Abstract
Type I interferons (IFN) are key mediators of innate immune activation, promoting upregulation of costimulatory molecules and Major Histocompatibility Complex (MHC) I/II on antigen-presenting cells (APCs). However, IFN also suppress endogenous translation to restrict viral replication. Critically, IFN-stimulated APCs lose the capacity to acquire new antigens, making the timing of IFN signaling a crucial determinant of vaccine efficacy. Here, we show that both DC-specific loss of IFNα/β receptor (IFNαR) and transient blockade of IFNαR before vaccination enhances vaccine uptake and expression within DCs, improves CD8⁺ T cell priming, and leads to superior tumor control. We also demonstrate that IFN signaling before vaccination, triggered by prior infection or administration of a different vaccine, impairs dendritic cell uptake of mRNA-LNP vaccines and reduces the magnitude of vaccine-specific CD8⁺ T cell responses. These findings highlight the dual-edged nature of IFN signaling and offer a potential strategy for enhancing vaccine-induced immunity.
Similar content being viewed by others
Introduction
mRNA-based vaccines have revolutionized vaccinology. Beyond its major impact in aiding to resolve the COVID-19 pandemic, the potential of mRNA-based therapeutics is vast, with applications extending from viruses like influenza1 and human immunodeficiency virus2,3 to cancer4,5,6. Notably, mRNA-based neoantigen therapies for hard-to-treat pancreatic cancer are already currently in clinical trials7. Therefore, it is critical to understand the underlying immune mechanisms governing their response and how to augment the efficacy of mRNA-lipid nanoparticle (mRNA-LNP) vaccines. It is well established that mRNA-LNPs induce type I interferons (IFN) (e.g., IFN-α, IFN-β) upon immunization8,9,10,11. IFN plays a critical role in antiviral immune responses by limiting viral spread and activating innate immunity to elicit long-lasting adaptive immunity12. IFN signaling has been shown to be required for CD8⁺ T cells to develop their cytotoxic capacity13 and plays a crucial role in augmenting antigen-presenting cell (APC) activation and the priming of adaptive immune responses14,15. IFN signaling in APCs at the time of antigen acquisition has been shown to significantly enhance the subsequent adaptive immune response through upregulation of costimulatory molecules and promotion of inflammatory cytokine production. Thus, IFN has canonically been thought to augment or induce immune responses. However, multiple studies have shown that chronic IFN signaling is detrimental to the overall immune response, leading to immune suppression16,17,18,19,20. It has also been shown that once a conventional dendritic cell (cDC) or APC is activated and matures, it can no longer acquire new material for presentation to the adaptive immune system21,22. Similarly, previous research showed that migratory, or mature, DCs are unable to acquire new cell-associated material for presentation to CD8⁺ T cells23. This suggests that the timing of APC activation may determine whether IFN acts as a stimulatory or inhibitory factor in APC-mediated antigen processing and presentation: a fully mature or activated APC is incapable of processing and presenting new antigens. Recent studies suggest that IFN signaling can also suppress adaptive immune responses during viral infections. Notably, transient blockade of IFN signaling using the IFNαR blocking antibody MAR1-5A324 (αIFNαR) has been shown to enhance virus-specific immune responses25. Similarly, a recent publication found that transient IFNαR blockade enhanced virus-specific immune responses in the context of an arthritogenic alphavirus26. Still, the underlying mechanisms remain unclear.
In the context of vaccination, transient IFNαR inhibition, as well as the use of IFNαR-deficient mice or tissue-specific IFNαR deletion, have yielded variable outcomes. A recent article demonstrated that blocking IFN signaling reduced overall CD8⁺ and CD4⁺ T cell responses following mRNA-LNP vaccination11. In contrast, a separate study found that transient IFNαR blockade during LCMV infection and vaccination enhanced the generation of virus- or vaccine-specific stem cell–like memory CD8⁺ T cells27. Thus, the role of IFN in vaccines, including mRNA-LNP-based vaccines, remains unclear, with evidence supporting both inhibitory and augmentative effects.
In this study, we aimed to elucidate the role of IFN signaling during mRNA-LNP-based vaccination by utilizing transient IFNαR blockade and mice with DC-specific loss of IFNαR. Given the inhibitory impacts of inflammatory cytokine signaling on DC antigen acquisition, we hypothesized that early IFN signaling within DCs impairs their ability to acquire and translate the new antigenic mRNA, thereby limiting antigen presentation and ultimately leading to a diminished adaptive immune response. Understanding this dynamic may provide insight into the timing and regulation of IFN signaling in optimizing vaccine efficacy.
Results
mRNA-LNP vaccine induced IFN reduces vaccine-specific mRNA expression by DCs in vitro
To assay if IFN signaling inhibited acquisition of mRNA-LNP, we generated bone marrow-derived DCs (BMDCs) using a novel culture method with stem cell factor (SCF) and FMS-like tyrosine kinase 3 ligand (Flt3L)28. This protocol enhances DC output and produces subsets of conventional DCs that closely resemble their in vivo-derived counterparts (Supplementary Fig. 1). To assess mRNA-LNP vaccine uptake, we incubated BMDCs with eGFP mRNA-LNP vaccines and measured eGFP expression by flow cytometry. 16-hour pretreatment of BMDCs with IFNα completely abrogated eGFP expression, resulting in no increase of eGFP expression above PBS incubated controls (Fig. 1A). This suggests that premature activation of BMDCs through IFNαR signaling inhibited mRNA-LNP acquisition and expression. Given the rapid nature of IFN signaling29, we hypothesized that once a cell senses mRNA-LNP, it produces IFN, which in turn suppresses mRNA-LNP uptake by surrounding DCs. To test this, we blocked IFNαR signaling at the time of mRNA-LNP administration using anti-IFNαR blocking antibody MAR1-5A3. Blocking IFN signaling during mRNA-LNP administration increased the proportion of DCs acquiring and expressing eGFP (Fig. 1A). When assessing the DC activation marker CD40, IFNα pretreatment and mRNA-LNP alone uniformly activated all DCs, whereas IFNαR blockade significantly reduced CD40 upregulation (Fig. 1B). These findings suggest that inhibiting premature IFN-mediated DC activation enhances mRNA-LNP uptake and expression by DCs in vitro.
A (Left) Representation flow plots of eGFP expression of BMDCs incubated with 2.5 μg eGFP mRNA-LNP alone, with mRNA-LNP and MAR1-5A3 antibody (αIFNαR, red), or with mRNA-LNP and 103 U/mL IFNα for 16 h (blue). (Right) eGFP expression as a percentage of all DCs incubated with 2.5 μg eGFP mRNA-LNP alone (white), with mRNA-LNP and MAR1-5A3 antibody (αIFNαR, red), or with mRNA-LNP and 103 U/mL IFNα for 16 h (blue). B (Left) Representative flow plots of CD40 MFI of BMDCs incubated with 2.5 μg eGFP mRNA-LNP alone, with mRNA-LNP and MAR1-5A3 antibody (αIFNαR, red), or with mRNA-LNP and 103 U/mL IFNα for 16 h (blue). (Right) CD40 MFI of DCs incubated with 2.5 μg eGFP mRNA-LNP alone (white), with mRNA-LNP and MAR1-5A3 antibody (αIFNαR, red), or with mRNA-LNP and 103 U/mL IFNα for 16 h (blue). Data represents pooled independent samples from four independent experiments (n = 11 for empty, n = 11 for mRNA alone, n = 12 for IFNα, and n = 11 for αIFNαR). Data were compared using one-way ANOVA.
