. To explore new resistance resources, we screened 100 hexaploid triticale accessions using the predominant Chinese P. striiformis f. sp. tritici races CYR32, CYR33 and CYR34 and found that most accessions showed high resistance, with the cultivar Rozovskaya displaying near-immunity. Through map-based cloning, we identified a resistance gene located on chromosome 6RL. Analysis of resequencing data from 117 rye accessions revealed two major haplotypes, both of which conferred near-immunity and broadly effective resistance to stripe rust in transgenic wheat. Sequence analysis and virus-induced gene silencing collectively confirmed the identity of this gene as Yr83. Yr83 encodes an atypical nucleotide-binding and leucine-rich repeat protein (NLR) fused to a Harbinger transposase-derived nuclease domain (HTDND). Truncation of the HTDND abolishes resistance, indicating that this domain is essential for Yr83-mediated immune function. Phylogenetic analysis showed that NLR–HTDND proteins are restricted to the Pooideae subfamily. For breeding applications, we employed a small 6RL translocation line that shows excellent agronomic performance, not only conferring strong resistance but also increasing spikelet number and grain number per spike. Our study reveals a transposase-integrated NLR as a valuable resource for wheat stripe rust resistance breeding.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Beddow, J. M. et al. Research investment implications of shifts in the global geography of wheat stripe rust. Nat. Plants 1, 15132 (2015.
Zhao, J. & Kang, Z. Fighting wheat rusts in China: a look back and into the future. Phytopathol. Res. 5, 6 (2023).
Sharma, D. et al. A single NLR gene confers resistance to leaf and stripe rust in wheat. Nat. Commun. 15, 9925 (2024).
Hafeez, A. N. et al. Creation and judicious application of a wheat resistance gene atlas. Mol. Plant 14, 1053–1070 (2021).
Schwessinger, B. Fundamental wheat stripe rust research in the 21st century. New Phytol. 213, 1625–1631 (2017).
Hu, C. et al. Resistance analyses on wheat stripe rust resistance genes to the predominant races of Puccinia striiformis f. sp. tritici in China. Sci. Agric. Sin. 55, 491–502 (2022).
Jones, J. & Dangl, J. J. N. The plant immune system. Nature 444, 323–329 (2006).
Wan, L. et al. TIR domains of plant immune receptors are NAD+-cleaving enzymes that promote cell death. Science 365, 799–803 (2019).
Wang, J. et al. Reconstitution and structure of a plant NLR resistosome conferring immunity. Science 364, eaav5870 (2019).
Sarris, P. F., Cevik, V., Dagdas, G., Jones, J. D. & Krasileva, K. V. Comparative analysis of plant immune receptor architectures uncovers host proteins likely targeted by pathogens. BMC Biol. 14, 8 (2016).
Sarris, P. F. et al. A plant immune receptor detects pathogen effectors that target WRKY transcription factors. Cell 161, 1089–1100 (2015).
Maqbool, A. et al. Structural basis of pathogen recognition by an integrated HMA domain in a plant NLR immune receptor. eLife 4, e08709 (2015).
Liu, S. et al. A telomere-to-telomere genome assembly coupled with multi-omic data provides insights into the evolution of hexaploid bread wheat. Nat. Genet. 57, 1008–1020 (2025).
Casola, C., Lawing, A. M., Betran, E. & Feschotte, C. PIF-like transposons are common in Drosophila and have been repeatedly domesticated to generate new host genes. Mol. Biol. Evol. 24, 1872–1888 (2007).
Emmons, S. W., Yesner, L., Ruan, K. S. & Katzenberg, D. Evidence for a transposon in Caenorhabditis elegans. Cell 32, 55–65 (1983).
Tang, W. et al. Transposase-derived proteins FHY3/FAR1 interact with PHYTOCHROME-INTERACTING FACTOR1 to regulate chlorophyll biosynthesis by modulating HEMB1 during deetiolation in Arabidopsis. Plant Cell 24, 1984–2000 (2012).
Mao, D. et al. The Harbinger transposon-derived gene PANDA epigenetically coordinates panicle number and grain size in rice. Plant Biotechnol. J. 20, 1154–1166 (2022).
