Abstract
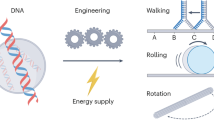
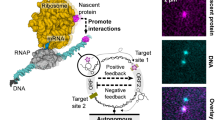
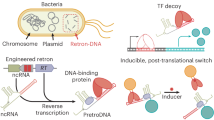
Biological transcription uses dynamic machinery modulated by transcription factors and auxiliary environmental cues to control multiple biological processes. Misregulation of the transcription machinery leads to diverse genetic disorders and diseases. Here we discuss the application of DNA nanostructures and circuits in developing synthetic in vitro transcription frameworks that mimic dynamic features, such as switchable blockage of transcription by topological barriers, transcription machineries revealing transient dissipative kinetics, and bistable programs or oscillatory transcription circuits driven by feedback loops, paving the way to exploring and validating mechanisms in native transcription and their potential biological applications. Possible applications of the transcription frameworks for sensing, and future perspectives for autonomous therapeutics and the design of artificial cells, are discussed.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Simon, I. et al. Serial regulation of transcriptional regulators in the yeast cell cycle. Cell 106, 697–708 (2001).
Accili, D. & Arden, K. C. FoxOs at the crossroads of cellular metabolism, differentiation, and transformation. Cell 117, 421–426 (2004).
Bain, G. et al. E2A proteins are required for proper B cell development and initiation of immunoglobulin gene rearrangements. Cell 79, 885–892 (1994).
Hager, G. L., McNally, J. G. & Misteli, T. Transcription dynamics. Mol. Cell. 35, 741–753 (2009).
De Nadal, E., Ammerer, G. & Posas, F. Controlling gene expression in response to stress. Nat. Rev. Genet. 12, 833–845 (2011).
Sztal, T. E. & Stainier, D. Y. R. Transcriptional adaptation: a mechanism underlying genetic robustness. Development 147, dev186452 (2020).
Vaquerizas, J. M., Kummerfeld, S. K., Teichmann, S. A. & Luscombe, N. M. A census of human transcription factors: function, expression and evolution. Nat. Rev. Genet. 10, 252–263 (2009).
Lenstra, T. L., Rodriguez, J., Chen, H. & Larson, D. R. Transcription dynamics in living cells. Annu. Rev. Biophys. 45, 25–47 (2016).
Weber, C. M. & Henikoff, S. Histone variants: dynamic punctuation in transcription. Genes Dev. 28, 672–682 (2014).
Fuxreiter, M. et al. Malleable machines take shape in eukaryotic transcriptional regulation. Nat. Chem. Biol. 4, 728–737 (2008).
Dauba, A., Braikia, F.-Z., Oudinet, C. & Khamlichi, A. A. Interleukin 7 regulates switch transcription in developing B cells. Cell. Mol. Immunol. 18, 776–778 (2021).
Rodionov, D. A. Comparative genomic reconstruction of transcriptional regulatory networks in bacteria. Chem. Rev. 107, 3467–3497 (2007).
Berger, S. L. The complex language of chromatin regulation during transcription. Nature 447, 407–412 (2007).
Spitz, F. & Furlong, E. E. M. Transcription factors: from enhancer binding to developmental control. Nat. Rev. Genet. 13, 613–626 (2012).
Petrykowska, H. M., Vockley, C. M. & Elnitski, L. Detection and characterization of silencers and enhancer-blockers in the greater CFTR locus. Genome Res. 18, 1238–1246 (2008).
Gaszner, M. & Felsenfeld, G. Insulators: exploiting transcriptional and epigenetic mechanisms. Nat. Rev. Genet. 7, 703–713 (2006).
Travers, A. Transcriptional switches: the role of mass action. Phys. Life Rev. 1, 57–69 (2004).
Nelson, D. E., Sée, V., Nelson, G. & White, M. R. Oscillations in transcription factor dynamics: a new way to control gene expression. Biochem. Soc. Trans. 32, 1090–1092 (2004).
