Abstract
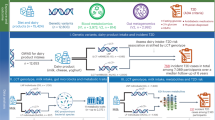
The neonatal gut is colonized by commensal bacteria, including lactobacilli, that play a critical role in promoting host health. While lactobacilli are known for producing bioactive metabolites, species-specific responses to neonatal diets remain unclear. We hypothesized that human milk and infant formula would differentially influence lactobacilli metabolite production. Six species (Lactobacillus acidophilus, Levilactobacillus brevis, Lactobacillus johnsonii, Lacticaseibacillus paracasei, Limosilactobacillus reuteri, Lacticaseibacillus rhamnosus) were cultured in defined media with human milk, formula, or water, and supernatants were analyzed via untargeted LC-MS/MS metabolomics. Principal coordinates analyses showed distinct metabolomic profiles among lactobacilli species and across dietary treatments. Human milk enhanced the production of several metabolites, including hydroxyphenyllactic acid, tyrosine, indoles, vaccenic acid, and di-peptides. Interestingly, several unique metabolites were upregulated in the presence of infant formula, including isonicotinic acid. These results highlight the differential effects of human milk and infant formula on the metabolomic profiles of lactobacilli species, emphasizing significant species-specific variation and pronounced production of potentially beneficial compounds in response to human milk. This work underscores the importance of understanding diet-microbe interactions to optimize neonatal gut health.
Similar content being viewed by others
Data availability
The raw data supporting the conclusions of this manuscript are openly available in the NIH Common Fund's National Metabolomics Data Repository (NMDR).
References
Hill, C. J. et al. Evolution of gut microbiota composition from birth to 24 weeks in the INFANTMET cohort. Microbiome 5, 4 (2017).
Tanaka, M. & Nakayama, J. Development of the gut microbiota in infancy and its impact on health in later life. Allergol. Int. 66, 515–522 (2017).
Yang, I. et al. The infant microbiome: implications for infant health and neurocognitive development. Nurs. Res. 65, 76–88 (2016).
Zhang, H., Zhang, Z., Liao, Y., Zhang, W. & Tang, D. The complex link and disease between the gut microbiome and the immune system in infants. Front. Cell Infect. Microbiol. 12, 924119 (2022).
Underwood, M. A., Mukhopadhyay, S., Lakshminrusimha, S. & Bevins, C. L. Neonatal intestinal dysbiosis. J. Perinatol. 40, 1597–1608 (2020).
Hoskinson, C. et al. Delayed gut microbiota maturation in the first year of life is a hallmark of pediatric allergic disease. Nat. Commun. 14, 4785 (2023).
Sanidad, K. Z. & Zeng, M. Y. Neonatal gut microbiome and immunity. Curr. Opin. Microbiol. 56, 30–37 (2020).
Stokholm, J. et al. Maturation of the gut microbiome and risk of asthma in childhood. Nat. Commun. 9, 141 (2018).
Depner, M. et al. Maturation of the gut microbiome during the first year of life contributes to the protective farm effect on childhood asthma. Nat. Med. 26, 1766–1775 (2020).
Carlson, A. L. et al. Infant gut microbiome associated with cognitive development. Biol. Psychiatry 83, 148–159 (2018).
Tzitiridou-Chatzopoulou, M., Kountouras, J. & Zournatzidou, G. The potential impact of the gut microbiota on neonatal brain development and adverse health outcomes. Children 11, 552 (2024).
Lu, J. et al. Effects of intestinal microbiota on brain development in humanized gnotobiotic mice. Sci. Rep. 8, 5443 (2018).
Gregory, K. E. et al. Influence of maternal breast milk ingestion on acquisition of the intestinal microbiome in preterm infants. Microbiome 4, 68 (2016).
Yi, D.Y. & Kim, S.Y. Human breast milk composition and function in human health: from nutritional components to microbiome and MicroRNAs. Nutrients 13, 362 (2021).
Zhang, X. et al. The composition and concordance of lactobacillus populations of infant gut and the corresponding breast-milk and maternal gut. Front. Microbiol. 11, 597911 (2020).
Martín, R. et al. Human milk is a source of lactic acid bacteria for the infant gut. J. Pediatr. 143, 754–758 (2003).
