Abstract
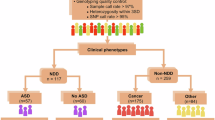
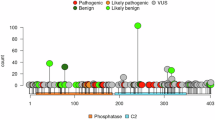
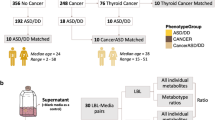
PTEN hamartoma tumor syndrome (PHTS), caused by germline PTEN variants, exhibits marked phenotypic heterogeneity, most notably cancer, neurodevelopmental disorders (NDD), or both. The basis for this divergence, even among carriers of identical PTEN variants, remains poorly defined. We performed whole-genome sequencing of 599 individuals with PHTS and family members, complemented by analyses of PTEN variant carriers from the All of Us Research Program. Analyses included both targeted evaluation of genes previously implicated in cancer and NDD and agnostic genome-wide single-variant and rare-variant burden testing. The analytic cohort comprised 543 PHTS probands, including individuals with NDD (n = 171), cancer (n = 221), both phenotypes (n = 21), or neither (n = 130) at the time of enrollment. Pathogenic or likely pathogenic variants in cancer-associated genes were identified in 37 (6.8%), most frequently in MITF, DICER1, and BRCA2, while 43 (7.9%) harbored variants in NDD-related genes, including DHCR7, POLG, and ARSA. Such secondary variants were less common in PTEN variant carriers in All of Us. Genome-wide analyses identified candidate modifier loci functionally linked to PTEN, including in ZNF713, TPTE2P1, and PDPK1. These findings demonstrate that PHTS phenotypes are shaped by complex gene–gene interactions beyond PTEN alone, informing mechanisms underlying the cancer–NDD dichotomy and advancing precision risk stratification.
Similar content being viewed by others
Data availability
The data generated in this study will be publicly available in the database of Genotypes and Phenotypes (dbGaP) upon acceptance. All of Us (AoU) data analyses were performed using the AoU Controlled Tier Dataset v8 dataset and executed using the AoU Research Workbench. All other raw data are available from the corresponding author on reasonable request.
Code availability
All software versions are indicated within the Methods. All custom scripts related to this study are available from the corresponding author upon reasonable request. All other bioinformatics and statistical tools are publicly available and are described within the manuscript text.
References
FitzGerald, G. et al. The future of humans as model organisms. Science 361, 552–553 (2018).
Topol, E. J. Individualized medicine from prewomb to tomb. Cell 157, 241–253 (2014).
Eichler, E. E. Genetic Variation, Comparative Genomics, and the Diagnosis of Disease. N. Engl. J. Med. 381, 64–74 (2019).
Tan, M. H. et al. Lifetime cancer risks in individuals with germline PTEN mutations. Clin. Cancer Res. 18, 400–407 (2012).
Hendricks, L. A. J. et al. Cancer risks by sex and variant type in PTEN Hamartoma Tumor Syndrome. J. Natl. Cancer Inst. https://doi.org/10.1093/jnci/djac188 (2022).
Yehia, L. et al. Longitudinal analysis of cancer risk in children and adults with germline PTEN variants. JAMA Netw. Open 6, e239705 (2023).
Yehia, L., Ngeow, J. & Eng, C. PTEN-opathies: from biological insights to evidence-based precision medicine. J. Clin. Invest 129, 452–464 (2019).
Satterstrom, F. K. et al. Large-scale exome sequencing study implicates both developmental and functional changes in the neurobiology of autism. Cell 180, 568–584.e523 (2020).
Trost, B. et al. Genomic architecture of autism from comprehensive whole-genome sequence annotation. Cell 185, 4409–4427.e4418 (2022).
Ni, Y. et al. Germline SDHx variants modify breast and thyroid cancer risks in Cowden and Cowden-like syndrome via FAD/NAD-dependant destabilization of p53. Hum. Mol. Genet 21, 300–310 (2012).
Yehia, L. et al. Copy number variation and clinical outcomes in patients with germline PTEN mutations. JAMA Netw. Open 3, e1920415 (2020).
Wei, R., Yehia, L., Ni, Y. & Eng, C. The mitochondrial genome as a modifier of autism versus cancer phenotypes in PTEN hamartoma tumor syndrome. HGG Adv. 4, 100199 (2023).