Inhibition of IFNαR signaling enhances dendritic cell acquisition of mRNA-LNPs and improves vaccine-specific CD8⁺ T cell responses
To determine whether blocking IFNαR signaling enhances DC acquisition of mRNA-LNPs in vivo, we first confirmed that IFN is produced following immunization with our mRNA-LNP vaccine. IFNα 2 and 4 were found in the serum of immunized WT C57BL/6 J mice at 8 h and was absent by 24 h post vaccination, in agreement with previous findings showing IFN production is early and transient post mRNA-LNP vaccination6 (Fig. 2A). Previous studies have shown that administering 100 μg of IFNαR-blocking antibody MAR1-5A3 reduces surface expression of IFNαR1 for five days, with receptor expression returning by day six25. We verified these results; however, we saw that MAR1-5A3 antibody used at 250 μg reduced IFNαR1 surface expression for at least 9 days post injection with expression not returning to control WT C57BL/6 J levels over the experimental time course (Fig. 2B). Next, we asked whether blocking the IFNαR receptor before immunization with an eGFP mRNA-LNP vaccine would increase DC mRNA-LNP acquisition and translation. To test this, we blocked IFNαR one day before immunizing mice with an eGFP mRNA-LNP vaccine. Splenic cDCs (Supplementary Fig. 2A) were isolated 24 h post-immunization and analyzed for eGFP expression by flow cytometry. IFNαR inhibition significantly increased the number of splenic DCs expressing eGFP compared to unblocked controls (Fig. 2C). In terms of DC activation, both eGFP and OVA mRNA-LNP vaccines induced upregulation of the activation markers CD40 and CD86 on splenic DCs; however, IFNαR blockade substantially reduced CD40 and CD86 expression compared to unblocked, immunized mice. Despite this reduction, activation levels in IFNαR-blocked, eGFP-vaccinated mice remained higher than in PBS-immunized controls (Fig. 2D). In addition, DCs that had acquired eGFP mRNA-LNP were activated, albeit less than unblocked controls (Supplementary Fig. 3A). Moreover, blocking IFNαR signaling during immunization did not alter overall DC numbers (Supplementary Fig. 3B). Thus, inhibition of IFNαR signaling leads to an increase in DC mediated mRNA-LNP acquisition and expression. With this in mind, we next asked whether blocking the IFNαR receptor before immunization with the OVA mRNA-LNP vaccine would increase vaccine-specific CD8⁺ T cells. Blocking IFNαR prior to immunization significantly increased vaccine-specific CD8⁺ T cells (Supplementary Figs. 2B, 3C, and Fig. 2E). In addition, vaccine-specific CD8⁺ T cells expressed higher surface levels of PD-1 (Fig. 2F), potentially pointing to increased T cell receptor signaling due to increased antigen presentation30,31. To ensure that blocking IFNαR increased mRNA-LNP acquisition and not merely mRNA translation, we immunized mice with mRNA-LNP incorporated with the fluorescent lipid 1,1’-dioctadecyl-3,3,3’,3’-tetramethylindodicarbocyanine (DiD) which incorporates into the endosomal membrane once acquired by cells32. IFNαR inhibition 24 h prior to immunization significantly increased the number of splenic DCs that acquired and expressed the fluorescent LNP compared to unblocked controls indicating an increase in mRNA-LNP acquisition by DCs (Fig. 2G). These findings support our hypothesis that inhibiting IFNαR signaling prevents bystander DC activation, allowing for enhanced mRNA-LNP uptake and subsequent adaptive immune priming.
A Serum levels of IFNα 2 and 4 at various timepoints after immunization with 2.5 μg of mRNA-LNP from four C57BL/6J wild-type mice (WT) pooled together. B (Left) Surface expression of IFNαR1 from mice given 250 μg of MAR1-5A3 (αIFNαR, red) (IP), or no MAR1-5A3 (black) and compared to an IFNαR knockout mouse (blue). (Right) Representative plot showing IFNαR1 MFI of each mouse at days 1, 3, 5, 7, and 9 post injection of MAR1-5A3 (red, n = 5) compared to a WT mouse with no MAR1-5A3 injection (black, n = 1) or an IFNαR knockout mouse (blue, n = 1). C (Left) Representative flow plot of splenic CD11c+MHCII+ DCs expressing eGFP from WT mice with or without MAR1-5A3. (Right) Percentage of eGFP expression in splenic DCs 24 h post mRNA-LNP immunization (5 μg eGFP mRNA-LNP or 2.5 μg OVA mRNA-LNP IV) in WT mice (black, n = 6), or WT mice given MAR1-5A3 prior to immunization (red, n = 8). D (Left) Representative flow plot showing CD40+CD86+ splenic DCs after immunization with mRNA-LNP with or without MAR1-5A3. (Right) Percentage of CD40+CD86+ splenic DCs from mice in Fig. 2C. E (Left) Representative flow plots of SIINFEKL-H2-Kb (OVA) specific CD8⁺ T cells. (Right) Percentage of OVA specific CD8⁺ T cells from WT mice (black, n = 8) and WT mice given MAR1-5A3 (red, n = 11). 2.5 μg of mRNA-LNP was administered IV 24 h after MAR1-5A3. F (Left) Representative flow plot showing PD-1 expression of OVA specific TCRβ+CD8α+ T cells. (Right) PD-1 expression of TCRβ+CD8α+ T cells from mice in Fig. 2E. G (Left) Representative flow plot of splenic CD11c+MHCII+ DCs expressing fluorescent LNP from WT mice with or without MAR1-5A3. (Right) Percentage of LNP expression in splenic DCs 24 h post mRNA-LNP immunization (5 μg DiD incorporated LNP eGFP mRNA-LNP or 2.5 μg OVA mRNA-LNP IV) in WT mice (black, n = 10), or WT mice given MAR1-5A3 prior to immunization (red, n = 11). Error bars in (A) indicate ± SD. All bars in (C)–(G) indicate the mean. Data in (C) and (E) were compared using a Mann–Whitney test. Data in (D) were compared with either an unpaired two-tailed students T test or a Mann–Whitney test. Data in (F) and (G) were compared by an unpaired two-tailed students T test.
DC specific loss of IFNαR increases vaccine-specific CD8⁺ T cell responses
To further test the notion that IFN signaling inhibits mRNA-LNP vaccine acquisition and antigen presentation specifically in DCs we assayed mRNA-LNP vaccine responses in mice that lack DC-specific expression of IFNαR. We utilized the CD11c cre (Itgaxcre)33 mice which have DC-specific expression of cre recombinase crossed to mice which contain LoxP sites flanking exon 3 of Ifnαr34 (Ifnαrfl/fl) to test DC-specific deletion of IFNαR. We immunized Itgaxcre Ifnαrfl/fl (IfnαrcKO) and Ifnαr fl/fl (IfnαrWT) mice with mRNA-LNP vaccines and assayed for eGFP expression at 24 h. DCs from IfnαrcKO mice had a significantly higher proportion of splenic DCs expressing eGFP compared to IfnαrWT controls (Fig. 3A). Similarly, IfnαrcKO DCs were significantly less activated as evidenced by surface expression of CD40 and CD86 when compared to IfnαrWT controls (Fig. 3B). Similar to blocking IFNαR signaling during immunization, DC-specific deletion of Ifnαr did not alter overall DC numbers after immunization (Supplementary Fig. 4A). These findings support the notion that IFNαR signaling within DCs limits mRNA-LNP uptake and expression. Next, we assayed if DC-specific loss of IFNαR could enhance CD8⁺ T cell responses to mRNA-LNPs. We immunized IfnαrcKO and IfnαrWT mice with OVA mRNA-LNP vaccines and measured vaccine-specific CD8⁺ T cell responses 7 days later. IfnαrcKO mice displayed a significant increase in vaccine-specific CD8⁺ specific T cells as compared to IfnαrWT control mice (Supplementary Fig. 4B and Fig. 3C). In addition, IfnαrcKO vaccine-specific CD8⁺ T cells displayed an increased surface expression of PD-1 as compared to IfnαrWT controls (Fig. 3D). Again, to ascertain if this effect was due to an increased ability for DCs to acquire the mRNA-LNP, we immunized Itgaxcre Ifnαrfl/fl (IfnαrcKO) and Ifnαr fl/fl (IfnαrWT) mice with mRNA-LNP incorporated with a fluorescently labeled LNP, as in Fig. 2G. Twenty four hours after immunization with fluorescently labeled mRNA-LNP, splenic DCs from IfnαrcKO mice displayed a significantly higher expression of the fluorescent LNP compared to IfnαrWT control DCs indicating an increased ability to acquire the mRNA-LNP (Fig. 3E). These findings resembled those observed when IFNαR was blocked systemically by the MAR1-5A3 antibody (Fig. 2C–G). Our results show that loss of IFNαR signaling specifically in DCs is sufficient to enhance mRNA-LNP vaccine immune responses through increased mRNA-LNP uptake and expression.
