Abstract
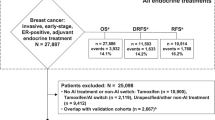
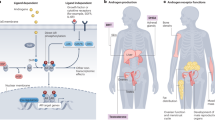
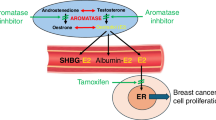
Aromatase inhibitors (AI) are standard therapy for hormone receptor–positive breast cancer in post-menopausal women, yet recurrence remains common. Our previous work suggests that an androgen‑dominated steroid environment may drive AI resistance. Although most androgen research has focused on classical genomic pathways in reproductive tissues, interest is growing in their non‑reproductive functions. In particular, the role of cytoplasmic AR has recently gained attention, and its connection to metabolic modulation remains largely unexplored in the context of breast cancer. Cytoplasmic AR was evaluated in a breast cancer microarray (n = 875), validated in an independent cohort (n = 30), and examined in metastatic biopsies (n = 12). LC‑MS/MS identified AR‑interacting proteins in AI‑resistant cells exposed to adrenal androgens, confirmed by co‑immunoprecipitation and imaging. High cytoplasmic AR predicted poor survival in post‑menopausal patients, especially luminal B cancers (p = 0.0085). AI‑resistant models showed diffuse AR localisation throughout the cytoplasm and nucleus accompanied by increased mitochondrial mass and membrane potential, and elevated oxidative phosphorylation and glycolysis. Label‑free mass spectrometry identified G3BP1, SLIRP and IGFBP5 as AR interactors linked to stress response, metabolic adaptation and ERα repression. The findings of this study highlight the prognostic potential of cytoplasmic AR immunoreactivity in specific breast cancer subtypes and uncover novel cytoplasmic AR protein interactions that may mediate metabolic adaptations during the development of endocrine-resistance.
Similar content being viewed by others
Data availability
All data supporting the findings of this study are available within the paper and its Supplementary Information.
References
Houghton, L. C. et al. The Steroid Metabolome and Breast Cancer Risk in Women with a Family History of Breast Cancer: The Novel Role of Adrenal Androgens and Glucocorticoids. Cancer Epidemiol. Biomark. Prev. 30, 89–96 (2021).
Trinh, L. et al. Pre-diagnostic serum metabolome and breast cancer risk: a nested case-control study. Breast Cancer Res 27, 156 (2025).
Pfeiler, G. et al. Impact of body mass index on estradiol depletion by aromatase inhibitors in postmenopausal women with early breast cancer. Br. J. Cancer 109, 1522–1527 (2013).
Hanamura, T. & Hayashi, S. -i Overcoming aromatase inhibitor resistance in breast cancer: possible mechanisms and clinical applications. Breast Cancer 25, 379–391 (2018).
Rechoum, Y. et al. AR collaborates with ERα in aromatase inhibitor-resistant breast cancer. Breast Cancer Res Treat. 147, 473–485 (2014).
Fujii, R. et al. Increased androgen receptor activity and cell proliferation in aromatase inhibitor-resistant breast carcinoma. J. Steroid Biochem. Mol. Biol. 144, 513–522 (2014).
Creevey, L. et al. Altered Steroid Milieu in AI-Resistant Breast Cancer Facilitates AR Mediated Gene-Expression Associated with Poor Response to Therapy. Mol. Cancer Therapeutics 18, 1731 (2019).
Bleach, R. et al. Steroid Ligands, the Forgotten Triggers of Nuclear Receptor Action; Implications for Acquired Resistance to Endocrine Therapy. Clin. Cancer Res. 27, 3980–3989 (2021).
Bleach, R. & McIlroy, M. The Divergent Function of Androgen Receptor in Breast Cancer; Analysis of Steroid Mediators and Tumor Intracrinology. Front Endocrinol. (Lausanne) 9, 594 (2018).
Davey, R. A. & Grossmann, M. Androgen Receptor Structure, Function and Biology: From Bench to Bedside. Clin. Biochem Rev. 37, 3–15 (2016).
Chia, K. et al. Non-canonical AR activity facilitates endocrine resistance in breast cancer. Endocr. Relat. Cancer 26, 251–264 (2019).
Agbana, S. & McIlroy, M. Extra-nuclear and cytoplasmic steroid receptor signalling in hormone dependent cancers. J. Steroid Biochem Mol. Biol. 243, 106559 (2024).