Kim, A. Y., Yehia, L. & Eng, C. Genomic diversity in functionally relevant genes modifies neurodevelopmental versus neoplastic risks in individuals with germline PTEN variants. NPJ Genom. Med 10, 43 (2025).
Yehia, L. et al. Distinct alterations in tricarboxylic acid cycle metabolites associate with cancer and autism phenotypes in Cowden syndrome and Bannayan–Riley–Ruvalcaba syndrome. Am. J. Hum. Genet 105, 813–821 (2019).
Yehia, L., Ni, Y., Sadler, T., Frazier, T. W. & Eng, C. Distinct metabolic profiles associated with autism spectrum disorder versus cancer in individuals with germline PTEN mutations. NPJ Genom. Med 7, 16 (2022).
Chen, H. et al. Control for population structure and relatedness for binary traits in genetic association studies via logistic mixed models. Am. J. Hum. Genet 98, 653–666 (2016).
Yehia, L., Keel, E. & Eng, C. The clinical spectrum of PTEN mutations. Annu Rev. Med 71, 103–116 (2020).
Valle, L., Rodriguez-Lopez, R., Robledo, M., Benitez, J. & Urioste, M. Concurrence of germline mutations in the APC and PTEN genes in a colonic polyposis family member. J. Clin. Oncol. 22, 2252–2253 (2004).
Zbuk, K. M. et al. Germline mutations in PTEN and SDHC in a woman with epithelial thyroid cancer and carotid paraganglioma. Nat. Clin. Pract. Oncol. 4, 608–612 (2007).
Plon, S. E. et al. Multiple tumors in a child with germ-line mutations in TP53 and PTEN. N. Engl. J. Med 359, 537–539 (2008).
Whitworth, J. et al. Multilocus inherited neoplasia alleles syndrome: a case series and review. JAMA Oncol. 2, 373–379 (2016).
Stradella, A. et al. Does multilocus inherited neoplasia alleles syndrome have severe clinical expression?. J. Med Genet 56, 521–525 (2019).
Paillerets, B. B., Lesueur, F. & Bertolotto, C. A germline oncogenic MITF mutation and tumor susceptibility. Eur. J. Cell Biol. 93, 71–75 (2014).
Bertolotto, C. et al. A SUMOylation-defective MITF germline mutation predisposes to melanoma and renal carcinoma. Nature 480, 94–98 (2011).
Yokoyama, S. et al. A novel recurrent mutation in MITF predisposes to familial and sporadic melanoma. Nature 480, 99–103 (2011).
Guhan, S. M. et al. Cancer risks associated with the germline MITF(E318K) variant. Sci. Rep. 10, 17051 (2020).
Fischer-Zirnsak, B. et al. Haploinsufficiency of the notch ligand DLL1 causes variable neurodevelopmental disorders. Am. J. Hum. Genet 105, 631–639 (2019).
Metsu, S. et al. A CGG-repeat expansion mutation in ZNF713 causes FRA7A: association with autistic spectrum disorder in two families. Hum. Mutat. 35, 1295–1300 (2014).
Walker, S. M., Downes, C. P. & Leslie, N. R. TPIP: a novel phosphoinositide 3-phosphatase. Biochem J. 360, 277–283 (2001).
Cordon-Barris, L. et al. Mutation of the 3-phosphoinositide-dependent protein kinase 1 (PDK1) substrate-docking site in the developing brain causes microcephaly with abnormal brain morphogenesis independently of Akt, leading to impaired cognition and disruptive behaviors. Mol. Cell Biol. 36, 2967–2982 (2016).
Mester, J. L. et al. Gene-specific criteria for PTEN variant curation: recommendations from the ClinGen PTEN Expert Panel. Hum. Mutat. 39, 1581–1592 (2018).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res 20, 1297–1303 (2010).
DePristo, M. A. et al. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nat. Genet 43, 491–498 (2011).
McLaren, W. et al. The Ensembl Variant Effect Predictor. Genome Biol. 17, 122 (2016).
Kandoth, C. G. mskcc/vcf2maf: vcf2maf v1.6.16 (v1.6.16), https://doi.org/10.5281/zenodo.1185418 (2018).