A (Left) Representative flow plot of splenic CD11c+MHCII+ DCs expressing eGFP from IFNαRWT or IFNαRcKO mice. (Right) Percentage of eGFP expression in splenic DCs 24 h post mRNA-LNP immunization (5 μg eGFP mRNA-LNP or 2.5 μg OVA mRNA-LNP IV) in IFNαRWT mice (black, n = 4), or IFNαRcKO mice (red, n = 8). B (Left) Representative flow plot showing CD40+CD86+ splenic DCs after immunization with mRNA-LNP. (Right) Percentage of CD40+CD86+ splenic DCs from mice in Fig. 3A. C (Left) Representative flow plots of SIINFEKL-H2-Kb (OVA) specific CD8⁺ T cells. (Right) Percentage of OVA specific CD8⁺ T cells from IFNαRWT mice (black, n = 17) and IFNαRcKO mice (red, n = 9). 2.5 μg of mRNA-LNP was administered IV. D (Left) Representative flow plot showing PD-1 expression of OVA specific TCRβ+CD8α+ T cells. (Right) PD-1 expression of TCRβ+CD8α+ T cells from mice in Fig. 3C. E (Left) Representative flow plot of splenic CD11c+MHCII+ DCs expressing fluorescent LNP from IFNαRWT or IFNαRcKO mice. (Right) Percentage of fluorescent LNP expression in splenic DCs 24 h post mRNA-LNP immunization (5 μg DiD incorporated LNP eGFP mRNA-LNP or 2.5 μg OVA mRNA-LNP IV) in IFNαRWT mice (black, n = 4), or IFNαRcKO mice (red, n = 7). All bars in (A)–(E) indicate mean. Data from (A), (B), (D), and (E) were compared using an unpaired two-tailed students T test. Data from (C) was compared by a Mann–Whitney test.
Inducing IFN before mRNA-LNP vaccination reduces vaccine efficacy
Previous studies have shown that DCs cease acquiring exogenous antigens after activation21. Based on this, we hypothesized that inducing IFN prior to administering our OVA mRNA-LNP vaccine would suppress the OVA-specific immune response. To test this hypothesis, we immunized mice with PBS or with eGFP mRNA-LNP to induce transient IFN production 24 h before administering the OVA mRNA-LNP vaccine (Fig. 4A). Immunization with eGFP mRNA-LNP 24 h before OVA mRNA-LNP immunization significantly reduced OVA-specific CD8⁺ T cell responses (Fig. 4B). However, when we blocked IFNαR signaling prior to administering both mRNA-LNP vaccines, generation of vaccine OVA-specific CD8+ T cells was restored (Fig. 4B). While surface level expression of PD-1 was not significantly different on vaccine-specific CD8⁺ T cells with or without MAR1-5A3 treatment, there is trend towards higher PD-1 MFI in IFNαR blocked mice (Fig. 4C).
A Experimental timeline for (B) and (C). Mice were injected IP with MAR1-5A3 antibody or PBS on Day −2. On Day −1, mice were immunized eGFP mRNA-LNP or PBS. The following day mice were immunized with OVA mRNA-LNP. Seven days later mouse spleens were analyzed for SIINFEKL-H2-Kb specific CD8⁺ T cells. B (Left) Representative flow plots of SIINFEKL-H2-Kb (OVA) specific CD8⁺ T cells. (Right) Percentage of OVA specific CD8⁺ T cells from C57BL/6J wild-type mice (WT) given only OVA mRNA-LNP (black, n = 5), eGFP mRNA-LNP then 24 h later given OVA mRNA-LNP (E.O) (blue, n = 6), or MAR1-5A3 then 24 h later given eGFP mRNA-LNP followed by OVA mRNA-LNP 24 h after eGFP (red, n = 6) (M.E.O). 2.5 μg of mRNA-LNP was administered IV and 250 μg of MAR1-5A3 was administered IP. C (Left) Representative flow plot showing PD-1 expression of OVA specific TCRβ+CD8α+ T cells. (Right) PD-1 expression of TCRβ+CD8α+ T cells from mice in Fig. 4B. D Experimental timeline for (E) and (F). Mice were injected with eGFP mRNA-LNP or PBS on Day −1. The following day mice were immunized with OVA mRNA-LNP. Seven days later mouse spleens were analyzed for SIINFEKL-H2-Kb specific CD8⁺ T cells. E (Left) Representative flow plots of OVA specific CD8⁺ T cells. (Right) Percentage of OVA specific CD8⁺ T cells from IFNαRWT mice given just OVA mRNA-LNP (black, n = 6), IFNαRWT mice given eGFP mRNA-LNP then 24 h later given OVA mRNA-LNP (blue, n = 6), and IFNαRcKO mice given eGFP mRNA-LNP then 24 h later given OVA mRNA-LNP (red, n = 6). F (Left) Representative flow plot showing PD-1 expression of OVA specific TCRβ+CD8α+ T cells. (Right) PD-1 expression of TCRβ+CD8α+ T cells from mice in Fig. 4E. All bars in (B), (C), (E), and (F) indicate mean. Data from (B) and (E) were compared using an unpaired two-tailed students T test. Data in (C) and (F) were compared with either an unpaired two-tailed students T test or a Mann–Whitney test.
To ascertain if these results were driven by IFN signaling specifically in DCs, we immunized IfnαrcKO and IfnαrWT mice with the eGFP mRNA-LNP vaccine followed by immunization with our OVA mRNA-LNP vaccine (Fig. 4D). As expected, IfnαrWT mice immunized with eGFP mRNA-LNP prior to OVA mRNA-LNP vaccination displayed significantly reduced OVA-specific CD8+ T cell responses compared to IfnαrWT mice immunized with OVA mRNA-LNP vaccine alone (Fig. 4E). In contrast, no decrease in CD8+ T cell response was observed in IfnαrcKO mice immunized with eGFP mRNA-LNP followed by OVA mRNA-LNP vaccines (Fig. 4E). Additionally, IfnαrcKO mice immunized with both eGFP mRNA-LNP and OVA mRNA-LNP vaccines displayed a significant increase in PD-1 expression on OVA vaccine-specific CD8⁺ T cells compared to those from IfnαrWT immunized with both eGFP mRNA-LNP and OVA mRNA-LNP vaccines or OVA mRNA-LNP vaccines alone (Fig. 4F). Thus, immunizing with mRNA-LNP vaccines when systemic IFN is present or recently induced, either due to a prior vaccination or other IFN inducing agent, reduces the effectiveness of the vaccine-specific CD8⁺ T cell response. This effect is likely due to premature DC activation.
Dendritic cells are essential for generating vaccine-specific CD8⁺ T cell responses to mRNA-LNP immunization
Because we found that IFNαR signaling in DCs alters how mRNA-LNPs are acquired and presented, we next wanted to determine whether DCs are the primary APCs responsible for presenting mRNA-LNP-derived antigens. To test this, we used the Itgax-HBEGF/EGFP (CD11c-DTR) mouse model, which allows for an inducible depletion of DCs via expression of the diphtheria toxin receptor (DTR) on CD11c+ cells. Upon administration of diphtheria toxin (DTX), CD11c+ DCs are selectively ablated through toxin-induced cell death (Fig. 5A). We then evaluated whether vaccine-specific CD8⁺ T cells could be generated in the absence of DCs. For this, we administered DTX to CD11c-DTR mice one day prior to immunization with OVA mRNA-LNPs and continued DTX treatment every other day until day 7, when we assessed CD8⁺ T cell responses. CD11c-DTR mice treated with DTX showed a marked reduction in vaccine-specific CD8+ T cells compared to DTX-treated WT mice and vehicle-treated CD11c-DTR mice (Fig. 5B). While we observed a slight increase in PD-1 expression on vaccine-specific DTX-treated WT mice, PD-1 levels remained equivalent amongst the experimental groups (Fig. 5C). These results demonstrate that DCs are essential for the mRNA-LNP vaccine response in our model and are critical for eliciting vaccine-specific CD8⁺ T cell responses.