Martín, R., Heilig, G. H., Zoetendal, E. G., Smidt, H. & Rodríguez, J. M. Diversity of the Lactobacillus group in breast milk and vagina of healthy women and potential role in the colonization of the infant gut. J. Appl. Microbiol. 103, 2638–2644 (2007).
Łubiech, K. & Twarużek, M. Lactobacillus bacteria in breast milk. Nutrients 12, 3783 (2020).
Satokari, R. M. et al. Diversity of bifidobacterium and lactobacillus spp. in breast-fed and formula-fed infants as assessed by 16S rDNA sequence differences. Microb. Ecol. Health Dis. 14, 97–105 (2002).
Chau, K. et al. Probiotics for infantile colic: a randomized, double-blind, placebo-controlled trial investigating Lactobacillus reuteri DSM 17938. J. Pediatr. 166, 74–78 (2015).
Baldassarre, M.E. et al. Effectiveness and safety of a probiotic-mixture for the treatment of infantile colic: a double-blind, randomized, placebo-controlled clinical trial with fecal real-time PCR and NMR-based metabolomics analysis. Nutrients 10, 212 (2018).
Indrio, F. et al. The effects of probiotics on feeding tolerance, bowel habits, and gastrointestinal motility in preterm newborns. J. Pediatr. 152, 801–806 (2008).
Zhang, W. et al. Clinical efficacy of probiotics on feeding intolerance in preterm infants: a systematic review and meta-analysis. Transl. Pediatr. 11, 229–238 (2022).
Dempsey, E. & Corr, S. C. Lactobacillus spp. for gastrointestinal health: current and future perspectives. Front. Immunol. 13, 840245 (2022).
Tang, H., Huang, W. & Yao, Y. F. The metabolites of lactic acid bacteria: classification, biosynthesis and modulation of gut microbiota. Micro Cell 10, 49–62 (2023).
Krishna, M. et al. Maternal Lactobacillus reuteri supplementation shifts the intestinal microbiome in mice and provides protection from experimental colitis in female offspring. FASEB Bioadv 4, 109–120 (2022).
Engevik, M. et al. Limosilactobacillus reuteri ATCC 6475 metabolites upregulate the serotonin transporter in the intestinal epithelium. Benef. Microbes 12, 583–599 (2021).
Engevik, M. A. et al. Immunomodulation of dendritic cells by Lactobacillus reuteri surface components and metabolites. Physiol. Rep. 9, e14719 (2021).
Chamberlain, M., O’Flaherty, S., Cobián, N. & Barrangou, R. Metabolomic analysis of Lactobacillus acidophilus, L. gasseri, L. crispatus, and Lacticaseibacillus rhamnosus strains in the presence of pomegranate extract. Front. Microbiol. 13, 863228 (2022).
Engevik, M. A. et al. Reuterin disrupts Clostridioides difficile metabolism and pathogenicity through reactive oxygen species generation. Gut Microbes 12, 1788898 (2020).
Hall, A. E., Engevik, M. A., Oezguen, N., Haag, A. & Versalovic, J. ClC transporter activity modulates histidine catabolism in Lactobacillus reuteri by altering intracellular pH and membrane potential. Microb. Cell Fact. 18, 212 (2019).
Engevik, M. A. et al. Microbial metabolic capacity for intestinal folate production and modulation of host folate receptors. Front. Microbiol. 10, 2305 (2019).
Engevik, K. A. et al. Phylogenetically diverse bacterial species produce histamine. Syst. Appl. Microbiol. 47, 126539 (2024).
Miao, H. et al. Lactobacillus species ameliorate membranous nephropathy through inhibiting the aryl hydrocarbon receptor pathway via tryptophan-produced indole metabolites. Br. J. Pharm. 181, 162–179 (2024).
Zhang, Q. et al. Lactobacillus plantarum-derived indole-3-lactic acid ameliorates colorectal tumorigenesis via epigenetic regulation of CD8(+) T cell immunity. Cell Metab. 35, 943–960.e949 (2023).