Safi, R. et al. Androgen receptor monomers and dimers regulate opposing biological processes in prostate cancer cells. Nat. Commun. 15, 7675 (2024).
Ricciardelli, C. et al. The Magnitude of Androgen Receptor Positivity in Breast Cancer Is Critical for Reliable Prediction of Disease Outcome. Clin. Cancer Res. 24, 2328 (2018).
Kolyvas, E. A., Caldas, C., Kelly, K. & Ahmad, S. S. Androgen receptor function and targeted therapeutics across breast cancer subtypes. Breast Cancer Res. 24, 79 (2022).
Gonzalez, M. W. & Kann, M. G. Chapter 4: Protein interactions and disease. PLoS Comput Biol. 8, e1002819–e1002819 (2012).
Hsiao, J. J., Smits, M. M., Ng, B. H., Lee, J. & Wright, M. E. Discovery Proteomics Identifies a Molecular Link between the Coatomer Protein Complex I and Androgen Receptor-dependent Transcription. J. Biol. Chem. 291, 18818–18842 (2016).
Stelloo, S. et al. Endogenous androgen receptor proteomic profiling reveals genomic subcomplex involved in prostate tumorigenesis. Oncogene 37, 313–322 (2018).
Dahiya, U. R. & Heemers, H. V. Analyzing the Androgen Receptor Interactome in Prostate Cancer: Implications for Therapeutic Intervention. Cells 11, https://doi.org/10.3390/cells11060936 (2022).
Hosseinzadeh, L. et al. The androgen receptor interacts with GATA3 to transcriptionally regulate a luminal epithelial cell phenotype in breast cancer. Genome Biol. 25, 44 (2024).
Jasavala, R. et al. Identification of putative androgen receptor interaction protein modules: cytoskeleton and endosomes modulate androgen receptor signaling in prostate cancer cells. Mol. Cell Proteom. 6, 252–271 (2007).
Mohammed, H. et al. Endogenous purification reveals GREB1 as a key estrogen receptor regulatory factor. Cell Rep. 3, 342–349 (2013).
KeeMing, C. et al. Non-canonical AR activity facilitates endocrine resistance in breast cancer. Endocr. -Relat. Cancer 26, 251–264 (2019).
Efstathiou, E. et al. Molecular characterization of enzalutamide-treated bone metastatic castration-resistant prostate cancer. Eur. Urol. 67, 53–60 (2015).
Bajpai, P., Koc, E., Sonpavde, G., Singh, R. & Singh, K. K. Mitochondrial localization, import, and mitochondrial function of the androgen receptor. J. Biol. Chem. 294, 6621–6634 (2019).
Barton, V. N. et al. Androgen Receptor Supports an Anchorage-Independent, Cancer Stem Cell-like Population in Triple-Negative Breast Cancer. Cancer Res. 77, 3455–3466 (2017).
de Kruijff, I. E. et al. Androgen receptor expression in circulating tumor cells of patients with metastatic breast cancer. Int J. Cancer 145, 1083–1089 (2019).
Shaw, F. L. et al. A detailed mammosphere assay protocol for the quantification of breast stem cell activity. J. Mammary Gland Biol. Neoplasia 17, 111–117 (2012).
Gonthier, K., Poluri, R. T. K. & Audet-Walsh, É. Functional genomic studies reveal the androgen receptor as a master regulator of cellular energy metabolism in prostate cancer. J. Steroid Biochem. Mol. Biol. 191, 105367 (2019).
Wustmann, N. et al. Molecular analysis of androgen receptor splice variant AR-V3 reveals eminent ambiguity regarding activity and clinical utility. Cancer Cell Int 25, 316 (2025).
ten Have, S., Boulon, S., Ahmad, Y. & Lamond, A. I. Mass spectrometry-based immuno-precipitation. Proteom. - Use ’s. guide Proteom. 11, 1153–1159 (2011).
Paltoglou, S. et al. Novel Androgen Receptor Coregulator GRHL2 Exerts Both Oncogenic and Antimetastatic Functions in Prostate Cancer. Cancer Res. 77, 3417–3430 (2017).
Mellacheruvu, D. et al. The CRAPome: a contaminant repository for affinity purification-mass spectrometry data. Nat. Methods 10, 730–736 (2013).
De Silva, D. et al. Interaction between androgen receptor and coregulator SLIRP is regulated by Ack1 tyrosine kinase and androgen. Sci. Rep. 9, 18637 (2019).