A (Left) Representative flow plots of splenic MHCII+CD11c+ cells. (Right) Percentage of splenic MHCII+CD11c+ cells in C57BL/6J wild-type mice (WT) not given diphtheria toxin (black, n = 5), WT mice given diphtheria toxin (purple, n = 7), CD11c DTR mice not given diphtheria toxin (blue, n = 4), and CD11c DTR mice given diphtheria toxin (red, n = 7). B (Left) Representative flow plots of SIINFEKL-H2-Kb (OVA) specific CD8⁺ T cells. (Right) Percentage of OVA specific CD8⁺ T cells from WT mice not given diphtheria toxin mice (black, n = 7), WT mice given diphtheria toxin (purple, n = 7), CD11c DTR mice not given diphtheria toxin (blue, n = 8), and CD11c DTR mice given diphtheria toxin (red, n = 7). Diphtheria toxin was administered on days −1, 1, 3, and 5 of mRNA-LNP administration at 4 ng per gram of mouse. Mice were immunized with 2.5 μg of either eGFP or OVA mRNA-LNP and CD8⁺ T cell were measured 7 days later. C (Left) Representative flow plot showing PD-1 expression of OVA specific TCRβ+CD8α+ T cells. (Right) PD-1 expression of TCRβ+CD8α+ T cells from mice in Fig. 5B. All bars in (A–C) indicate mean. Data was compared in (A) with an unpaired two-tailed students T test. Data from (B) and (C) were compared with either an unpaired two-tailed students T test or a Mann–Whitney test.
cDC1s are the major DC subset priming the CD8⁺ T cell mRNA-LNP vaccine immune response
We next wanted to determine if a specific subset of cDC was the major antigen-presenting cell to create OVA vaccine-specific CD8⁺ cells. To assay this, we used an Irf8 + 32 enhancer knockout mouse (Δ32) which lack the conventional dendritic cell type 1 (cDC1) subset35 and the Zeb2 triple enhancer knockout mouse (Δ1 + 2 + 3) which lack the conventional dendritic cell type 2 (cDC2) subset (Fig. 6A, B)36. We immunized Δ32 and Δ1 + 2 + 3 mice with OVA mRNA-LNP vaccines and assayed vaccine-specific CD8⁺ T cell responses 7 days later. At 7 days post immunization, Δ32 mice had an almost complete loss in the ability to produce vaccine-specific CD8⁺ T cells (Fig. 6C). While priming of vaccine-specific CD8+ specific T cells was nearly absent in Δ32 mice, Δ1 + 2 + 3 mice displayed a striking increase in vaccine-specific CD8+ T cells compared to WT controls (Fig. 6C). We also found that the few vaccine-specific CD8+ T cells produced in Δ32 mice displayed significantly lower surface expression of PD-1 compared to both vaccine-induced CD8+ T cells from WT and Δ1 + 2 + 3 mice (Fig. 6D). In summary, our data indicate that the cDC1 subset is largely responsible for driving mRNA-LNP vaccine immune responses.
A Representative flow plots of the cDC1 population indicated by XCR1+Sirpα- and the cDC2 population indicated by XCR1-Sirpα+ in C57BL/6J wild-type (WT) mice (black), Δ1 + 2 + 3 mice (blue), and Δ32 mice (red). B (Left) Data showing the percentage of cDC1 cells in WT, Δ1 + 2 + 3, or Δ32 mice. (Right) Data showing the percentage of cDC2 cells in WT, Δ1 + 2 + 3, or Δ32 mice. All cells are quantified by gating strategies used in Fig. 6A. C (Left) Representative flow plot of SIINFEKL-H2-Kb (OVA) specific CD8⁺ T cells. (Right) Percentage of OVA specific CD8⁺ T cells from WT mice immunized with OVA mRNA-LNP (black, n = 9), Δ1 + 2 + 3 mice immunized with OVA mRNA-LNP (blue, n = 7), and Δ32 mice immunized with OVA mRNA-LNP (red, n = 6) measured 7 days post vaccination. All mice were immunized with 2.5 μg of mRNA-LNP IV. D (Left) Representative flow plot showing PD-1 expression of OVA specific TCRβ+CD8α+ T cells. (Right) PD-1 expression of TCRβ+CD8α+ T cells from mice in Fig. 6C. All bars in (B–D) indicate mean. Data from (B) to (D) were compared using either an unpaired two-tailed students T test or a Mann–Whitney test.
Virus induced responses reduces mRNA-vaccine efficacy
Because IFN inhibits mRNA-LNP vaccine efficacy, we asked if IFN induced during viral infection could inhibit subsequent mRNA-LNP vaccine efficacy. To test this, we used a non-lethal viral infection model of mouse adapted Dengue virus (D220). Although D220 infection in C57BL/6J mice does not lead to weight loss, mice that are infected with this virus can still induce inflammatory cytokines including type I and II interferons as well as TNFα37,38,39. WT B6 mice were injected with MAR1-5A3 antibody or PBS one day prior to inoculation with D220 virus. On the following day (Day 0), mice were immunized with either OVA mRNA-LNP or control eGFP mRNA-LNP. Seven days post-immunization, vaccine-specific CD8⁺ T cell responses were assessed (Fig. 7A). We tracked weight of these mice throughout the experiment and saw no changes between mock infected and infected mice (Fig. 7B). We found that mice infected with D220, had a significant reduction in vaccine-specific CD8⁺ T cells populations compared to mock infected mice (Fig. 7C). Mice injected with MAR1-5A3 antibody had equivalent vaccine-specific CD8⁺ T cell populations when compared to unblocked mice (Fig. 7C). However, IFNαR blockade in D220-infected mice resulted in increased PD-1 expression compared to both uninfected mice and unblocked D220-infected mice (Fig. 7D). Dengue infection is known to produce other inflammatory cytokines such as TNFα, IL-6, and IL-1β37,38,39 which can activate DCs and decrease their ability to acquire the mRNA-LNP vaccine. Thus, inhibiting IFNαR signaling alone in an infection model may not be sufficient to restore the response. However, in contexts where IFN is produced in isolation, such as with mRNA-based vaccines, IFNαR blockade can be effective.
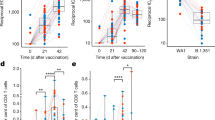
A Experimental timeline for infectious challenge. Mice were injected IP with MAR1-5A3 antibody (αIFNαR) or PBS on Day −2. On Day −1, mice were inoculated with D220 virus. The following day mice were immunized with control eGFP mRNA-LNP or OVA mRNA-LNP. Seven days later mouse spleens were analyzed for SIINFEKL-H2-Kb specific CD8⁺ T cells. B Percent weight change of C57BL/6J wild-type mice (WT) either only given mRNA-LNP (black), mRNA-LNP after being infected with Dengue virus (blue), or given MAR1-5A3 24 h before infection with Dengue virus then immunized with mRNA-LNP a day later (red). C (Left) Representative flow plot of SIINFEKL-H2-Kb (OVA) specific CD8⁺ T cells. (Right) Percentage of OVA specific CD8⁺ T cells from WT mice given OVA mRNA-LNP only (black, n = 4), Dengue infected mice given OVA mRNA-LNP (blue, n = 8), or mice given MAR1-5A3 24 h prior to infection with Dengue virus then immunized with OVA mRNA-LNP a day later (red, n = 8). D (Left) Representative flow plot showing PD-1 expression of OVA specific TCRβ+CD8α+ T cells. (Right) PD-1 expression of TCRβ+CD8α+ T cells from mice in Fig. 7C. MAR1-5A3 was administered at 250 μg IP, mRNA-LNP was administered at 2.5 μg IV, and mice were infected with Dengue virus with 6.5 × 106 FFU IM. Error bars in (B) indicate ± SD. All bars in (C, D) indicate mean. Data in (C) was compared using either an unpaired two-tailed students T test or a Mann–Whitney test. Data in (D) was compared using an unpaired two-tailed students T test.
Vaccine-specific CD8⁺ T cells produced in setting of IFNαR blockade display increased ability to control tumors
It has been shown that lack of IFNαR expression on CD8⁺ T cells abrogates their cytolytic ability13. Therefore, while we see an increase in vaccine-specific CD8⁺ T cells, it is possible that IFNαR blockade could inhibit their cytolytic capacity. To assess whether IFNαR inhibition altered CD8⁺ T cell function, we used an in vivo T cell cytotoxicity assay that monitored the capacity of naturally arising CTLs to kill peptide-pulsed splenocytes labeled with cell trace violet (CTV). WT C57BL/6J IFNαR blocked and unblocked mice immunized with OVA mRNA-LNP vaccines equivalently killed CTVhi-labelled SIINFEKL peptide-pulsed splenocytes but not CTVlo-labelled irrelevant peptide-pulsed splenocytes 10 days following immunization (Fig. 8A). In contrast, WT C57BL/6J mice immunized with eGFP mRNA-LNP were incapable of eliminating both CTVhi-labelled SIINFEKL peptide-pulsed splenocytes and CTVlo-labelled irrelevant peptide-pulsed splenocytes (Fig. 8A). Next, we tested if IFNαR blockade influenced inflammatory cytokine production. We found that blocking IFNαR prior to immunization did not alter production of IFNγ in vaccine-specific CD8⁺ T cells (Fig. 8B). Thus, IFNαR blockade did not diminish the effector capacity of vaccine-elicited CD8⁺ T cells.