Fong, W. et al. Lactobacillus gallinarum-derived metabolites boost anti-PD1 efficacy in colorectal cancer by inhibiting regulatory T cells through modulating IDO1/Kyn/AHR axis. Gut 72, 2272–2285 (2023).
Le, B., Anh, P.T.N. & Yang, S.H. Enhancement of the anti-inflammatory effect of mustard kimchi on RAW 264.7 macrophages by the lactobacillus plantarum fermentation-mediated generation of phenolic compound derivatives. Foods 9, 1815 (2020).
Menard, S. et al. Lactic acid bacteria secrete metabolites retaining anti-inflammatory properties after intestinal transport. Gut 53, 821–828 (2004).
Guo, S. et al. Secreted metabolites of bifidobacterium infantis and lactobacillus acidophilus protect immature human enterocytes from IL-1beta-induced inflammation: a transcription profiling analysis. PLoS One 10, e0124549 (2015).
Gu, Q. et al. Lactobacillus plantarum ZJ316 alleviates ulcerative colitis by inhibiting inflammation and regulating short-chain fatty acid levels and the gut microbiota in a mouse model. Food Funct. 14, 3982–3993 (2023).
Vemuri, R. et al. Lactobacillus acidophilus DDS-1 modulates intestinal-specific microbiota, short-chain fatty acid and immunological profiles in aging mice. Nutrients 11, 1297 (2019).
Xie, Z., Li, M., Qian, M., Yang, Z. & Han, X. Co-cultures of lactobacillus acidophilus and bacillus subtilis enhance mucosal barrier by modulating gut microbiota-derived short-chain fatty acids. Nutrients 14, 3574 (2022).
Xu, B. et al. Milk-derived Lactobacillus with high production of short-chain fatty acids relieves antibiotic-induced diarrhea in mice. Food Funct. 15, 5329–5342 (2024).
Makki, K., Deehan, E. C., Walter, J. & Backhed, F. The impact of dietary fiber on gut microbiota in host health and disease. Cell Host Microbe 23, 705–715 (2018).
Stewart, M. L., Savarino, V. & Slavin, J. L. Assessment of dietary fiber fermentation: effect of Lactobacillus reuteri and reproducibility of short-chain fatty acid concentrations. Mol. Nutr. Food Res. 53, S114–120 (2009).
Sinha, A. K. et al. Dietary fibre directs microbial tryptophan metabolism via metabolic interactions in the gut microbiota. Nat. Microbiol. 9, 1964–1978 (2024).
Huang, Z., Boekhorst, J., Fogliano, V., Capuano, E. & Wells, J. M. Impact of high-fiber or high-protein diet on the capacity of human gut microbiota to produce tryptophan catabolites. J. Agric Food Chem. 71, 6956–6966 (2023).
Filannino, P., Bai, Y., Di Cagno, R., Gobbetti, M. & Ganzle, M. G. Metabolism of phenolic compounds by Lactobacillus spp. during fermentation of cherry juice and broccoli puree. Food Microbiol. 46, 272–279 (2015).
Odiase, E. et al. The gut microbiota differ in exclusively breastfed and formula-fed United States infants and are associated with growth status. J. Nutr. 153, 2612–2621 (2023).
Inchingolo, F. et al. Difference in the intestinal microbiota between breastfeed infants and infants fed with artificial milk: a systematic review. Pathogens 13, 336 (2024).
Mai, T.T., Tran, D.Q., Roos, S., Rhoads, J.M. & Liu, Y. Human breast milk promotes the secretion of potentially beneficial metabolites by probiotic Lactobacillus reuteri DSM 17938. Nutrients 11, 1358 (2019).
Repa, A. et al. Probiotics (Lactobacillus acidophilus and Bifidobacterium infantis) prevent NEC in VLBW infants fed breast milk but not formula [corrected]. Pediatr. Res. 77, 381–388 (2015).
Klopp, A. et al. Modes of infant feeding and the risk of childhood asthma: a prospective birth cohort study. J. Pediatr. 190, 192–199.e192 (2017).
El-Heneidy, A., Abdel-Rahman, M.E., Mihala, G., Ross, L.J. & Comans, T.A. Milk other than breast milk and the development of asthma in children 3 years of age. a birth cohort study (2006–2011). Nutrients 10, 1798 (2018).
