Abstract
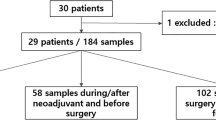
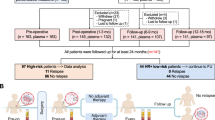
Neoadjuvant systemic therapy (NST) is a standard treatment approach for patients with early-stage breast cancer, particularly those with stage II-III disease and aggressive subtypes such as HER2-positive and triple-negative breast cancer. While NST improves surgical outcomes and provides prognostic information, accurately assessing preoperative treatment response remains a clinical challenge. Circulating tumor DNA (ctDNA) has emerged as a promising non-invasive biomarker for monitoring disease dynamics and guiding therapeutic decisions. In this study, we aimed to evaluate whether ctDNA analysis in patients with stage II-III breast cancer (n = 20) could serve as a surrogate for invasive biopsies in molecular profiling and as a tool for monitoring response to NST. At baseline, ctDNA was detectable in the majority of patients by droplet digital (dd)PCR (15/18, 83%) and all patients with longitudinal follow-up had ctDNA clearance after NST (13/13; 100%). A positive correlation was observed between the allele fraction in ctDNA, histologic grade and molecular subtype, suggesting that ctDNA levels may be influenced by tumor biology. None of the three patients with undetectable baseline ctDNA had distant relapse, regardless of whether they achieved pathologic complete response (pCR), compared to 5/15 (33%) with detectable baseline ctDNA. These findings suggest that ctDNA assessment at baseline may provide additional prognostic information to define the risk of patients after NST. While ctDNA shows promise in capturing tumor burden and biological characteristics, its role in predicting pCR and long-term outcomes requires further investigation.
Similar content being viewed by others
Data availability
The assembled somatic mutational data of the primary tumor, plasma, residual disease as well as distant/ local relapse from the entire cohort are accessible through cBioPortal at https://www.cbioportal.org/study/summary?id=breast_msk_cfDNA_2026. All codes used in this manuscript are available for academic use on GitHub at https://github.com/ndbrown6/MSK-Early-Breast/.
References
Korde, L. A. et al. Neoadjuvant chemotherapy, endocrine therapy, and targeted therapy for breast cancer: ASCO Guideline. J. Clin. Oncol. 39, 1485–1505 (2021).
Curigliano, G. et al. Understanding breast cancer complexity to improve patient outcomes: The St Gallen International Consensus Conference for the Primary Therapy of Individuals with Early Breast Cancer 2023. Ann. Oncol. 34, 970–986 (2023).
Loibl, S. et al. Early breast cancer: ESMO clinical practice guideline for diagnosis, treatment and follow-up. Ann. Oncol. 35, 159–182 (2024).
Houssami, N., Macaskill, P., von Minckwitz, G., Marinovich, M. L. & Mamounas, E. Meta-analysis of the association of breast cancer subtype and pathologic complete response to neoadjuvant chemotherapy. Eur. J. Cancer 48, 3342–3354 (2012).
Spring, L. M. et al. Pathologic complete response after neoadjuvant chemotherapy and impact on breast cancer recurrence and survival: a comprehensive meta-analysis. Clin. Cancer Res. 26, 2838–2848 (2020).
Parsons, H. A. et al. Sensitive detection of minimal residual disease in patients treated for early-stage breast cancer. Clin. Cancer Res. 26, 2556–2564 (2020).
Cortazar, P. et al. Pathological complete response and long-term clinical benefit in breast cancer: the CTNeoBC pooled analysis. Lancet 384, 164–172 (2014).
Wan, J. C. M. et al. Liquid biopsies for residual disease and recurrence. Med. 2, 1292–1313 (2021).
Wan, J. C. M. et al. Liquid biopsies come of age: towards implementation of circulating tumour DNA. Nat. Rev. Cancer 17, 223–238 (2017).
Forshew, T. et al. Noninvasive identification and monitoring of cancer mutations by targeted deep sequencing of plasma DNA. Sci. Transl. Med 4, 136ra168 (2012).
Bettegowda, C. et al. Detection of circulating tumor DNA in early- and late-stage human malignancies. Sci. Transl. Med. 6, 224ra224 (2014).
Razavi, P. et al. High-intensity sequencing reveals the sources of plasma circulating cell-free DNA variants. Nat. Med. 25, 1928–1937 (2019).
Pascual, J. et al. ESMO recommendations on the use of circulating tumour DNA assays for patients with cancer: a report from the ESMO Precision Medicine Working Group. Ann. Oncol. 33, 750–768 (2022).
Turner, N. C. et al. Circulating tumour DNA analysis to direct therapy in advanced breast cancer (plasmaMATCH): a multicentre, multicohort, phase 2a, platform trial. Lancet Oncol. 21, 1296–1308 (2020).
Fribbens, C. et al. Plasma ESR1 mutations and the treatment of estrogen receptor-positive advanced breast cancer. J. Clin. Oncol. 34, 2961–2968 (2016).
Bidard, F. C. et al. Elacestrant (oral selective estrogen receptor degrader) versus standard endocrine therapy for estrogen receptor-positive, human epidermal growth factor receptor 2-negative advanced breast cancer: results from the randomized phase III EMERALD trial. J. Clin. Oncol. 40, 3246–3256 (2022).