A (Left) Representative flow plot showing CTV low (100 nM) labeled cells pulsed with an irrelevant peptide (IAPYYEAL; 1 μM) and CTV high (1 μM) labeled cells pulsed with OVA peptide (SIINFEKL; 1 μM) that were administered 10 days post mRNA-LNP immunization. One million of each CTV labeled cell population was injected IV. C57BL/6J wild-type mice (WT) were given MAR1-5A3 (αIFNαR, red) or not (black) 24 h before immunization with eGFP or OVA mRNA-LNP. MAR1-5A3 was administered at 250 μg IP and mRNA-LNP was administered at 2.5 μg IV. (Right) Percentage of the CTV high labeled population pulsed with OVA in WT mice given MAR1-5A3 (red, n = 5) or not (black, n = 6). B Percentage of IFNγ producing vaccine-specific CD8+ T cells from mice immunized with 2.5 μg of eGFP mRNA-LNP (white, n = 1 and 2), OVA mRNA-LNP with MAR1-5A3 24 h prior to mRNA-LNP (red n = 4 and 8), or not (black, n = 4 and 8). Spleens were isolated 7 days post immunization and cells were stimulated with SIINFEKL-H2-Kb peptide (10 μM) for 5 h (right), or irrelevant peptide as control (left). C Experimental timeline for tumor vaccination in (D). Mice were injected IP with MAR1-5A3 antibody or PBS on Day −1. The following day mice were immunized with control eGFP mRNA-LNP or OVA mRNA-LNP. Fourteen days later mice were implanted with B16-mOVA and followed for tumor growth. D Tumor growth curves of B16-mOVA C57BL/6J WT mice given MAR1-5A3 (red line, n = 8) or not (black line, n = 7) 24 h before immunization with OVA mRNA-LNP. Control mice were administered eGFP mRNA-LNP (dotted line, n = 5). Tumor measurements in (D) are pooled. All bars in (A) and (B) indicate mean. Data in (A) was compared using a Mann–Whitney test. Error bars in (D) indicate S.E.M. Data in (D) was compared using a Two-way ANOVA followed by a Tukey’s multiple comparisons test.
To further investigate the function of vaccine-induced CD8+ T cells generated when IFNαR is inhibited, we used the B16F10 melanoma model engineered to express membrane-bound ovalbumin (B16 mOVA, Supplementary Fig. 5)40. WT B6 mice were injected intraperitoneally with MAR1-5A3 antibody or PBS and then immunized with either OVA or eGFP mRNA-LNP vaccines the following day. Fourteen days later, B16-mOVA tumor cells were implanted subcutaneously into the flank and were followed for tumor growth (Fig. 8C). As expected, mice immunized with the control eGFP mRNA-LNP failed to reject tumors (Fig. 8D). Mice immunized with OVA mRNA-LNP alone displayed delayed outgrowth; some tumors progressed indicating that the vaccine-induced CD8⁺ T cell response was insufficient to maintain tumor control. In contrast, mice treated with IFNαR-blocking antibody showed sustained control of B16-mOVA tumors over the course of the experiment, suggesting that CD8⁺ T cells generated in the absence of IFNαR signaling had an enhanced capacity to control tumor growth (Fig. 8D).
Discussion
Type I interferons are produced upon sensing of viral products such as viral RNA or DNA and induce many antiviral interferon-stimulated genes (ISGs)12. For instance, IFN activate Protein Kinase R (PKR), which phosphorylates eukaryotic initiation factor 2α (eIF2α). This phosphorylation halts cap-dependent translation, leading to a global shutdown of protein synthesis in infected and neighboring cells to prevent viral replication41. Additionally, IFN signaling induces 2’,5’-oligoadenylate synthetase (OAS) and RNase L, which degrade cellular and viral RNA, further suppressing translation41. In addition, it has been shown that an activated DC loses its ability to acquire new material21,23. Because IFN activates DCs and induces a myriad of ISGs inhibiting transcription and translation, APCs that sense IFN could be blunted in their ability to acquire and/or translate mRNA-LNPs. Indeed, APCs that were activated before exposure to FITC-dextran were unable to internalize it21. Similarly, systemic malaria infection severely inhibited antigen cross-presentation by DCs, in agreement with our hypothesis22. On the other hand, when DCs are stimulated by IFN, they upregulate several co-stimulatory molecules that enhance T cell activation. Key costimulatory molecules such as CD80 (B7-1), CD86 (B7-2), and CD40 promote T cell priming, expansion, and cytokine production. Additionally, IFN stimulation increases the expression of MHC class I and II, enhancing antigen presentation to CD8⁺ and CD4⁺ T cells, respectively12,15,42. Together, these co-stimulatory molecules enhance T cell priming and activation, linking innate immune sensing with adaptive immune responses. These contrasting roles of IFN are possibly why many studies seem to show opposing effects on the adaptive immune response when IFN is present. We propose that IFN primarily evolved as a mechanism to inhibit viral replication and dissemination and secondarily within APCs to enhance adaptive immunity. However, the timing of IFN sensing is critical in determining its impact on immune responses. IFN signaling in APCs during antigen acquisition can help limit viral replication while still allowing the APCs to acquire and present antigens. If IFN signaling occurs during antigen uptake, APCs can effectively process and present viral antigens while simultaneously initiating antiviral defenses. However, if IFN is sensed after a period before the virus or antigen is acquired, the APC diverts to inhibiting viral dissemination. This is accomplished by inhibiting additional antigen uptake and shutting down translation ensuring the DC presents material already acquired, thereby preventing the virus from replicating and spreading systemically. Thus, IFN functions in a delicate balance between antiviral defense and immune activation, with its timing shaping the overall effectiveness of the innate antigen presentation response. During vaccination, the primary goal is to maximize the spread and uptake of the vaccine by as many APCs as possible. In this context, IFN signaling within APCs can be detrimental, as it may suppress antigen acquisition and/or mRNA-LNP translation. Here we show that inhibiting IFN signaling enhances the ability of more APCs to take up, translate, and process the vaccine, leading to a more effective immune response. Since mRNA-LNPs are already potent stimulators of APCs through pattern recognition receptor (PRR) activation6,10, the additional immune stimulation provided by IFN signaling is unnecessary and hinders optimal vaccine efficacy as we have shown herein.
We have demonstrated when mRNA-LNPs are targeted to APCs, IFN signaling inhibits adaptive immune responses. However, it should be noted that different formulations of LNPs could potentially target different cell types and locations. Indeed, it has been shown that altering the cationic ratio of mRNA to LNP, targets LNPs to different organs6. Our mRNA-LNP targets the spleen and to APCs in particular. Therefore, inhibiting IFN signaling allows for more mRNA-LNP acquisition. If an mRNA-LNP vaccine were designed to exclusively target fibroblasts or epithelial cells, IFN signaling within DCs may be necessary for recognizing the vaccine as foreign. In this scenario, since DCs would not be directly activated through PRR signaling, they might rely on IFN signaling from mRNA-LNP expressing cells to enhance antigen presentation and stimulate the adaptive immune response. Given that mRNA-LNP vaccines are still a relatively new technology, it is essential to investigate how different LNP formulations are distributed and sensed across various cell types. Understanding this distribution could help refine strategies for modulating IFN signaling, potentially identifying opportunities to inhibit IFN where it may be detrimental or leverage it where it is beneficial.