McGuire, W. & Anthony, M. Y. Donor human milk versus formula for preventing necrotising enterocolitis in preterm infants: systematic review. Arch. Dis. Child Fetal Neonatal Ed. 88, F11–14 (2003).
Altobelli, E., Angeletti, P.M., Verrotti, A. & Petrocelli, R. The impact of human milk on necrotizing enterocolitis: a systematic review and meta-analysis. Nutrients 12, 1322 (2020).
Boyd, C. A., Quigley, M. A. & Brocklehurst, P. Donor breast milk versus infant formula for preterm infants: systematic review and meta-analysis. Arch. Dis. Child Fetal Neonatal Ed. 92, F169–175 (2007).
Cortez, J. et al. Maternal milk feedings reduce sepsis, necrotizing enterocolitis and improve outcomes of premature infants. J. Perinatol. 38, 71–74 (2018).
Fernandez, L. et al. The human milk microbiota: origin and potential roles in health and disease. Pharm. Res. 69, 1–10 (2013).
Yang, B. et al. Bifidobacterium and Lactobacillus composition at species level and gut microbiota diversity in infants before 6 weeks. Int. J. Mol. Sci. 20, 3306 (2019).
Urbanska, M. & Szajewska, H. The efficacy of Lactobacillus reuteri DSM 17938 in infants and children: a review of the current evidence. Eur. J. Pediatr. 173, 1327–1337 (2014).
Roggero, P. et al. Analysis of immune, microbiota and metabolome maturation in infants in a clinical trial of Lactobacillus paracasei CBA L74-fermented formula. Nat. Commun. 11, 2703 (2020).
Murakami, K., Habukawa, C., Nobuta, Y., Moriguchi, N. & Takemura, T. The effect of Lactobacillus brevis KB290 against irritable bowel syndrome: a placebo-controlled double-blind crossover trial. Biopsychosoc. Med. 6, 16 (2012).
Manzoni, P. et al. Routine Lactobacillus rhamnosus GG administration in VLBW infants: a retrospective, 6-year cohort study. Early Hum. Dev. 87(Suppl 1), S35–38 (2011).
Suzuki, Y. et al. Identification of antioxidants produced by Lactobacillus plantarum. Biosci. Biotechnol. Biochem. 77, 1299–1302 (2013).
Gu, X. et al. The antioxidant activity and metabolomic analysis of the supernatant of Streptococcus alactolyticus strain FGM. Sci. Rep. 14, 8413 (2024).
Vogie, K., Mantick, N. & Carlson, G. Metabolism and toxicity of the styrene metabolite 4-vinylphenol in CYP2E1 knockout mice. J. Toxicol. Environ. Health A 67, 145–152 (2004).
Carlson, G. P., Ullman, M., Mantick, N. A. & Snyder, P. W. 4-Vinylphenol-induced pneumotoxicity and hepatotoxicity in mice. Toxicol. Pathol. 30, 565–569 (2002).
Girawale, S.D., Meena, S.N., Yadav, A.A. & Kodam, K.M. Biocatalysis at its finest: Bacillus paralicheniformis produces 4-vinylphenol with potent biological activities. Biocatal. Agric. Biotechnol. 52, 102837 (2023).
Rodriguez, A., Meadows, J. A., Sun, N., Simmons, B. A. & Gladden, J. M. Evaluation of bacterial hosts for conversion of lignin-derived p-coumaric acid to 4-vinylphenol. Micro Cell Fact. 20, 181 (2021).
Santamaria, L., Reveron, I., de Felipe, F.L., de Las Rivas, B. & Munoz, R. Ethylphenol formation by lactobacillus plantarum: identification of the enzyme involved in the reduction of vinylphenols. Appl. Environ. Microbiol. 84, e01064-18 (2018).
Ehrlich, A. M. et al. Indole-3-lactic acid associated with Bifidobacterium-dominated microbiota significantly decreases inflammation in intestinal epithelial cells. BMC Microbiol. 20, 357 (2020).
Meng, D. et al. Indole-3-lactic acid, a metabolite of tryptophan, secreted by Bifidobacterium longum subspecies infantis is anti-inflammatory in the immature intestine. Pediatr. Res. 88, 209–217 (2020).