Fang, X. et al. Cell fate potentials and switching kinetics uncovered in a classic bistable genetic switch. Nat. Commun. 9, 2787 (2018).
Lee, T. I. & Young, R. A. Transcriptional regulation and its misregulation in disease. Cell 152, 1237–1251 (2013).
Ament, S. A. et al. Transcriptional regulatory networks underlying gene expression changes in Huntington’s disease. Mol. Syst. Biol. 14, e7435 (2018).
Boyadjiev, S. A. & Jabs, E. W. Online Mendelian Inheritance in Man (OMIM) as a knowledgebase for human developmental disorders. Clin. Genet. 57, 253–266 (2000).
Pandelakis, M., Delgado, E. & Ebrahimkhani, M. R. CRISPR-based synthetic transcription factors in vivo: the future of therapeutic cellular programming. Cell Syst. 10, 1–14 (2020).
Toniatti, C., Bujard, H., Cortese, R. & Ciliberto, G. Gene therapy progress and prospects: transcription regulatory systems. Gene Ther. 11, 649–657 (2004).
Villicaña, C., Cruz, G. & Zurita, M. The basal transcription machinery as a target for cancer therapy. Cancer Cell Int. 14, 18 (2014).
Elowitz, M. B. & Leibler, S. A synthetic oscillatory network of transcriptional regulators. Nature 403, 335–338 (2000).
Atkinson, M. R., Savageau, M. A., Myers, J. T. & Ninfa, A. J. Development of genetic circuitry exhibiting toggle switch or oscillatory behavior in Escherichia coli. Cell 113, 597–607 (2003).
Friedland, A. E. et al. Synthetic gene networks that count. Science 324, 1199–1202 (2009).
Saw, P. E. & Song, E. Advancements in clinical RNA therapeutics: present developments and prospective outlooks. Cell Rep. Med. 5, 101555 (2024).
Fabrini, G. et al. Co-transcriptional production of programmable RNA condensates and synthetic organelles. Nat. Nanotechnol. 19, 1665–1673 (2024).
Li, H. et al. RNA as a stable polymer to build controllable and defined nanostructures for material and biomedical applications. Nano Today 10, 631–655 (2015).
Lopez, R., Wang, R. & Seelig, G. A molecular multi-gene classifier for disease diagnostics. Nat. Chem. 10, 746–754 (2018).
Kaminski, M. M., Abudayyeh, O. O., Gootenberg, J. S., Zhang, F. & Collins, J. J. CRISPR-based diagnostics. Nat. Biomed. Eng. 5, 643–656 (2021).
Zhang, D. Y. & Winfree, E. Control of DNA strand displacement kinetics using toehold exchange. J. Am. Chem. Soc. 131, 17303–17314 (2009).
Zhang, D. Y., Turberfield, A. J., Yurke, B. & Winfree, E. Engineering entropy-driven reactions and networks catalyzed by DNA. Science 318, 1121–1125 (2007).
Zhang, D. Y. & Seelig, G. Dynamic DNA nanotechnology using strand-displacement reactions. Nat. Chem. 3, 103–113 (2011).
Wang, F., Liu, X. & Willner, I. DNA switches: from principles to applications. Angew. Chem. Int. Ed. 54, 1098–1129 (2015).
Leroy, J. L., Guéron, M., Mergny, J. L. & Hélène, C. Intramolecular folding of a fragment of the cytosine-rich strand of telomeric DNA into an i-motif. Nucleic Acids Res. 22, 1600–1606 (1994).
Nonin, S. & Leroy, J. L. Structure and conversion kinetics of a bi-stable DNA i-motif: broken symmetry in the [d(5mCCTCC)]4 tetramer. J. Mol. Biol. 261, 399–414 (1996).
Miyake, Y. et al. MercuryII-mediated formation of thymine−HgII−thymine base pairs in DNA duplexes. J. Am. Chem. Soc. 128, 2172–2173 (2006).