Our results also suggest that immunization during an ongoing immune response is detrimental. We found that administering a vaccine too soon, either during an ongoing infection or shortly after a different vaccination, significantly diminished the immune response to the new vaccine. In the case of two mRNA-LNP vaccines administered just one day apart, the diminished immune response appeared to be at least partially due to premature IFN activation of DCs, impairing their ability to acquire and express the second vaccine. This was supported by our finding that transient IFNαR blockade rescued the adaptive immune response. However, when Dengue virus infection was followed the next day by mRNA-LNP vaccination, the reduction in vaccine-specific immunity could not be rescued by IFNαR blockade. This suggests that mRNA-LNP immunizations following an infection cannot be rescued by inhibiting IFN signaling alone. Indeed, it has been shown that dengue infection induces a low but systemic TNFα response37, which could activate DCs and prevent them from effectively processing the subsequent mRNA-LNP vaccine. These findings highlight the importance of timing in vaccination strategies and describe a mechanistic hypothesis for the reduced response to vaccines when infected. Administering vaccines too closely together or during an ongoing infection, even if asymptomatic, can significantly impair vaccine efficacy by disrupting antigen presentation and adaptive immune priming. While our results are primarily focused on the CD8+ T cell response, we identified minor changes in the CD4+ T cell response. MAR1-5A3 administration did improve CD4+ T cell response by percentage of TCRβ+CD4+CXCR5+PD-1+B220- T follicular helper (Tfh) cells (Supplementary Fig. 6). Our model proved to mainly be a CD8+ T cell effect which still demonstrates an important effect that can be used for mRNA-LNP vaccine development.
Finally, our results demonstrate that transient blockade of IFNαR signaling does not compromise, but enhances, the cytotoxic capacity of the vaccine-specific CD8+ T cell response, contrary to previously published reports10,13. When vaccinated mice were injected with vaccine-specific peptide-pulsed splenocytes, they exhibited cytotoxic T cell responses that were comparable to those of controls. Additionally, IFNαR blocked vaccine-specific CD8+ T cells displayed enhanced protection against tumor outgrowth. Thus, in the context of mRNA-LNP vaccination against patient-specific neoantigens, inhibiting IFNαR signaling transiently could potentially augment neoantigen-elicited CD8+ T cell responses and lead to better tumor control. Altogether, our findings reveal that IFN signaling plays a context and timing-dependent role in shaping the immune response to mRNA-LNP vaccines. These insights provide a compelling rationale for optimizing vaccine schedules and formulation strategies to transiently modulate innate signaling pathways, particularly IFNαR, to maximize antigen acquisition, mRNA expression, T cell priming, and therapeutic efficacy.
Methods
Study design
This study aimed to determine the role of type I interferon signaling on DCs in the process of creating an adaptive immune response to mRNA-LNP vaccines. We hypothesized that early IFN signaling within DCs impairs their ability to acquire new antigenic material, thereby limiting antigen presentation and ultimately leading to a diminished adaptive immune response. We used a combination of in vitro experiments using bone marrow-derived dendritic cells as well as in vivo experiments using a variety of mouse models that are described in each respective results section. We inhibited type I interferon signaling in various ways as well as administering mRNA-LNP to cells and mice to then measure adaptive immune responses by flow cytometry. This was used to determine the ability to acquire mRNA-LNP and create a strong adaptive immune response to the antigen in question. No randomization was done, and blinding was not necessary as most data was quantified by objective readouts by flow cytometry. Sample sizes were determined by previous experiments, and the number of biological replicates are indicated by “n” in each figure legend. Each experiment was replicated at least once with consistent results. Outliers were defined by a ROUT test with Q = 1%. One outlier was determined and excluded in the study under Fig. 4 within the OVA group.
Mice
WT C57BL/6J (Jax stock no. 000664), CD11cCre mice (stock no. 008068), IFNαRfl/fl mice (stock no. 028256), Zeb2 triple enhancer mutant mice (Δ 1, 2, 3; stock no. 037704), and CD11c DTR mice (stock no. 004509) were purchased from The Jackson Laboratory and bred in-house. Irf8+32–/– (Δ32) mice, which are homozygous for the deletion of a downstream enhancer element of Irf8, were gifted by Dr. Kenneth Murphy at Washington University in St. Louis and described previously32. All mice were housed under a 12-h dark/light cycle, and housing was maintained at an ambient temperature of 72 °F. Mice were age-matched and sex-matched and between 8 and 16 weeks of age when used for experiments. Mice were humanely euthanized by CO2 anesthesia followed by cervical dislocation in accordance with the Guide for the Care and Use of Laboratory Animals of the NIH under approval by the Institutional Animal Care and Use Committee (IACUC) at Saint Louis University School of Medicine (Assurance Number: D16-00141).
mRNA and LNP generation
The DNA plasmid used for in vitro transcription was based on a previously published Zika virus vaccine design (PMC5388441). Briefly, the gene of interest (GOI) was flanked by a 5′ untranslated region (UTR; 5′-GGGAAATAAGAGAGAAAAGAAGAGTAAGAAGAAATATAAGAGCCACC-3′) and a 3′ UTR (5′-TAATAGGCTGGAGCCTCGGTGGCCATGCTTCTTGCCCCTTGGGCCTCCCCCCAGCCCCTCCTCCCCTTCCTGCACCCGTACCCCCGTGGTCTTTGAATAAAGTCTGA-3′), followed by a poly(A) tail consisting of 120 adenosines with a single guanosine interruption in the middle. The GOI was cloned into a multiple cloning site without a secretion signal sequence. The expression vector includes the T7 Promoter, 5’ and 3’ untranslated region (UTR) flanking the ZsGreen1, eGFP, or cytosolic Ovalbumin (OVA), and the AG CleanCap nucleotide sequence after the TATA box sequence for downstream in vitro mRNA transcription. The plasmid was transformed into DH5α competent E. coli cells (NEB Cat No. C2988J) and plated onto Ampicillin agar plates for resistant colonies to grow. Colonies were harvested and checked for correct insert size by colony PCR and sequenced. Correct plasmids were grown in 250 mL LB ampicillin cultures and purified via endotoxin free plasmid maxi kits (Qiagen Cat No. 12362).
After plasmid purification, the plasmid was digested using the restriction enzymes BglII and XhoI at 37 °C overnight. The insert was isolated via gel purification using Gel/PCR DNA fragments extraction kits (IBI Scientific cat IB47020). These fragments were used for in vitro mRNA transcription via mRNA Kit with CleanCap Reagent AG (NEB Cat No. E2080S) utilizing CleanCap for the 5’ cap and pseudo-uridine in place of uridine as per manufacturers protocol, and left overnight at 37 °C. Encapsulation of the mRNA was performed on a NanoAssemblr Ignite Instrument (Cat No. NIN0001). The mRNA was dissolved in PNI Formulation Buffer (Cat No. NWW0043) and run through a NanoAssemblr Ignite NxGen Cartridge (Cat No. NIN0061) with the lipid nanoparticle GenVoy ILM or GenVoy-ILM with Dye formulations (Cat No. NWW0042) at a flow ratio of 3:1 and a total flow rate of 12 mL/min to generate the LNP encapsulated mRNA vaccines. For downflow processing, the size of the LNP particles was determined by polydispersity index (PDI) by Dynamic Light Scattering (DLS). The mRNA-LNPs were subsequently concentrated and purified by diluting the samples in PBS and filtrating via centrifugal filtration using a filtration tube (Millipore Sigma Amicon Ultra-15 Cat No. C7715). Measurement of the mRNA concentration within the LNP and encapsulation efficiency was performed using a Quant-IT RiboGreen Assay (Thermo Fisher Scientific Cat No. R11490).
In vitro DC cultures
Bone marrow (BM) was harvested from femurs and tibias from WT C57BL/6J mice. Bones were crushed with a mortar and pestle in Iscove’s modified Dulbecco’s media (IMDM) supplemented with 10% FCS (Gibco) and were passed through 70-µm filters. Red blood cells were lysed using ammonium chloride-potassium bicarbonate (ACK) buffer. After RBC lysis, cells were brought up in I10F, counted via the Countess-3 (Fisher) cell counter. Cells were then plated at 105 cells/mL in I10F supplemented with 10% SCF conditioned media for 3 days. Cells were then washed in I10F and replated and cultured in I10F supplemented with 5% Flt3L-Fc conditioned media for 8 days. On day 6 of Flt3L-Fc culture, some DCs were cultured with recombinant IFNα at 103 U/mL for 16 h prior to receiving mRNA-LNP. On Day 7, DCs were cultured with 2.5 μg ZsGreen or eGFP mRNA-LNP with or without 10 μg MAR1-5A3 anti-IFNαR blocking antibody and cultured for 24 h. At 24 h following mRNA-LNP addition, cells were harvested and stained with fluorescently labeled antibodies at 4 °C in magnetic-activated cell-sorting (MACS) buffer (PBS + 0.5% BSA + 2 mM EDTA) in the presence of 10% Fc block supernatant from 2.4G2 cells and analyzed via flow cytometry.