Coombes, R. C. et al. Personalized detection of circulating tumor DNA antedates breast cancer metastatic recurrence. Clin. Cancer Res. 25, 4255–4263 (2019).
Garcia-Murillas, I. et al. Assessment of molecular relapse detection in early-stage breast cancer. JAMA Oncol. 5, 1473–1478 (2019).
Garcia-Murillas, I. et al. Mutation tracking in circulating tumor DNA predicts relapse in early breast cancer. Sci. Transl. Med. 7, 302ra133 (2015).
Radovich, M. et al. Association of circulating tumor DNA and circulating tumor cells after neoadjuvant chemotherapy with disease recurrence in patients with triple-negative breast cancer: preplanned secondary analysis of the BRE12-158 randomized clinical trial. JAMA Oncol. 6, 1410–1415 (2020).
Parsons, H. A. et al. Circulating tumor DNA association with residual cancer burden after neoadjuvant chemotherapy in triple-negative breast cancer in TBCRC 030. Ann. Oncol. 34, 899–906 (2023).
Magbanua, M. J. M. et al. Circulating tumor DNA in neoadjuvant-treated breast cancer reflects response and survival. Ann. Oncol. 32, 229–239 (2021).
Agostinetto, E., Nader-Marta, G. & Ignatiadis, M. Circulating tumor DNA in breast cancer: a biomarker for patient selection. Curr. Opin. Oncol. 35, 426–435 (2023).
Nader-Marta, G. et al. Circulating tumor DNA for predicting recurrence in patients with operable breast cancer: a systematic review and meta-analysis. ESMO Open 9, 102390 (2024).
Thierry, A. R., El Messaoudi, S., Gahan, P. B., Anker, P. & Stroun, M. Origins, structures, and functions of circulating DNA in oncology. Cancer Metastasis Rev. 35, 347–376 (2016).
Mouliere, F. et al. Enhanced detection of circulating tumor DNA by fragment size analysis. Sci. Transl. Med. 10, eaat4921 (2018).
Cailleux, F. et al. Circulating tumor DNA after neoadjuvant chemotherapy in breast cancer is associated with disease relapse. JCO Precis. Oncol. 6, e2200148 (2022).
Abbosh, C. et al. Phylogenetic ctDNA analysis depicts early-stage lung cancer evolution. Nature 545, 446–451 (2017).
Magbanua, M. J. M. et al. Clinical significance and biology of circulating tumor DNA in high-risk early-stage HER2-negative breast cancer receiving neoadjuvant chemotherapy. Cancer Cell 41, 1091–1102.e1094 (2023).
Coakley, M. et al. Comparison of circulating tumor DNA assays for molecular residual disease detection in early-stage triple-negative breast cancer. Clin. Cancer Res. 30, 895–903 (2024).
Zhang, D. et al. Role of circulating tumor DNA in early-stage triple-negative breast cancer: a systematic review and meta-analysis. Breast Cancer Res. 27, 38 (2025).
Santonja, A. et al. Comparison of tumor-informed and tumor-naive sequencing assays for ctDNA detection in breast cancer. EMBO Mol. Med. 15, e16505 (2023).
Elliott, M. J. et al. Longitudinal evaluation of circulating tumor DNA in patients undergoing neoadjuvant therapy for early breast cancer using a tumor-informed assay. Nat. Commun. 16, 1837 (2025).
Mittendorf, E. A. et al. Peri-operative atezolizumab in early-stage triple-negative breast cancer: final results and ctDNA analyses from the randomized phase 3 IMpassion031 trial. Nat. Med. 31, 2397–2404 (2025).
Black, J. R. M. et al. Ultrasensitive ctDNA detection for preoperative disease stratification in early-stage lung adenocarcinoma. Nat. Med. 31, 70–76 (2025).
Kurtz, D. M. et al. Enhanced detection of minimal residual disease by targeted sequencing of phased variants in circulating tumor DNA. Nat. Biotechnol. 39, 1537–1547 (2021).
Cabel, L. et al. 293P Ultra-sensitive ctDNA detection and monitoring in early breast cancer using PhasED-Seq. Ann. Oncol. 35, S338 (2024).
Zviran, A. et al. Genome-wide cell-free DNA mutational integration enables ultra-sensitive cancer monitoring. Nat. Med. 26, 1114–1124 (2020).
Elliott, M. J. et al. Ultrasensitive detection and monitoring of circulating tumor DNA using structural variants in early-stage breast cancer. Clin. Cancer Res. 31, 1520–1532 (2025).
Janni, W. et al. Detection of minimal residual disease and prediction of recurrence in breast cancer using a plasma-only circulating tumor DNA assay. ESMO Open 10, 104296 (2025).
Turner, N. C. et al. Results of the c-TRAK TN trial: a clinical trial utilising ctDNA mutation tracking to detect molecular residual disease and trigger intervention in patients with moderate- and high-risk early-stage triple-negative breast cancer. Ann. Oncol. 34, 200–211 (2023).