Akkiprik, M., Hu, L., Sahin, A., Hao, X. & Zhang, W. The subcellular localization of IGFBP5 affects its cell growth and migration functions in breast cancer. BMC Cancer 9, 103 (2009).
Williams, M. M. et al. Steroid Hormone Receptor and Infiltrating Immune Cell Status Reveals Therapeutic Vulnerabilities of ESR1-Mutant Breast Cancer. Cancer Res 81, 732–746 (2021).
Riva, C. et al. Immunohistochemical study of androgen receptors in breast carcinoma. Evid. their Freq. Expr. lobular carcinoma Virchows Arch. 447, 695–700 (2005).
Lasley, B. L. et al. Circulating dehydroepiandrosterone sulfate levels in women who underwent bilateral salpingo-oophorectomy during the menopausal transition. Menopause 18, 494–498 (2011).
Nanba, A. T. et al. 11-Oxygenated C19 Steroids Do Not Decline With Age in Women. J. Clin. Endocrinol. Metab. 104, 2615–2622 (2019).
Turcu, A. F. et al. 11-Oxygenated Androgens Are Biomarkers of Adrenal Volume and Testicular Adrenal Rest Tumors in 21-Hydroxylase Deficiency. J. Clin. Endocrinol. Metab. 102, 2701–2710 (2017).
O’Reilly, M. W. et al. Hyperandrogenemia predicts metabolic phenotype in polycystic ovary syndrome: the utility of serum androstenedione. J. Clin. Endocrinol. Metab. 99, 1027–1036 (2014).
Hong, S. H. et al. Non-alcoholic fatty liver disease is associated with hyperandrogenism in women with polycystic ovary syndrome. Sci. Rep. 13, 13397 (2023).
Cerdas Perez, S. Menopause and diabetes. Climacteric 26, 216–221 (2023).
Navarro, G. et al. Extranuclear Actions of the Androgen Receptor Enhance Glucose-Stimulated Insulin Secretion in the Male. Cell Metab. 23, 837–851 (2016).
Xu, W. et al. Architecture of androgen receptor pathways amplifying glucagon-like peptide-1 insulinotropic action in male pancreatic beta cells. Cell Rep. 42, 112529 (2023).
Andrisse, S. et al. Hepatocyte androgen receptor in females mediates androgen-induced hepatocellular glucose mishandling and systemic insulin resistance. bioRxiv, 2021.2006.2009.447759 https://doi.org/10.1101/2021.06.09.447759 (2021).
Schiffer, L., Kempegowda, P., Arlt, W. & O’Reilly, M. W. MECHANISMS IN ENDOCRINOLOGY: The sexually dimorphic role of androgens in human metabolic disease. Eur. J. Endocrinol. 177, R125–R143 (2017).
O’Reilly, M. W. et al. 11-Oxygenated C19 Steroids Are the Predominant Androgens in Polycystic Ovary Syndrome. J. Clin. Endocrinol. Metab. 102, 840–848 (2017).
Yin, L., Luo, M., Wang, R., Ye, J. & Wang, X. Mitochondria in Sex Hormone-Induced Disorder of Energy Metabolism in Males and Females. Front Endocrinol. (Lausanne) 12, 749451 (2021).
Horwath, O., Moberg, M., Hirschberg, A. L., Ekblom, B. & Apro, W. Molecular Regulators of Muscle Mass and Mitochondrial Remodeling Are Not Influenced by Testosterone Administration in Young Women. Front Endocrinol. (Lausanne) 13, 874748 (2022).
Yin, L., Qi, S. & Zhu, Z. Advances in mitochondria-centered mechanism behind the roles of androgens and androgen receptor in the regulation of glucose and lipid metabolism. Front Endocrinol. (Lausanne) 14, 1267170 (2023).
Yi, S. et al. Melatonin ameliorates excessive PINK1/Parkin-mediated mitophagy by enhancing SIRT1 expression in granulosa cells of PCOS. Am. J. Physiol. Endocrinol. Metab. 319, E91–E101 (2020).
Rossetti, M. L. & Gordon, B. S. The role of androgens in the regulation of muscle oxidative capacity following aerobic exercise training. Appl Physiol. Nutr. Metab. 42, 1001–1007 (2017).