O’Hagan, M. P. et al. Photocleavable ortho-nitrobenzyl-protected DNA architectures and their applications. Chem. Rev. 123, 6839–6887 (2023).
Kamiya, Y. & Asanuma, H. Light-driven DNA nanomachine with a photoresponsive molecular engine. Acc. Chem. Res. 47, 1663–1672 (2014).
Lubbe, A. S., Szymanski, W. & Feringa, B. L. Recent developments in reversible photoregulation of oligonucleotide structure and function. Chem. Soc. Rev. 46, 1052–1079 (2017).
Montagne, K., Plasson, R., Sakai, Y., Fujii, T. & Rondelez, Y. Programming an in vitro DNA oscillator using a molecular networking strategy. Mol. Syst. Biol. 7, 466 (2011).
Baccouche, A., Montagne, K., Padirac, A., Fujii, T. & Rondelez, Y. Dynamic DNA-toolbox reaction circuits: a walkthrough. Methods 67, 234–249 (2014).
Seeman, N. C. & Sleiman, H. F. DNA nanotechnology. Nat. Rev. Mater. 3, 17068 (2017).
Goodman, R. P. et al. Reconfigurable, braced, three-dimensional DNA nanostructures. Nat. Nanotechnol. 3, 93–96 (2008).
Wang, Z.-G., Elbaz, J. & Willner, I. DNA machines: bipedal walker and stepper. Nano Lett. 11, 304–309 (2011).
Bath, J. & Turberfield, A. J. DNA nanomachines. Nat. Nanotechnol. 2, 275–284 (2007).
Yurke, B., Turberfield, A. J., Mills, A. P., Simmel, F. C. & Neumann, J. L. A DNA-fuelled molecular machine made of DNA. Nature 406, 605–608 (2000).
Omabegho, T., Sha, R. & Seeman, N. C. A bipedal DNA Brownian motor with coordinated legs. Science 324, 67–71 (2009).
Lu, C.-H., Cecconello, A., Elbaz, J., Credi, A. & Willner, I. A three-station DNA catenane rotary motor with controlled directionality. Nano Lett. 13, 2303–2308 (2013).
Lohmann, F., Ackermann, D. & Famulok, M. Reversible light switch for macrocycle mobility in a DNA rotaxane. J. Am. Chem. Soc. 134, 11884–11887 (2012).
Lohmann, F., Weigandt, J., Valero, J. & Famulok, M. Logic gating by macrocycle displacement using a double-stranded DNA [3]rotaxane shuttle. Angew. Chem. Int. Ed. 53, 10372–10376 (2014).
Yang, Y., Endo, M., Hidaka, K. & Sugiyama, H. Photo-controllable DNA origami nanostructures assembling into predesigned multiorientational patterns. J. Am. Chem. Soc. 134, 20645–20653 (2012).
Wu, N. & Willner, I. pH-Stimulated reconfiguration and structural isomerization of origami dimer and trimer systems. Nano Lett. 16, 6650–6655 (2016).
Wang, J. et al. Active generation of nanoholes in DNA origami scaffolds for programmed catalysis in nanocavities. Nat. Commun. 10, 4963 (2019).
Goodman, R. P. et al. Rapid chiral assembly of rigid DNA building blocks for molecular nanofabrication. Science 310, 1661–1665 (2005).
Qian, L., Winfree, E. & Bruck, J. Neural network computation with DNA strand displacement cascades. Nature 475, 368–372 (2011).
Qian, L. & Winfree, E. Scaling up digital circuit computation with DNA strand displacement cascades. Science 332, 1196–1201 (2011).
Lv, H. et al. DNA-based programmable gate arrays for general-purpose DNA computing. Nature 622, 292–300 (2023).
Okumura, S. et al. Nonlinear decision-making with enzymatic neural networks. Nature 610, 496–501 (2022).
Yue, L., Wang, S., Zhou, Z. & Willner, I. Nucleic acid based constitutional dynamic networks: from basic principles to applications. J. Am. Chem. Soc. 142, 21577–21594 (2020).