Cristina, M. O. et al. Mechanisms and therapeutic potential of key anti-inflammatory metabiotics: trans-vaccenic acid, indole-3-lactic acid, thiamine, and butyric acid. Probiotics Antimicrob Proteins 17, 2805–2818 (2025).
Kim, K., Kim, H. & Sung, G.Y. Effects of indole-3-lactic acid, a metabolite of tryptophan, on IL-4 and IL-13-induced human skin-equivalent atopic dermatitis models. Int. J. Mol. Sci. 23, 13520 (2022).
Qian, X. et al. Bifidobacteria with indole-3-lactic acid-producing capacity exhibit psychobiotic potential via reducing neuroinflammation. Cell Rep. Med. 5, 101798 (2024).
Bassett, C. M. et al. Dietary vaccenic acid has antiatherogenic effects in LDLr-/- mice. J. Nutr. 140, 18–24 (2010).
Valenzuela, C.A., Baker, E.J., De Souza, C.O., Miles, E.A. & Calder, P.C. Differential effects of ruminant and industrial 18-carbon trans-monounsaturated fatty acids (trans vaccenic and elaidic) on the inflammatory responses of an endothelial cell line. Molecules 26, 5313 (2021).
Jacome-Sosa, M. et al. Vaccenic acid suppresses intestinal inflammation by increasing anandamide and related N-acylethanolamines in the JCR:LA-cp rat. J. Lipid Res. 57, 638–649 (2016).
Ory, R. L., Hood, D. W. & Lyman, C. M. The role of glutamine in the synthesis of arginine by Lactobacillus arabinosus. J. Biol. Chem. 207, 267–273 (1954).
Oleskin, A. et al. Lactic-acid bacteria supplement fermented dairy products with human behavior-modifying neuroactive compounds. J. Pharm. Nutr. Sci. 4, 199–206 (2014).
Wu, G., Fang, Y. Z., Yang, S., Lupton, J. R. & Turner, N. D. Glutathione metabolism and its implications for health. J. Nutr. 134, 489–492 (2004).
Barker-Tejeda, T. C. et al. Comparative characterization of the infant gut microbiome and their maternal lineage by a multi-omics approach. Nat. Commun. 15, 3004 (2024).
Vallès, Y. et al. Microbial succession in the gut: directional trends of taxonomic and functional change in a birth cohort of Spanish infants. PLoS Genet. 10, e1004406 (2014).
Palmer, C., Bik, E. M., DiGiulio, D. B., Relman, D. A. & Brown, P. O. Development of the human infant intestinal microbiota. PLoS Biol. 5, e177 (2007).
Hattoufi, K. et al. Association of gut microbiota and type of feeding: Molecular analysis of a cohort of preterm moroccan newborns. Pediatr. Neonatol. S1875-9572, 00195-0 (2025).
Dawod, B., Marshall, J. S. & Azad, M. B. Breastfeeding and the developmental origins of mucosal immunity: how human milk shapes the innate and adaptive mucosal immune systems. Curr. Opin. Gastroenterol. 37, 547–556 (2021).
Lopez, D. A. et al. Breastfeeding duration is associated with domain-specific improvements in cognitive performance in 9-10-year-old children. Front. Public Health 9, 657422 (2021).
Bene, J. et al. Differences in circulating carnitine status of preterm infants fed fortified human milk or preterm infant formula. J. Pediatr. Gastroenterol. Nutr. 57, 673–676 (2013).
Whitfield, J., Smith, T., Sollohub, H., Sweetman, L. & Roe, C. R. Clinical effects of L-carnitine supplementation on apnea and growth in very low birth weight infants. Pediatrics 111, 477–482 (2003).
Shortland, G. J. et al. Randomised controlled trial of L-carnitine as a nutritional supplement in preterm infants. Arch. Dis. Child Fetal Neonatal Ed. 78, F185–188 (1998).
Rajamani, S. et al. The vitamin riboflavin and its derivative lumichrome activate the LasR bacterial quorum-sensing receptor. Mol. Plant Microbe Interact. 21, 1184–1192 (2008).