Miller, D. T. et al. ACMG SF v3.2 list for reporting of secondary findings in clinical exome and genome sequencing: A policy statement of the American College of Medical Genetics and Genomics (ACMG). Genet Med 25, 100866 (2023).
Thorvaldsdottir, H., Robinson, J. T. & Mesirov, J. P. Integrative Genomics Viewer (IGV): high-performance genomics data visualization and exploration. Brief. Bioinform. 14, 178–192 (2013).
Acknowledgements
We are sincerely indebted to the generosity of the families and patients in PTEN clinics across the United States who contributed their time and effort to this study. We thank the clinical research team at the PTEN Multidisciplinary Clinic and Center of Excellence at the Cleveland Clinic for administrative support. We would also like to thank the PTEN Hamartoma Tumor Syndrome Foundation and the PTEN Research Foundation for their continued support in PTEN research. We gratefully acknowledge All of Us participants for their contributions, without whom this research would not have been possible. We also thank the National Institutes of Health’s All of Us Research Program for making available the participant data examined in this study. This work has been partly funded by the PTEN Research Foundation, a charity governed by English law (charity number 117358) to Professor Charis Eng under grant number CCF-19-001. We are also grateful for other funding sources, including the American Cancer Society (RPG-02-151-01-CCE and Clinical Research Professorship), National Institute of Child Health and Human Development (R01HD105049), Breast Cancer Research Foundation, Doris Duke Distinguished Clinical Scientist Award, the Ambrose Monell Foundation, and the Zacconi Center of PTEN Research Excellence (all to C.E.). C.E. was the Sondra J. and Stephen R. Hardis Chair of Cancer Genomic Medicine at the Cleveland Clinic and an American Cancer Society Clinical Research Professor. The Developmental Synaptopathies Consortium (U54NS092090) is part of the National Center for Advancing Translational Sciences (NCATS) Rare Diseases Clinical Research Network (RDCRN) and is supported by the RDCRN Data Management and Coordinating Center (DMCC) (U2CTR002818). RDCRN is an initiative of the Office of Rare Diseases Research (ORDR), NCATS, funded through a collaboration between NCATS and the National Institute of Neurological Disorders and Stroke of the National Institutes of Health (NINDS), Eunice Kennedy Shriver National Institute of Child Health & Human Development (NICHD) and National Institute of Mental Health (NIMH). The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health (NIH).
Author information
Authors and Affiliations
Consortia
Contributions
L.Y. and C.E. conceptualized and designed the project. L.Y., L.L., G.I., V.M., and Y.N. performed the experimental procedures and data analyses. L.Y., Y.N., and C.E. interpreted the data. A.H., J.A.M., D.M.R., M.S., and C.E. performed patient accrual and phenotyping. C.E. and Y.N. supervised the project. All authors drafted, critically revised, and gave final approval of the paper. L.Y. and L.L. are co-first authors of this study. C.E. passed away on the 13th day of August 2024.
Corresponding author
Ethics declarations
Competing interests
T.W.F. has received funding or research support from, acted as a consultant to, received travel support from, and/or received a speaker’s honorarium from the PTEN Research Foundation, SYNGAP Research Fund, Malan Syndrome Foundation, ADNP Kids Research Foundation, Quadrant Biosciences, Autism Speaks, Impel NeuroPharma, F. Hoffmann-La Roche AG Pharmaceuticals, the Cole Family Research Fund, Simons Foundation, Ingalls Foundation, Forest Laboratories, Ecoeos, IntegraGen, Kugona LLC, Shire Development, Bristol-Myers Squibb, Roche Pharma, MaraBio, Scioto Biosciences, Linus Biotechnology, National Institutes of Health, and the Brain and Behavior Research Foundation, has equity options in Quadrant Biosciences/Autism Analytica, MaraBio, has an investor stake in Autism EYES LLC, is employed by CentralReach, and is a co-owner of iSCAN-R and AI-Measures. C.E. was an Associate Editor for npj Genomic Medicine but played no role in the peer-review or decision to publish this manuscript. All other authors declare no competing financial or non-financial interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yehia, L., Li, L., Idumah, G. et al. Genomic modifiers of malignant and neurodevelopmental phenotypes in individuals with PTEN hamartoma tumor syndrome. npj Genom. Med. (2026). https://doi.org/10.1038/s41525-026-00556-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41525-026-00556-1