Wang, S. et al. Controlling the catalytic functions of DNAzymes within constitutional dynamic networks of DNA nanostructures. J. Am. Chem. Soc. 139, 9662–9671 (2017).
Zhou, Z., Yue, L., Wang, S., Lehn, J.-M. & Willner, I. DNA-based multiconstituent dynamic networks: hierarchical adaptive control over the composition and cooperative catalytic functions of the systems. J. Am. Chem. Soc. 140, 12077–12089 (2018).
Yue, L. et al. Consecutive feedback-driven constitutional dynamic networks. Proc. Natl Acad. Sci. USA 116, 2843–2848 (2019).
Del Grosso, E., Franco, E., Prins, L. J. & Ricci, F. Dissipative DNA nanotechnology. Nat. Chem. 14, 600–613 (2022).
Deng, J., Liu, W., Sun, M. & Walther, A. Dissipative organization of DNA oligomers for transient catalytic function. Angew. Chem. Int. Ed. 61, e202113477 (2022).
Wang, J. et al. DNAzyme- and light-induced dissipative and gated DNA networks. Chem. Sci. 12, 11204–11212 (2021).
Zhou, Z., Ouyang, Y., Wang, J. & Willner, I. Dissipative gated and cascaded DNA networks. J. Am. Chem. Soc. 143, 5071–5079 (2021).
Dong, J., Ouyang, Y., Wang, J., O’Hagan, M. P. & Willner, I. Assembly of dynamic gated and cascaded transient DNAzyme networks. ACS Nano 16, 6153–6164 (2022).
Del Grosso, E., Amodio, A., Ragazzon, G., Prins, L. J. & Ricci, F. Dissipative synthetic DNA-based receptors for the transient loading and release of molecular cargo. Angew. Chem. Int. Ed. 57, 10489–10493 (2018).
Yue, L., Wang, S., Wulf, V. & Willner, I. Stiffness-switchable DNA-based constitutional dynamic network hydrogels for self-healing and matrix-guided controlled chemical processes. Nat. Commun. 10, 4774 (2019).
Wang, C., Yue, L. & Willner, I. Controlling biocatalytic cascades with enzyme–DNA dynamic networks. Nat. Catal. 3, 941–950 (2020).
Zhou, C., Duan, X. & Liu, N. A plasmonic nanorod that walks on DNA origami. Nat. Commun. 6, 8102 (2015).
Cecconello, A., Besteiro, L. V., Govorov, A. O. & Willner, I. Chiroplasmonic DNA-based nanostructures. Nat. Rev. Mater. 2, 17039 (2017).
Wang, C. et al. Gated dissipative dynamic artificial photosynthetic model systems. J. Am. Chem. Soc. 143, 12120–12128 (2021).
Ouyang, Y., Zhang, P., Manis-Levy, H., Paltiel, Y. & Willner, I. Transient dissipative optical properties of aggregated Au nanoparticles, CdSe/Zns quantum dots, and supramolecular nucleic acid-stabilized Ag nanoclusters. J. Am. Chem. Soc. 143, 17622–17632 (2021).
Ashkenasy, G., Hermans, T. M., Otto, S. & Taylor, A. F. Systems chemistry. Chem. Soc. Rev. 46, 2543–2554 (2017).
Ludlow, R. F. & Otto, S. Systems chemistry. Chem. Soc. Rev. 37, 101–108 (2008).
Mattia, E. & Otto, S. Supramolecular systems chemistry. Nat. Nanotechnol. 10, 111–119 (2015).
Zhang, P. et al. Topologically switchable and gated transcription machinery. Chem. Sci. 13, 10555–10565 (2022).
Jiao, K. et al. Programming switchable transcription of topologically constrained DNA. J. Am. Chem. Soc. 142, 10739–10746 (2020).
Ohuchi, S., Mori, Y. & Nakamura, Y. Evolution of an inhibitory RNA aptamer against T7 RNA polymerase. FEBS Open Bio 2, 203–207 (2012).