Abate, M. et al. Mitochondria as playmakers of apoptosis, autophagy and senescence. Semin Cell Dev. Biol. 98, 139–153 (2020).
Ahmad, I. & Newell-Fugate, A. E. Role of androgens and androgen receptor in control of mitochondrial function. Am. J. Physiol. Cell Physiol. 323, C835–C846 (2022).
Atawia, I. M. et al. Inhibition of Wnt/beta-catenin pathway overcomes therapeutic resistance to abiraterone in castration-resistant prostate cancer. Mol. Carcinog. 62, 1312–1324 (2023).
Wang, G., Wang, J. & Sadar, M. D. Crosstalk between the androgen receptor and beta-catenin in castrate-resistant prostate cancer. Cancer Res 68, 9918–9927 (2008).
Hatchell, E. C. et al. SLIRP, a small SRA binding protein, is a nuclear receptor corepressor. Mol. Cell 22, 657–668 (2006).
Tourriere, H. et al. RasGAP-associated endoribonuclease G3Bp: selective RNA degradation and phosphorylation-dependent localization. Mol. Cell Biol. 21, 7747–7760 (2001).
Buchanan, G. et al. Decreased androgen receptor levels and receptor function in breast cancer contribute to the failure of response to medroxyprogesterone acetate. Cancer Res. 65, 8487–8496 (2005).
Hickey, T. E. et al. The androgen receptor is a tumor suppressor in estrogen receptor-positive breast cancer. Nat. Med. 27, 310–320 (2021).
Amen, T. & Kaganovich, D. Stress granules inhibit fatty acid oxidation by modulating mitochondrial permeability. Cell Rep. 35, 109237 (2021).
Somasekharan, S. P. et al. G3BP1-linked mRNA partitioning supports selective protein synthesis in response to oxidative stress. Nucleic Acids Res. 48, 6855–6873 (2020).
Song, C. et al. IGFBP5 promotes diabetic kidney disease progression by enhancing PFKFB3-mediated endothelial glycolysis. Cell Death Dis. 13, 340 (2022).
Alessio, N. et al. IGFBP5 is released by senescent cells and is internalized by healthy cells, promoting their senescence through interaction with retinoic receptors. Cell Commun. Signal 22, 122 (2024).
Feeley, L. P., Mulligan, A. M., Pinnaduwage, D., Bull, S. B. & Andrulis, I. L. Distinguishing luminal breast cancer subtypes by Ki67, progesterone receptor or TP53 status provides prognostic information. Mod. Pathol. 27, 554–561 (2014).
Shevchenko, A., Wilm, M., Vorm, O. & Mann, M. Mass Spectrometric Sequencing of Proteins from Silver-Stained Polyacrylamide Gels. Anal. Chem. 68, 850–858 (1996).
Chen, E. Y. et al. Enrichr: interactive and collaborative HTML5 gene list enrichment analysis tool. BMC Bioinforma. 14, 128 (2013).
Kuleshov, M. V. et al. Enrichr: a comprehensive gene set enrichment analysis web server 2016 update. Nucleic Acids Res 44, W90–W97 (2016).
Xie, Z. et al. Gene Set Knowledge Discovery with Enrichr. Curr. Protoc. 1, e90 (2021).
Acknowledgements
M. Mcllroy is supported by Beaumont Hospital Cancer Research and Development Trust, project number: 2077.We owe a huge debt of gratitude to all patients who have contributed to this study, without whom translational research of this nature would not be possible.
Author information
Authors and Affiliations
Contributions
M.McI devised the project, the main conceptual ideas and proof outline. R.B., E.B., and J.Q. carried out the experiments, performed the analysis, drafted the manuscript and designed the figures. S.S. performed MS experiment and analysis, K.S., L.Y. performed digital pathology analysis. M.O., L.G., and S.A. performed experiments, data analysis and generated graphs. C.L.C., H.H.M., J.K.R., N.S., K.S., and A.C.V. evaluated primary and metastatic samples by IHC, analysed and generated graphs. M.McI, J.P., and M.O.R. supervised the project. All authors provided critical feedback and contributed to writing, reviewing, and editing the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Bleach, R., Bozkurt, E., Xin, J. et al. Androgen receptor localisation and protein interactions provide insight into steroid mediated metabolic shifts in endocrine resistant breast cancer. npj Breast Cancer (2026). https://doi.org/10.1038/s41523-026-00924-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41523-026-00924-1