Fig. 1: Scaling and self-similarity of de novo DNA methylation.
From: Scaling and self-similarity in the formation of the embryonic epigenome
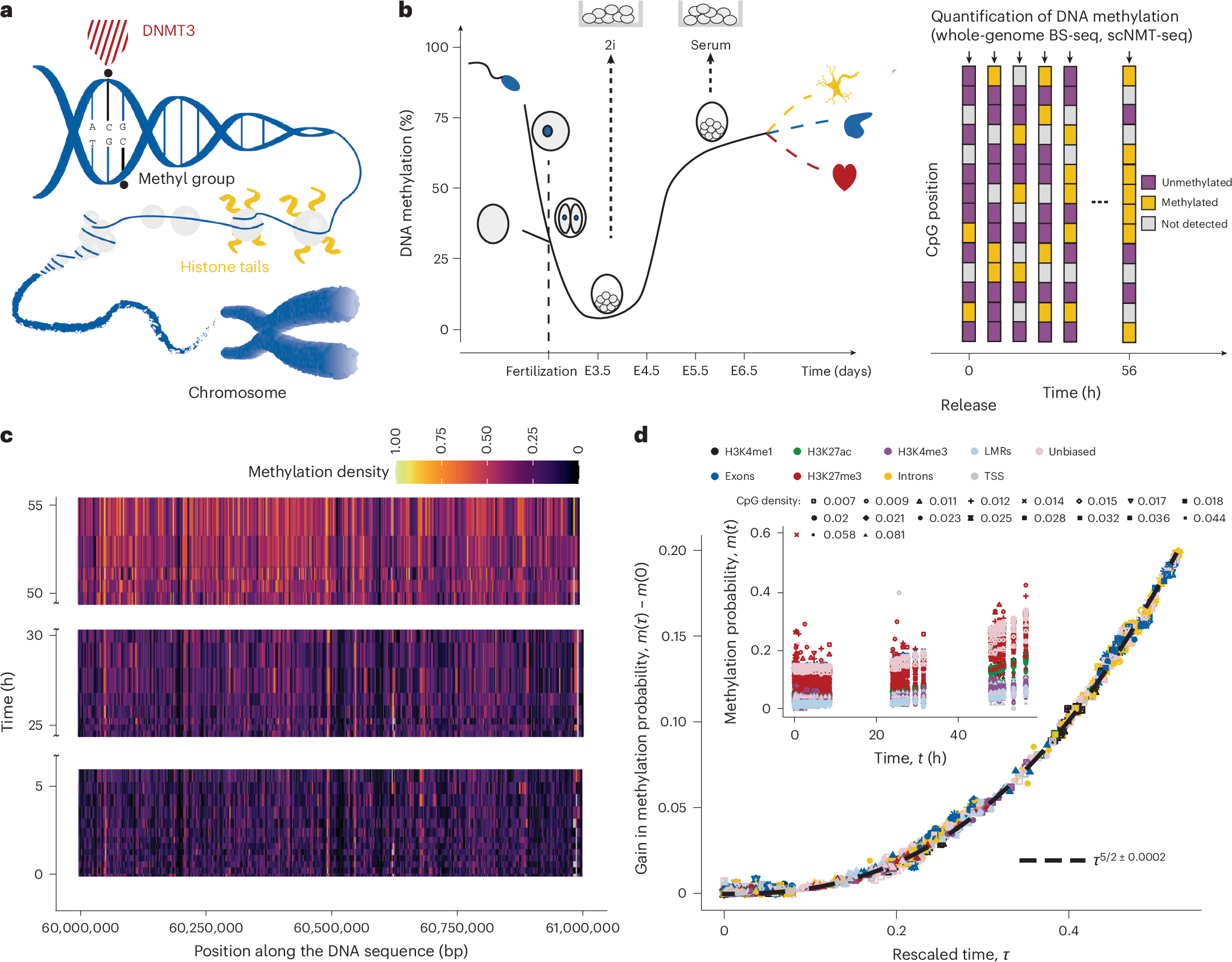
a, Schematic showing epigenetic processes involved in regulating cell fate. b, Schematic showing the time evolution of global DNA methylation levels during early mouse development (left) and the quantification of DNA methylation in the 2i release experiment. In single-cell BS-seq, a given CpG site can be measured as methylated, unmethylated or undetected. c, Heat map showing DNA methylation levels across an exemplary genomic domain over time. d, Inset: average probability that a CpG site is methylated increases at different rates in genomic regions with different functional annotation (colour; Extended Data Table 1) and CpG density (shape). H3K4me1, histone H3 Lys4 monomethylation; H3K4me3, histone H3 Lys4 trimethylation; H3K27me3, histone H3 Lys27 monomethylation; H3K27ac, histone H3 Lys27 acetylation; LMR, lowly methylated regions; TSS, transcription start sites; Unbiased, tiling of the genome into windows of equal coverage. The main plot shows the curves that were parameterized as m(t) = m(0) + btc; τ = t/b is shown on the x axis and m(τ) − m(0) on the y axis (see ‘Quantification of DNA methylation dynamics’ section). The gain in average DNA methylation follows a single power law with an exponent of 5/2 (dashed line; Methods). The exponent is robust with respect to log transformation and its uncertainty denotes 95% confidence intervals.