Lloyd, J. et al. Dynamic control of aptamer–ligand activity using strand displacement reactions. ACS Synth. Biol. 7, 30–37 (2018).
Hertz, L. M. et al. The effect of pseudoknot base pairing on cotranscriptional structural switching of the fluoride riboswitch. Nucleic Acids Res. 52, 4466–4482 (2024).
Martos, A., Jiménez, M., Rivas, G. & Schwille, P. Towards a bottom-up reconstitution of bacterial cell division. Trends Cell Biol. 22, 634–643 (2012).
Rothfield, L., Taghbalout, A. & Shih, Y.-L. Spatial control of bacterial division-site placement. Nat. Rev. Microbiol. 3, 959–968 (2005).
Rizzoli, S. O. Synaptic vesicle recycling: steps and principles. EMBO J. 33, 788–822 (2014).
Bugyi, B. & Carlier, M.-F. Control of actin filament treadmilling in cell motility. Annu. Rev. Biophys. 39, 449–470 (2010).
Parsons, J. T., Horwitz, A. R. & Schwartz, M. A. Cell adhesion: integrating cytoskeletal dynamics and cellular tension. Nat. Rev. Mol. Cell Biol. 11, 633–643 (2010).
Ragazzon, G. & Prins, L. J. Energy consumption in chemical fuel-driven self-assembly. Nat. Nanotechnol. 13, 882–889 (2018).
Weißenfels, M., Gemen, J. & Klajn, R. Dissipative self-assembly: fueling with chemicals versus light. Chem 7, 23–37 (2021).
Pappas, C. G., Sasselli, I. R. & Ulijn, R. V. Biocatalytic pathway selection in transient tripeptide nanostructures. Angew. Chem. Int. Ed. 54, 8119–8123 (2015).
Te Brinke, E. et al. Dissipative adaptation in driven self-assembly leading to self-dividing fibrils. Nat. Nanotechnol. 13, 849–855 (2018).
Kariyawasam, L. S. & Hartley, C. S. Dissipative assembly of aqueous carboxylic acid anhydrides fueled by carbodiimides. J. Am. Chem. Soc. 139, 11949–11955 (2017).
Li, Z., Wang, J. & Willner, I. Transient out-of-equilibrium nucleic acid-based dissipative networks and their applications. Adv. Funct. Mater. 32, 2200799 (2022).
Liu, Q. et al. DNA-based dissipative assembly toward nanoarchitectonics. Adv. Funct. Mater. 32, 2201196 (2022).
Dong, J. & Willner, I. Dynamic transcription machineries guide the synthesis of temporally operating DNAzymes, gated and cascaded DNAzyme catalysis. ACS Nano 17, 687–696 (2023).
Li, Z., Wang, J. & Willner, I. Alternate strategies to induce dynamically modulated transient transcription machineries. ACS Nano 17, 18266–18279 (2023).
Simmel, F. C., Yurke, B. & Singh, H. R. Principles and applications of nucleic acid strand displacement reactions. Chem. Rev. 119, 6326–6369 (2019).
Li, Z., Wang, J. & Willner, I. Autoinhibited transient, gated and cascaded dynamic transcription of RNAs. Sci. Adv. 8, eabq5947 (2022).
Sun, M., Deng, J. & Walther, A. Communication and cross-regulation between chemically fueled sender and receiver reaction networks. Angew. Chem. Int. Ed. 62, e202214499 (2022).
Dong, J. & Willner, I. Transient transcription machineries modulate dynamic functions of G-quadruplexes: temporal regulation of biocatalytic circuits, gene replication and transcription. Angew. Chem. Int. Ed. 62, e202307898 (2023).
Mitchell, J. C., Harris, J. R., Malo, J., Bath, J. & Turberfield, A. J. Self-assembly of chiral DNA nanotubes. J. Am. Chem. Soc. 126, 16342–16343 (2004).