Lopez, B.R., Palacios, O.A., Bashan, Y., Hernandez-Sandoval, F.E. & de-Bashan, L.E. Riboflavin and lumichrome exuded by the bacterium Azospirillum brasilense promote growth and changes in metabolites in Chlorella sorokiniana under autotrophic conditions. Algal Res. 44, 101671 (2019).
Yamamoto, K. & Asano, Y. Efficient production of lumichrome by Microbacterium sp. strain TPU 3598. Appl. Environ. Microbiol. 81, 7360–7367 (2015).
Phillips, D. A. et al. Identification of lumichrome as a sinorhizobium enhancer of alfalfa root respiration and shoot growth. Proc. Natl. Acad. Sci. USA 96, 12275–12280 (1999).
Dakora, F. D., Matiru, V. N. & Kanu, A. S. Rhizosphere ecology of lumichrome and riboflavin, two bacterial signal molecules eliciting developmental changes in plants. Front. Plant Sci. 6, 700 (2015).
Robledo, D.A. & Muhammad, G. Evaluation of the cytotoxic properties of lumichrome-derived compounds on breast adenocarcinoma, colorectal cancer, and liver carcinoma.Cent. Asian J. Med. Nat. Sci. 3, 203–208 (2022).
Chantarawong, W., Kuncharoen, N., Tanasupawat, S. & Chanvorachote, P. Lumichrome inhibits human lung cancer cell growth and induces apoptosis via a p53-dependent mechanism. Nutr. Cancer 71, 1390–1402 (2019).
Seferovic, M. D. et al. Maternal diet alters human milk oligosaccharide composition with implications for the milk metagenome. Sci. Rep. 10, 22092 (2020).
Xu, R. et al. The human milk microbiome is minimally associated with breastfeeding practices. Sci. Rep. 15, 19308 (2025).
Moossavi, S. et al. Composition and variation of the human milk microbiota are influenced by maternal and early-life factors. Cell Host Microbe 25, 324–335.e324 (2019).
Tran, L. C. et al. The metabolome of human milk is altered differentially by Holder pasteurization and high hydrostatic pressure processing. Front. Nutr. 10, 1107054 (2023).
Ten-Domenech, I. et al. The effect of Holder pasteurization on the lipid and metabolite composition of human milk. Food Chem. 384, 132581 (2022).
Waller, M. E. et al. Analyzing the responses of enteric bacteria to neonatal intensive care supplements. Int. J. Microbiol 2024, 3840327 (2024).
Horvath, T. D. et al. Interrogation of the mammalian gut–brain axis using LC–MS/MS-based targeted metabolomics with in vitro bacterial and organoid cultures and in vivo gnotobiotic mouse models. Nat. Protoc. 18, 490–529 (2023).
Engevik, M. A., Stripe, L. K., Baatz, J. E., Wagner, C. L. & Chetta, K. E. Identifying single-strain growth patterns of human gut microbes in response to preterm human milk and formula. Food Funct. 13, 5571–5589 (2022).
Engevik, M. A. et al. The metabolic profile of Bifidobacterium dentium reflects its status as a human gut commensal. BMC Microbiol. 21, 154 (2021).
Engevik, M. A. et al. Bifidobacterium dentium-derived y-glutamylcysteine suppresses ER-mediated goblet cell stress and reduces TNBS-driven colonic inflammation. Gut Microbes 13, 1–21 (2021).
Davoodabadi, A., Dallal, M., Ebrahimi, M. & Rajabi, Z. Colonization of Lactobacilli among the fecal microflora of healthy Iranian breast-fed infants. J. Clin. Gastroenterol. Treat. 6, 072 (2020).
Demirok, N., Durak, M. & Arici, M. Probiotic lactobacilli in faeces of breastfed babies. Food Sci. Technol. 42, e58221 (2021).
Haarman, M. & Knol, J. Quantitative real-time PCR analysis of fecal Lactobacillus species in infants receiving a prebiotic infant formula. Appl. Environ. Microbiol. 72, 2359–2365 (2006).
Al Kassaa, I. et al. Characterization of lactobacilli strains isolated from baby’s feces for their potential immunobiotic application. Iran. J. Microbiol. 11, 379–388 (2019).