Tang, Z. X. et al. in Wild Crop Relatives: Genomic and Breeding Resources: Cereals (ed. Kole, C.) 367–396 (Springer Berlin Heidelberg, 2011).
Liu, Y. et al. Past innovations and future possibilities in plant chromosome engineering. Plant Biotechnol. J. 23, 695–708 (2025).
Wang, C. et al. The Yr9 gene encoding a CC-NBS-LRR protein in the 1RS-1BL translocation confers wheat stripe rust resistance. Sci. China Life Sci. 68, 2804–2806 (2025).
Li, J. et al. Identification and characterization of a new stripe rust resistance gene Yr83 on rye chromosome 6R in wheat. Theor. Appl. Genet. 133, 1095–1107 (2020).
Li, G. et al. Molecular and cytogenetic dissection of stripe rust resistance gene Yr83 from rye 6R and generation of resistant germplasm in wheat breeding. Front. Plant Sci. 13, 1035784 (2022).
Duan, Y. et al. The physical location of stripe rust resistance genes on chromosome 6 of rye (Secale cereale L.) AR106BONE. Front. Plant Sci. 13, 928014 (2022).
Rabanus-Wallace, M. T. et al. Chromosome-scale genome assembly provides insights into rye biology, evolution and agronomic potential. Nat. Genet. 53, 564–573 (2021).
Sun, Y. et al. Population genomic analysis reveals domestication of cultivated rye from weedy rye. Mol. Plant 15, 552–561 (2022).
Moskal, K., Kowalik, S., Podyma, W., Łapiński, B. & Boczkowska, M. The pros and cons of rye chromatin introgression into wheat genome. Agronomy 11, 456 (2021).
Zhu, S. et al. Molecular cytogenetic analyses of two new wheat–rye 6RL translocation lines with resistance to wheat powdery mildew. Crop J. 11, 584–592 (2023).
An, D. et al. Molecular cytogenetic identification of a new wheat–rye 6R chromosome disomic addition line with powdery mildew resistance. PLoS ONE 10, e0134534 (2015).
Wen, A. et al. Genetic mapping of a major gene in triticale conferring resistance to bacterial leaf streak. Theor. Appl. Genet. 131, 649–658 (2018).
Dyda, M., Tyrka, M., Golebiowska, G., Rapacz, M. & Wedzony, M. Genetic mapping of adult-plant resistance genes to powdery mildew in triticale. J. Appl. Genet. 63, 73–86 (2022).
Karbarz, M. et al. Quantitative trait loci mapping of adult-plant resistance to powdery mildew in triticale. Ann. Appl. Biol. 177, 223–231 (2020).
Wei, F., Wing, R. A. & Wise, R. P. Genome dynamics and evolution of the Mla (powdery mildew) resistance locus in barley. Plant Cell 14, 1903–1917 (2002).
Trognitz, F. C. & Trognitz, B. R. Survey of resistance gene analogs in Solanum caripense, a relative of potato and tomato, and update on R gene genealogy. Mol. Genet. Genom. 274, 595–605 (2005).
Kim, S. et al. New reference genome sequences of hot pepper reveal the massive evolution of plant disease-resistance genes by retroduplication. Genome Biol. 18, 210 (2017).
Murray, M. & Thompson, W. F. Rapid isolation of high molecular weight plant DNA. Nucleic Acids Res. 8, 4321–4325 (1980).
Maccaferri, M. et al. Durum wheat genome highlights past domestication signatures and future improvement targets. Nat. Genet. 51, 885–895 (2019).
Kato, A. et al. Chromosome painting for plant biotechnology. Methods Mol. Biol. 701, 67–96 (2011).
Han, F., Gao, Z. & Birchler, J. A. Reactivation of an inactive centromere reveals epigenetic and structural components for centromere specification in maize. Plant Cell 21, 1929–1939 (2009).
Bolger, A., Lohse, M. & Usadel, B. J. B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics. 30, 2114–2120 (2014).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics. 29, 15–21 (2013).
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Mansfeld, B. & Grumet, R. QTLseqr: an R package for bulk segregant analysis with next-generation sequencing. Plant Genome 11, 180006 (2018).