Allison, K. H. et al. Estrogen and progesterone receptor testing in breast cancer: ASCO/CAP guideline update. J. Clin. Oncol. 38, 1346–1366 (2020).
Wolff, A. C. et al. Human epidermal growth factor receptor 2 testing in breast cancer: American Society of Clinical Oncology/College of American Pathologists Clinical Practice Guideline Focused Update. J. Clin. Oncol. 36, 2105–2122 (2018).
Symmans, W. F. et al. Measurement of residual breast cancer burden to predict survival after neoadjuvant chemotherapy. J. Clin. Oncol. 25, 4414–4422 (2007).
Yau, C. et al. Residual cancer burden after neoadjuvant chemotherapy and long-term survival outcomes in breast cancer: a multicentre pooled analysis of 5161 patients. Lancet Oncol. 23, 149–160 (2022).
Ashley, C. W. et al. High-sensitivity mutation analysis of cell-free DNA for disease monitoring in endometrial cancer. Clin. Cancer Res. 29, 410–421 (2023).
Zehir, A. et al. Mutational landscape of metastatic cancer revealed from prospective clinical sequencing of 10,000 patients. Nat. Med. 23, 703–713 (2017).
Nguyen, B. et al. Genomic characterization of metastatic patterns from prospective clinical sequencing of 25,000 patients. Cell 185, 563–575.e511 (2022).
Schwartz, C. J. et al. Morphologic and genomic characteristics of breast cancers occurring in individuals with lynch syndrome. Clin. Cancer Res. 28, 404–413 (2022).
Selenica, P. et al. APOBEC mutagenesis, kataegis, chromothripsis in EGFR-mutant osimertinib-resistant lung adenocarcinomas. Ann. Oncol. 33, 1284–1295 (2022).
Mandelker, D. et al. Genomic profiling reveals germline predisposition and homologous recombination deficiency in pancreatic acinar cell carcinoma. J. Clin. Oncol. 41, 5151–5162 (2023).
Gupta, A. et al. APOBEC3 mutagenesis drives therapy resistance in breast cancer. Nat. Genet. 57, 1452–1462 (2025).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Weigelt, B. et al. Diverse BRCA1 and BRCA2 reversion mutations in circulating cell-free DNA of therapy-resistant breast or ovarian cancer. Clin. Cancer Res. 23, 6708–6720 (2017).
Blanco-Heredia, J. et al. Converging and evolving immuno-genomic routes toward immune escape in breast cancer. Nat. Commun. 15, 1302 (2024).
Acknowledgements
Research reported in this publication was supported by a Cancer Center Support Grant of the NIH/NCI (P30 CA008748), the Marie-Josée and Henry R. Kravis Center for Molecular Oncology, by FDA 1U01FD007909-01A1 (F.P.), NIH/NCI MSK SPORE P50 CA247749-01 (F.P., BW), NIH R01 CA245069 (S.C.) and Breast Cancer Research Foundation (B.W., F.P., S.C.) grants.
Author information
Authors and Affiliations
Contributions
Conceptualization, T.A.K., J.S.R.-F. and B.W.; methodology, T.A.K., B.W., J.S.R.-F. and D.B.S.; formal analysis, F.P., S.H.K., D.N.B. and A.M.; investigation, T.A.K., G.P. and S.C.; data curation, A.M., S.H.K., F.P., T.B., B.W., and D.N. B.; writing, review and editing B.W., S.H.K., F.P., D.N.B. and A.M.; supervision, D.N.B. and T.A.K. All authors have read and agreed to the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
A.M. reports personal fees from Menarini/Stemline, Roche, Lilly and AstraZeneca, outside the submitted work; reports travel support from Menarini/Stemline, AstraZeneca and Daiichi-Sankyo, outside the submitted work; and is supported by the ESMO José Baselga Fellowship for Clinician Scientists sponsored by AstraZeneca, outside the submitted work. F.P. reports membership on the Scientific Advisory Board of MultiplexDx and has received consulting fees and served on advisory boards for AstraZeneca. D.B.S. has consulted/received honoraria from Pfizer, Fog Pharma, PaigeAI, BridgeBio, Scorpion Therapeutics, FORE Therapeutics, Function Oncology, Pyramid, and Elsie Biotechnologies, Inc., Meliora Therapeutics, Inc. B.W. reports research funding from Repare Therapeutics and SAGA Diagnostics, paid to the institution. J.S.R.-F. is an employee of AstraZeneca and owns AstraZeneca stocks. Prior Conflicts of Interest in the last 2 years include the receipt of personal fees for the following activities: Board Membership at Grupo Oncoclinicas, consultant for Goldman Sachs Merchant Banking, consultant for Bain Capital, consultant for and SAB member of Paige.ai, consultant for and SAB member of Repare Therapeutics, Consultant of SAGA Diagnostics, Consultant of Personalis, and consultant at Multiplex Dx.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Marra, A., Kim, S.H., Pareja, F. et al. Tracking response to neoadjuvant systemic therapy through circulating tumor DNA analysis in breast cancer. npj Breast Cancer (2026). https://doi.org/10.1038/s41523-026-00921-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41523-026-00921-4