CD8⁺ T cell tetramer staining
Spleens were harvested at different timepoints indicated by each experiment, respectively. Whole spleens were mashed and digested in collagenase B (250 µg/mL) and DNase (10 µg/mL) in Iscove’s modified Dulbecco’s media (IMDM) supplemented with 10% FCS (Gibco) for 45 min at 37 °C with shaking. Red blood cells were lysed with ammonium chloride–potassium bicarbonate (ACK) lysis buffer. Cells were subsequently counted with a Countess 3 (Invitrogen). SIINFEKL-H2-Kb biotinylated monomers were purchased from the NIH. Then, the peptide–MHC multimers were incubated with PE- and APC-conjugated streptavidin (SA) at a concentration of 1:9 for 30 min at 4 °C protected from light in separate reactions. SA-labelled tetramers were then incubated with 25 µM D-biotin for 20 min at 4 °C protected from light to quench free fluorochrome-labelled SA. 3 × 106 splenocytes were incubated in MACS buffer containing the Fc-blocking antibody produced from the 2.4G2 cell line for 5 min at 4 °C. Fluorochrome-conjugated tetramers were added to the splenocytes at a concentration of 1:100 and incubated at 4 °C for 30 min. Surface antibodies were added without washing at a final concentration of 1:200 and stained for another 30 min at 4 °C.
Antibodies and flow cytometry
Flow cytometry was completed on a FACS CantoII (BD) and analyzed using FlowJo analysis software (Tree Star). Staining was performed at 4 °C in the presence of Fc block (10% 2.4G2 supernatant) in MACS buffer. The following antibodies were used; from Biolegend: KLRG1 (clone: 2F1/KLRG1), CD45.1 (clone: A20), MHCII (clone: M5/114.15.2), IFNαR (clone: MAR1-5A3), CD40 (clone: FGK45), Vα2 (clone: B20.1), Sirpα (clone: P84), CD45.2 (clone: 104), TCRβ (clone: H57-597), PD-1 (clone: 29F.1A12), CD86 (clone: GL-1), Ly6C (clone: HK1.4), CD4 (clone: Rm4-5), CD44 (clone: IM7), CD11c (clone: N418), XCR1 (clone: ZET), B220 (clone: RA3-6B2), CD8α (clone: 53-6.7), IL-7Rα (clone: A7R34), F4/80 (clone: BM8), IFNγ (clone: XMG1.2), CXCR5 (clone: L138D7), PE streptavidin (cat: 405204), and APC streptavidin (cat: 405207). The anti-OVA antibody was purchased from Millipore (Millipore AB1225).
Mice immunizations
The blocking αIFNαR antibody (MAR1-5A3; Leinco) was administered to mice at 250 μg intraperitoneally (IP). Blocking IFNαR with MAR1-5A3 occurred one day before immunizing with mRNA-LNP. OVA mRNA-LNP was immunized in mice at 2.5 μg intravenously (IV) through the tail vein. ZsGreen or eGFP mRNA-LNP was immunized at either 5 μg or 2.5 μg intravenously through the tail vein indicated by each experiment, respectively. We labeled all experiments that used ZsGreen or eGFP mRNA-LNP as just eGFP mRNA-LNP to conserve space. There was no difference seen between these different constructs.
ELISA
For IFNα ELISA, whole serum was isolated from mice at hour 0 before immunization as well as hours 8 and 24 post mRNA-LNP vaccination. Whole serum was stored at −80 °C and was analyzed for IFNα 2 and 4 with an IFN alpha ELISA KIT (Invitrogen: Cat. BMS6027).
Dengue infection
WT C57BL/6J mice were used in the Dengue infection studies. Mice were administered with 250 μg MAR1-5A3 antibody intraperitoneally, one day before being infected with a mouse adapted dengue virus strain grown in C6/26 cells (D220, a gift from Eva Harris, UC Berkeley)43. Infections were performed intramuscularly into the hind leg of the mouse with 6.5 × 106 FFU per mouse. One day post-infection, mice were immunized with mRNA-LNP at 2.5 μg intravenously. Body weight was recorded every other day until the mice were sacrificed on day 7 post mRNA-LNP immunization.
Tumor line and growth experiment
The tumor line used was the B16F10 melanoma membrane expressing ovalbumin (B16-mOVA). The B16-mOVA line was engineered to express membrane ovalbumin (mOVA) using a MSCV-mOVA-IRES-Thy1.1 vector as previously described41. B16F10 tumor cells were retrovirally transduced with a pMSCV vector expressing mOVA (B16mOVA; Fig. 8). Clone 2 was selected by expression of surface OVA (Millipore AB1225) using flow cytometry. WT C57BL/6J mice were either given 250 μg MAR1-5A3 antibody intraperitoneally or PBS then immunized with 2.5 μg of OVA mRNA-LNP intravenously a day later. Control mice were immunized with 2.5 μg of eGFP mRNA-LNP. Fourteen days post immunization, mice were subcutaneously injected with 106 B16mOVA tumor cells into the flank. Tumor growth was measured with a caliper, and tumor area was calculated by the multiplication of two perpendicular diameters. Growth measurements were taken on days 0, 4, 7, 11, 14, 18, and 21 post inoculation. In accordance with our IACUC-approved protocol, maximal tumor diameter was 20 mm in one direction, and in no experiments was this limit exceeded.
Intracellular interferon gamma stain
Spleens were harvested 7 days post mRNA-LNP immunization. Whole spleens were mashed and digested in collagenase B (250 µg/mL) and DNase (10 µg/mL) in Iscove’s modified Dulbecco’s media (IMDM) supplemented with 10% FCS (Gibco) for 45 min at 37 °C with shaking. Red blood cells were lysed with ammonium chloride–potassium bicarbonate (ACK) lysis buffer. Cells were subsequently counted with a Countess 3 (Invitrogen). Cells were then incubated with Brefeldin A (BFA) (10 μg/mL) and SIINFEKL-H2-Kb (10 μM), or BFA with irrelevant peptide for 5 h at 37 °C. Cells were then stained normally at 4 °C for 30 min for surface antibodies. Cells were then washed before being fixed with the FoxP3 Fix/Perm Buffer set (Biolegend 421403) for 20 min at room temperature. Then cells were stained with intracellular antibody IFNγ in perm buffer at 4 °C for 1 h. Cells were then washed once with perm buffer and once with MACS buffer before being used for flow cytometry.
Statistical analysis
Statistical analysis was performed using GraphPad Prism software version 10. Unless otherwise noted, a one-way ANOVA, Unpaired two-tailed Students T test, or Mann–Whitney test was used to determine significant differences between samples, and all center values correspond to the mean. P ≤ 0.05 was considered statistically significant. Investigators were blinded to the treatments of the mice during sample preparation and data collection.
Data availability
All data generated or analyzed during this study are included in this published article and its supplementary information files.
References
Arevalo, C. P. et al. A multivalent nucleoside-modified mRNA vaccine against all known influenza virus subtypes. Science 378, 899–904 (2022).
Xie, Z. et al. mRNA-LNP HIV-1 trimer boosters elicit precursors to broad neutralizing antibodies. Science 384, eadk0582 (2024).
Barbier, A. J., Jiang, A. Y., Zhang, P., Wooster, R. & Anderson, D. G. The clinical progress of mRNA vaccines and immunotherapies. Nat. Biotechnol. 40, 840–854 (2022).
Rojas, L. A. et al. Personalized RNA neoantigen vaccines stimulate T cells in pancreatic cancer. Nature 618, 144–150 (2023).
Liu, C. et al. mRNA-based cancer therapeutics. Nat. Rev. Cancer 23, 526–543 (2023).
Kranz, L. M. et al. Systemic RNA delivery to dendritic cells exploits antiviral defence for cancer immunotherapy. Nature 534, 396–401 (2016).
Sethna, Z. et al. RNA neoantigen vaccines prime long-lived CD8+ T cells in pancreatic cancer. Nature 639, 1042–1051 (2025).
Sittplangkoon, C. et al. mRNA vaccine with unmodified uridine induces robust type I interferon-dependent anti-tumor immunity in a melanoma model. Front. Immunol. 13, 983000 (2022).