Rothemund, P. W. K. et al. Design and characterization of programmable DNA nanotubes. J. Am. Chem. Soc. 126, 16344–16352 (2004).
Agarwal, S. & Franco, E. Enzyme-driven assembly and disassembly of hybrid DNA–RNA nanotubes. J. Am. Chem. Soc. 141, 7831–7841 (2019).
Agarwal, S., Klocke, M. A., Pungchai, P. E. & Franco, E. Dynamic self-assembly of compartmentalized DNA nanotubes. Nat. Commun. 12, 3557 (2021).
Schultz, D., Wolynes, P. G., Jacob, E. B. & Onuchic, J. N. Deciding fate in adverse times: sporulation and competence in Bacillus subtilis. Proc. Natl Acad. Sci. USA 106, 21027–21034 (2009).
Oppenheim, A. B., Kobiler, O., Stavans, J., Court, D. L. & Adhya, S. Switches in bacteriophage lambda development. Annu. Rev. Genet. 39, 409–429 (2005).
Davidson, E. H. et al. A genomic regulatory network for development. Science 295, 1669–1678 (2002).
Peter, I. S. & Davidson, E. H. Assessing regulatory information in developmental gene regulatory networks. Proc. Natl Acad. Sci. USA 114, 5862–5869 (2017).
Kim, J., Khetarpal, I., Sen, S. & Murray, R. M. Synthetic circuit for exact adaptation and fold-change detection. Nucleic Acids Res. 42, 6078–6089 (2014).
Subsoontorn, P., Kim, J. & Winfree, E. Ensemble Bayesian analysis of bistability in a synthetic transcriptional switch. ACS Synth. Biol. 1, 299–316 (2012).
Schaffter, S. W. & Schulman, R. Building in vitro transcriptional regulatory networks by successively integrating multiple functional circuit modules. Nat. Chem. 11, 829–838 (2019).
Climent-Catala, A., Ouldridge, T. E., Stan, G.-B. V. & Bae, W. Building an RNA-based toggle switch using inhibitory RNA aptamers. ACS Synth. Biol. 11, 562–569 (2022).
Kim, J., White, K. S. & Winfree, E. Construction of an in vitro bistable circuit from synthetic transcriptional switches. Mol. Syst. Biol. 2, 68 (2006).
Kim, J. & Winfree, E. Synthetic in vitro transcriptional oscillators. Mol. Syst. Biol. 7, 465 (2011).
Weitz, M. et al. Diversity in the dynamical behaviour of a compartmentalized programmable biochemical oscillator. Nat. Chem. 6, 295–302 (2014).
Franco, E. et al. Timing molecular motion and production with a synthetic transcriptional clock. Proc. Natl Acad. Sci. USA 108, E784–E793 (2011).
Panda, S., Hogenesch, J. B. & Kay, S. A. Circadian rhythms from flies to human. Nature 417, 329–335 (2002).
Green, L. N. et al. Autonomous dynamic control of DNA nanostructure self-assembly. Nat. Chem. 11, 510–520 (2019).
Sneyd, J. et al. On the dynamical structure of calcium oscillations. Proc. Natl Acad. Sci. USA 114, 1456–1461 (2017).
Dick, T. E. et al. Cardiorespiratory coupling: common rhythms in cardiac, sympathetic, and respiratory activities. Prog. Brain Res. 209, 191–205 (2014).
Fujii, T. & Rondelez, Y. Predator–prey molecular ecosystems. ACS Nano 7, 27–34 (2013).
Dong, J. & Willner, I. Photochemically triggered, transient, and oscillatory transcription machineries guide temporal modulation of fibrinogenesis. J. Am. Chem. Soc. 147, 2216–2227 (2025).
Patino Diaz, A. et al. Programmable cell-free transcriptional switches for antibody detection. J. Am. Chem. Soc. 144, 5820–5826 (2022).
Dong, J. et al. Target-triggered transcription machinery for ultra-selective and sensitive fluorescence detection of nucleoside triphosphates in one minute. Biosens. Bioelectron. 100, 333–340 (2018).