Kim, D., Paggi, J. M., Park, C., Bennett, C. & Salzberg, S. L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat. Biotechnol. 37, 907–915 (2019).
Pertea, M. et al. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat. Biotechnol. 33, 290–295 (2015).
Frazee, A. C. et al. Ballgown bridges the gap between transcriptome assembly and expression analysis. Nat. Biotechnol. 33, 243–246 (2015).
Li, H. & Durbin, R. J. B. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics. 25, 1754–1760 (2009).
Meng, L., Li, H., Zhang, L. & Wang, J. QTL IciMapping: integrated software for genetic linkage map construction and quantitative trait locus mapping in biparental populations. Crop J. 3, 269–283 (2015).
Chen, H. et al. Firefly luciferase complementation imaging assay for protein–protein interactions in plants. Plant Physiol. 146, 368–376 (2008).
Chen, S., Zou, Y., Tong, X. & Xu, C. A tomato NBS-LRR gene Mi-9 confers heat-stable resistance to root-knot nematodes. J. Integr. Agric. 24, 2869–2875 (2025).
Bai, X. et al. Transcription factor BZR2 activates chitinase Cht20.2 transcription to confer resistance to wheat stripe rust. Plant Physiol. 187, 2749–2762 (2021).
Bolser, D. M., Staines, D. M., Perry, E. & Kersey, P. J. Ensembl Plants: integrating tools for visualizing, mining, and analyzing plant genomic data. Methods Mol. Biol. 1533, 115–140 (2017).
Yuan, S., Chan, H. C. S., Filipek, S. & Vogel, H. PyMOL and Inkscape bridge the data and the data visualization. Structure 24, 2041–2042 (2016).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
R Core Team. R: a language and environment for statistical computing. MSOR Connections 1, (2014).
Yu, G., Lam, T. T., Zhu, H. & Guan, Y. Two methods for mapping and visualizing associated data on phylogeny using Ggtree. Mol. Biol. Evol. 35, 3041–3043 (2018).
Kumar, S., Stecher, G., Li, M., Knyaz, C. & Tamura, K. MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 35, 1547–1549 (2018).
Ramírez, F., Dündar, F., Diehl, S., Grüning, B. A. & Manke, T. deepTools: a flexible platform for exploring deep-sequencing data. Nucleic Acids Res. 42, W187–191 (2014).
Acknowledgements
We thank Z. Ni (China Agricultural University) for providing the triticale germplasm, C. Xu (IGDB, CAS) for providing the MiC-4 expression plasmid, Y. Chen (IGDB, CAS) for his contributions to the protein structure prediction and model assembly, Z. Liu (IGDB, CAS) for providing field mixtures of Pst and Q. Li (Northwest A&F University) for providing Pst races. This work was supported by the National Key Research and Development Program of China (grant no. 2022YFF1003303 to F.H.) and the ‘Biological Breeding—National Science and Technology Major Project’ (grant no. 2023ZD04025 to F.H.).
Author information
Authors and Affiliations
Contributions
F.H. and C.W. designed the research. C.W. contributed to the execution and analysis of most of the experiments. S.F. created the translocation line 6R/6A and characterized its agronomic traits. C.Y. and M.W. were responsible for the genome resequencing data and transcriptome analysis. Y.C. did the genetic transformation of wheat under the guidance of X.Y. C.Z., J.Y. and Y.L. provided assistance with GISH technology, genetic marker development and protein work, respectively. Z.W. and T.W. helped characterize stripe rust resistance in the field. R.F., Y.W., and W.Y. provided constructive suggestions for this work. F.H., X.Y. and Y.L. supervised this study. F.H., Y.L. and C.W. wrote the final version of the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Plants thanks Jorge Dubcovsky, Joshua Hegarty, Yinghui Li and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–18.
Supplementary Tables (download XLSX )
Supplementary Tables 1–8.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wang, C., Fu, S., Yi, C. et al. An NLR–transposase fusion gene from rye provides broadly effective resistance to stripe rust in wheat. Nat. Plants (2026). https://doi.org/10.1038/s41477-026-02248-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41477-026-02248-1