Arunachalam, P. S. et al. Systems vaccinology of the BNT162b2 mRNA vaccine in humans. Nature 596, 410–416 (2021).
Li, C. et al. Mechanisms of innate and adaptive immunity to the Pfizer-BioNTech BNT162b2 vaccine. Nat. Immunol. 23, 543–555 (2022).
Kim, S. et al. Innate immune responses against mRNA vaccine promote cellular immunity through IFN-β at the injection site. Nat. Commun. 15, 7226 (2024).
McNab, F., Mayer-Barber, K., Sher, A., Wack, A. & O’Garra, A. Type I interferons in infectious disease. Nat. Rev. Immunol. 15, 87–103 (2015).
Kolumam, G. A., Thomas, S., Thompson, L. J., Sprent, J. & Murali-Krishna, K. Type I interferons act directly on CD8 T cells to allow clonal expansion and memory formation in response to viral infection. J. Exp. Med. 202, 637–650 (2005).
Sikora, A. G. et al. IFN-α enhances peptide vaccine-induced CD8+ T cell numbers, effector function, and antitumor activity. J. Immunol. 182, 7398–7407 (2009).
Ngoi, S. M., Tovey, M. G. & Vella, A. T. Targeting Poly I:C to the TLR3-independent pathway boosts effector CD8 T cell differentiation through IFNα/β. J. Immunol. 181, 7670–7680 (2008).
Hall, J. C. & Rosen, A. Type I interferons: crucial participants in disease amplification in autoimmunity. Nat. Rev. Rheumatol. 6, 40–49 (2010).
Su, A. I. et al. Genomic analysis of the host response to hepatitis C virus infection. Proc. Natl. Acad. Sci. USA 99, 15669–15674 (2002).
O’Garra, A. et al. The immune response in tuberculosis. Annu. Rev. Immunol. 31, 475–527 (2013).
McNab, F. W. et al. TPL-2-ERK1/2 signaling promotes host resistance against intracellular bacterial infection by negative regulation of type I IFN production. J. Immunol. 191, 1732–1743 (2013).
Berry, M. P. R. et al. An interferon-inducible neutrophil-driven blood transcriptional signature in human tuberculosis. Nature 466, 973–977 (2010).
Sallusto, F., Cella, M., Danieli, C. & Lanzavecchia, A. Dendritic cells use macropinocytosis and the mannose receptor to concentrate macromolecules in the major histocompatibility complex class II compartment: downregulation by cytokines and bacterial products. J. Exp. Med. 182, 389–400 (1995).
Wilson, N. S. et al. Systemic activation of dendritic cells by Toll-like receptor ligands or malaria infection impairs cross-presentation and antiviral immunity. Nat. Immunol. 7, 165–172 (2006).
Theisen, D. J. et al. Batf3-dependent genes control tumor rejection induced by dendritic cells independently of cross-presentation. Cancer Immunol. Res. 7, 29–39 (2019).
Sheehan, K. C. F. et al. Blocking monoclonal antibodies specific for mouse IFN-alpha/beta receptor subunit 1 (IFNAR-1) from mice immunized by in vivo hydrodynamic transfection. J. Interferon Cytokine Res. 26, 804–819 (2006).
Palacio, N. et al. Early type I IFN blockade improves the efficacy of viral vaccines. J. Exp. Med. 217, e20191220 (2020).
Bullock, C. B. et al. Type I interferon signaling in dendritic cells limits direct antigen presentation and CD8+ T cell responses against an arthritogenic alphavirus. mBio. 15, e02930-24 (2024).
Broomfield, B. J. et al. Transient inhibition of type I interferon enhances CD8+ T cell stemness and vaccine protection. J. Exp. Med 222, e20241148 (2025).
Ou, F. et al. Enhanced in vitro type 1 conventional dendritic cell generation via the recruitment of hematopoietic stem cells and early progenitors by Kit ligand. Eur. J. Immunol. 53, e2250201 (2023).
Ryman, K. D. et al. Sindbis virus translation is inhibited by a PKR/RNase L-independent effector induced by alpha/beta interferon priming of dendritic cells. J. Virol. 79, 1487–1499 (2005).
Ahn, E. et al. Role of PD-1 during effector CD8 T cell differentiation. Proc. Natl. Acad. Sci. USA 115, 4749–4754 (2018).
Zahm, C. D., Colluru, V. T. & McNeel, D. G. Vaccination with high-affinity epitopes impairs antitumor efficacy by increasing PD-1 expression on CD8+ T cells. Cancer Immunol. Res. 5, 630–641 (2017).
Zheng, L., Bandara, S. R., Tan, Z. & Leal, C. Lipid nanoparticle topology regulates endosomal escape and delivery of RNA to the cytoplasm. Proc. Natl. Acad. Sci. USA 120, e2301067120 (2023).
Caton, M. L., Smith-Raska, M. R. & Reizis, B. Notch-RBP-J signaling controls the homeostasis of CD8- dendritic cells in the spleen. J. Exp. Med. 204, 1653–1664 (2007).
Prigge, J. R. et al. Type-I-IFNs act upon hematopoietic progenitors to protect and maintain hematopoiesis during Pneumocystis lung infection in mice. J. Immunol. 195, 5347–5357 (2015).
Durai, V. et al. Cryptic activation of an Irf8 enhancer governs cDC1 fate specification. Nat. Immunol. 20, 1161–1173 (2019).
Liu, T.-T. et al. Ablation of cDC2 development by triple mutations within the Zeb2 enhancer. Nature https://doi.org/10.1038/s41586-022-04866-z (2022).
Shresta, S., Sharar, K. L., Prigozhin, D. M., Beatty, P. R. & Harris, E. Murine model for dengue virus-induced lethal disease with increased vascular permeability. J. Virol. 80, 10208–10217 (2006).
Chen, H.-W. et al. The roles of IRF-3 and IRF-7 in innate antiviral immunity against dengue virus. J. Immunol. 191, 4194–4201 (2013).
Chen, H.-C., Hofman, F. M., Kung, J. T., Lin, Y.-D. & Wu-Hsieh, B. A. Both virus and tumor necrosis factor alpha are critical for endothelium damage in a mouse model of dengue virus-induced hemorrhage. J. Virol. 81, 5518–5526 (2007).
Ferris, S. T. et al. cDC1 prime and are licensed by CD4+ T cells to induce anti-tumour immunity. Nature https://doi.org/10.1038/s41586-020-2611-3 (2020).
Yan, N. & Chen, Z. J. Intrinsic antiviral immunity. Nat. Immunol. 13, 214–222 (2012).
Busselaar, J., Sijbranda, M. & Borst, J. The importance of type I interferon in orchestrating the cytotoxic T-cell response to cancer. Immunol. Lett. 270, 106938 (2024).
Orozco, S. et al. Characterization of a model of lethal dengue virus 2 infection in C57BL/6 mice deficient in the alpha/beta interferon receptor. J. Gen. Virol. 93, 2152–2157 (2012).
Acknowledgements
Stephen T. Ferris is supported by the President’s Research Fund of Saint Louis University School of Medicine (S.F.). Elise Alspach, PhD, is supported by a Research Scholar Grant, RSG-24-1251974-01-IBCD, from the American Cancer Society (https://doi.org/10.53354/ACS.RSG-24-1251974-01-IBCD.pc.gr.193734), and is the recipient of a Cancer Research Institute CLIP Grant (CRI5509). We thank the NIH Tetramer Core Facility (NIH Contract 75N93020D00005 and RRID:SCR_026557) for providing H2-Kb chicken ova 257-264 SIINFEKL Monomer.
Author information
Authors and Affiliations
Contributions
Conceptualization: S.T.F., T.A.L., and A.D.; Methodology: S.T.F., T.A.L., E.A., and L.V.T.; Investigation: T.A.L., S.T.F., S.B., J.A.C., Y.D., and W.G.; Visualization: T.A.L. and S.T.F.; Funding acquisition: S.T.F.; Project administration: S.T.F. and R.J.D.; Supervision: S.T.F.; Writing—original draft: T.A.L. and S.T.F.; Writing—review & editing: T.A.L., S.T.F., E.A., L.V.T., and W.G.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Lobb, T.A., Dickson, A., Guo, W. et al. Type I interferon restricts mRNA vaccine efficacy through suppression of antigen uptake in cDCs. npj Vaccines 11, 41 (2026). https://doi.org/10.1038/s41541-025-01362-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41541-025-01362-z