Wang, L.-j., Liang, L., Liu, B.-j., Jiang, B. & Zhang, C.-y. A controlled T7 transcription-driven symmetric amplification cascade machinery for single-molecule detection of multiple repair glycosylases. Chem. Sci. 12, 5544–5554 (2021).
Cao, J. et al. Harnessing a previously unidentified capability of bacterial allosteric transcription factors for sensing diverse small molecules in vitro. Sci. Adv. 4, eaau4602 (2018).
Chou, L. Y. T. & Shih, W. M. In vitro transcriptional regulation via nucleic-acid-based transcription factors. ACS Synth. Biol. 8, 2558–2565 (2019).
Jung, J. K., Archuleta, C. M., Alam, K. K. & Lucks, J. B. Programming cell-free biosensors with DNA strand displacement circuits. Nat. Chem. Biol. 18, 385–393 (2022).
Lee, H. et al. Plug-and-play protein biosensors using aptamer-regulated in vitro transcription. Nat. Commun. 15, 7973 (2024).
Huang, F. et al. Three compartment liposome fusion: functional protocells for biocatalytic cascades and operation of dynamic DNA machineries. Adv. Funct. Mater. 33, 2302814 (2023).
Kröck, L. & Heckel, A. Photoinduced transcription by using temporarily mismatched caged oligonucleotides. Angew. Chem. Int. Ed. 44, 471–473 (2005).
Fischer, A. et al. DNA-tetrahedra corona-modified hydrogel microcapsules: “smart” ATP- or microRNA-responsive drug carriers. Small 18, 2204108 (2022).
Knott, G. J. & Doudna, J. A. CRISPR–Cas guides the future of genetic engineering. Science 361, 866–869 (2018).
Dzieciol, A. J. & Mann, S. Designs for life: protocell models in the laboratory. Chem. Soc. Rev. 41, 79–85 (2012).
Kurihara, K. et al. A recursive vesicle-based model protocell with a primitive model cell cycle. Nat. Commun. 6, 8352 (2015).
Elani, Y., Law, R. V. & Ces, O. Vesicle-based artificial cells as chemical microreactors with spatially segregated reaction pathways. Nat. Commun. 5, 5305 (2014).
Vriezema, D. M. et al. Positional assembly of enzymes in polymersome nanoreactors for cascade reactions. Angew. Chem. Int. Ed. 46, 7378–7382 (2007).
Huang, X., Patil, A. J., Li, M. & Mann, S. Design and construction of higher-order structure and function in proteinosome-based protocells. J. Am. Chem. Soc. 136, 9225–9234 (2014).
Paleos, C. M., Tsiourvas, D., Sideratou, Z. & Pantos, A. Formation of artificial multicompartment vesosome and dendrosome as prospected drug and gene delivery carriers. J. Control. Release 170, 141–152 (2013).
Qiao, Y., Li, M., Booth, R. & Mann, S. Predatory behaviour in synthetic protocell communities. Nat. Chem. 9, 110–119 (2017).
Chan, V., Novakowski, S. K., Law, S., Klein-Bosgoed, C. & Kastrup, C. J. Controlled transcription of exogenous mRNA in platelets using protocells. Angew. Chem. Int. Ed. 54, 13590–13593 (2015).
Sun, L. et al. Guiding protein delivery into live cells using DNA-programmed membrane fusion. Chem. Sci. 9, 5967–5975 (2018).
Acknowledgements
This research is supported by the Israel Science Foundation (grant 2049/20).
Author information
Authors and Affiliations
Contributions
J.D. and I.W. contributed to discussions and co-wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Chemistry thanks Alessandro Porchetta, Anthony Genot and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Dong, J., Willner, I. Synthetic dynamic transcription frameworks and their applications. Nat. Chem. 18, 227–245 (2026). https://doi.org/10.1038/s41557-025-02046-w
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41557-025-02046-w