Fortmann, I. et al. Lactobacillus acidophilus/bifidobacterium infantis probiotics are beneficial to extremely low gestational age infants fed human milk. Nutrients 12, 850 (2020).
Waki, N., Matsumoto, M., Fukui, Y. & Suganuma, H. Effects of probiotic Lactobacillus brevis KB290 on incidence of influenza infection among schoolchildren: an open-label pilot study. Lett. Appl. Microbiol. 59, 565–571 (2014).
Haschke-Becher, E. et al. Urinary D-lactate excretion in infants receiving Lactobacillus johnsonii with formula. Ann. Nutr. Metab. 53, 240–244 (2008).
Wang, S. et al. A randomized, double blind, parallel, placebo-controlled study to investigate the efficacy of Lactobacillus paracasei N1115 in gut development of young children. Food Sci. Nutr. 9, 6020–6030 (2021).
Spreckels, J.E. et al. Lactobacillus reuteri Colonisation of extremely preterm infants in a randomised placebo-controlled trial. Microorganisms 9, 915 (2021).
Vendt, N. et al. Growth during the first 6 months of life in infants using formula enriched with Lactobacillus rhamnosus GG: double-blind, randomized trial. J. Hum. Nutr. Diet. 19, 51–58 (2006).
Vivatvakin, B. & Kowitdamrong, E. Randomized control trial of live Lactobacillus acidophilus plus Bifidobacterium infantis in treatment of infantile acute watery diarrhea. J. Med Assoc. Thai 89, S126–133 (2006).
Savino, F. et al. Lactobacillus reuteri DSM 17938 in infantile colic: a randomized, double-blind, placebo-controlled trial. Pediatrics 126, e526–533 (2010).
Basturk, A., Isik, I., Atalay, A. & Yilmaz, A. Investigation of the efficacy of Lactobacillus rhamnosus GG in infants with Cow’s milk protein allergy: a randomised double-blind placebo-controlled trial. Probiotics Antimicrob. Proteins 12, 138–143 (2020).
Schymanski, E. L. et al. Identifying small molecules via high resolution mass spectrometry: communicating confidence. Environ. Sci. Technol. 48, 2097–2098 (2014).
Acknowledgements
We would like to express our deepest appreciation to the families that participated in this study. The Texas Children’s Research Institute provides salary support to staff members who work in the Virginia and L.E. Simmons Family Foundation Mass Spectrometry Laboratory and purchased all of the reagents, consumables and durable supplies described.
Funding
This study was supported by T32DK124191-01A1 (AG), NATS NIH KL2TR001452 (KEC), Darby Pediatrics pilot grant (KEC), UL1TR001450 (KEC), 3P20 P20 GM130457 (COBRE in Digestive & Liver Disease, MUSC; GM130457-04S1 supplement), P30 DK123704 (Digestive Disease Research Center, MUSC). The Virginia and L.E. Simmons Family Foundation Mass Spectrometry Laboratory housed within the Texas Children’s Research Institute - Microbiome Center is supported by an NIH S10 Shared Instrumentation Grant 1S10ODO36416-01 (TDH), and by a NIH P30DK056338 (TDH).
Author information
Authors and Affiliations
Contributions
Concept and design (A.G., B.P., M.A.E.); intellectual contribution (K.E.C., T.D.H., M.A.E.); data acquisition and analysis (A.G., B.P.), interpretation (A.G., K.E.C., M.A.E.); drafting manuscript (A.G.), editing manuscript (B.P., K.E.C., T.D.H., M.A.E.); funding (K.E.C., T.D.H., M.A.E.).
Corresponding author
Ethics declarations
Competing interests
T.D.H. is an Editorial Board Member and is contracted as an Associate Academic Editor for Cell Press – STAR Protocols. T.D.H. is a voluntary participant in the SCIEX Global Thought Leaders in Mass Spectrometry Program. All other authors have no competing interests to declare.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Gutierrez, A., Chetta, K.E., Puckett, B. et al. Human milk and infant formula influence lactobacilli metabolism in a species-specific manner. npj Sci Food (2026). https://doi.org/10.1038/s41538-026-00759-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41538-026-00759-x