Abstract
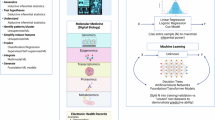
Mathematical modelling has proven to be a valuable tool in predicting the delivery and efficacy of molecular, antibody-based, nano and cellular therapy in solid tumours. Mathematical models based on our understanding of the biological processes at subcellular, cellular and tissue level are known as mechanistic models that, in turn, are divided into continuous and discrete models. Continuous models are further divided into lumped parameter models — for describing the temporal distribution of medicine in tumours and normal organs — and distributed parameter models — for studying the spatiotemporal distribution of therapy in tumours. Discrete models capture interactions at the cellular and subcellular levels. Collectively, these models are useful for optimizing the delivery and efficacy of molecular, nanoscale and cellular therapy in tumours by incorporating the biological characteristics of tumours, the physicochemical properties of drugs, the interactions among drugs, cancer cells and various components of the tumour microenvironment, and for enabling patient-specific predictions when combined with medical imaging. Artificial intelligence-based methods, such as machine learning, have ushered in a new era in oncology. These data-driven approaches complement mechanistic models and have immense potential for improving cancer detection, treatment and drug discovery. Here we review these diverse approaches and suggest ways to combine mechanistic and artificial intelligence-based models to further improve patient treatment outcomes.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Jain, R. K. Normalizing tumor microenvironment to treat cancer: bench to bedside to biomarkers. J. Clin. Oncol. 31, 2205–2218 (2013). This review article describes how different components of the TME are abnormal, how these abnormalities contribute to resistance to drug delivery and efficacy, and how repairing these abnormalities can improve drug delivery and efficacy.
Jain, R. K. Determinants of tumor blood flow: a review. Cancer Res. 48, 2641–2658 (1988).
Nagy, J., Chang, S.-H., Shih, S.-C., Dvorak, A. & Dvorak, H. Heterogeneity of the tumor vasculature. Semin. Thromb. Hemost. 36, 321–331 (2010).
Jain, R. K. Antiangiogenesis strategies revisited: from starving tumors to alleviating hypoxia. Cancer Cell 26, 605–622 (2014).
Mitchell, M. J., Jain, R. K. & Langer, R. Engineering and physical sciences in oncology: challenges and opportunities. Nat. Rev. Cancer 17, 659–675 (2017).
Nia, H. T., Munn, L. L. & Jain, R. K. Probing the physical hallmarks of cancer. Nat. Methods https://doi.org/10.1038/s41592-024-02564-4 (2025). This review article describes different types of physical forces in tumours, how to measure them, how to overcome the challenges posed by these physical constraints in tumours and how mathematical modelling has helped measure these forces.
Stylianopoulos, T. et al. Causes, consequences, and remedies for growth-induced solid stress in murine and human tumors. Proc. Natl Acad. Sci. USA 109, 15101–15108 (2012).
Voutouri, C. & Stylianopoulos, T. Accumulation of mechanical forces in tumors is related to hyaluronan content and tissue stiffness. PLoS ONE 13, e0193801 (2018).
Linke, J. A., Munn, L. L. & Jain, R. K. Compressive stresses in cancer: characterization and implications for tumour progression and treatment. Nat. Rev. Cancer 24, 768–791 (2024).
Padera, T. P. et al. Cancer cells compress intratumour vessels. Nature 427, 695–695 (2004).
Stylianopoulos, T., Munn, L. L. & Jain, R. K. Reengineering the physical microenvironment of tumors to improve drug delivery and efficacy: from mathematical modeling to bench to bedside. Trends Cancer 4, 292–319 (2018). This review describes the barriers posed by the physical microenvironment of tumours to effective drug delivery and provides evidence that reengineering the tumour microenvironment can enhance treatment efficacy.
Jain, R. K. & Baxter, L. T. Mechanisms of heterogeneous distribution of monoclonal antibodies and other macromolecules in tumors: significance of elevated interstitial pressure. Cancer Res. 48, 7022–7032 (1988). This paper presented the first mathematical model for the transport of fluid and macromolecules in tumours and predicted the spatial profile of interstitial fluid pressure in tumours subsequently confirmed by direct measurements in tumours in rats.
Jain, R. K., Tong, R. T. & Munn, L. L. Effect of vascular normalization by antiangiogenic therapy on interstitial hypertension, peritumor edema, and lymphatic metastasis: insights from a mathematical model. Cancer Res. 67, 2729–2735 (2007).
McDonald, T. O. et al. Computational approaches to modelling and optimizing cancer treatment. Nat. Rev. Bioeng. 1, 695–711 (2023).
Nikmaneshi, M. R., Jain, R. K. & Munn, L. L. Computational simulations of tumor growth and treatment response: benefits of high-frequency, low-dose drug regimens and concurrent vascular normalization. PLoS Comput. Biol. 19, e1011131 (2023). This paper developed a hybrid continuous–discrete model that simulates the TME, showing that metronomic therapy improves drug delivery, especially when combined with anti-angiogenic treatments.
Vavourakis, V. et al. A validated multiscale in-silico model for mechano-sensitive tumour angiogenesis and growth. PLoS Comput. Biol. 13, e1005259 (2017).
Yanagisawa, H., Sugimoto, M. & Miyashita, T. Mathematical simulation of tumour angiogenesis: angiopoietin balance is a key factor in vessel growth and regression. Sci. Rep. 11, 419 (2021).
Slavkova, K. P. et al. Mathematical modelling of the dynamics of image-informed tumor habitats in a murine model of glioma. Sci. Rep. 13, 2916 (2023).
Mirramezani, M. & Shadden, S. C. A distributed lumped parameter model of blood flow. Ann. Biomed. Eng. 48, 2870–2886 (2020).
Zhu, J. et al. Translational pharmacokinetic/pharmacodynamic modeling and simulation of oxaliplatin and irinotecan in colorectal cancer. Pharmaceutics 15, 2274 (2023).
Michor, F. & Beal, K. Improving cancer treatment via mathematical modeling: surmounting the challenges is worth the effort. Cell 163, 1059–1063 (2015).
Malinzi, J., Basita, K. B., Padidar, S. & Adeola, H. A. Prospect for application of mathematical models in combination cancer treatments. Inform. Med. Unlocked 23, 100534 (2021).
Baxter, L. T., Zhu, H., Mackensen, D. G. & Jain, R. K. Physiologically based pharmacokinetic model for specific and nonspecific monoclonal antibodies and fragments in normal tissues and human tumor xenografts in nude mice. Cancer Res. 54, 1517–1528 (1994).
De Montigny, J. et al. An in silico hybrid continuum-/agent-based procedure to modelling cancer development: interrogating the interplay amongst glioma invasion, vascularity and necrosis. Methods 185, 94–104 (2021).
Vavourakis, V., Stylianopoulos, T. & Wijeratne, P. A. In-silico dynamic analysis of cytotoxic drug administration to solid tumours: effect of binding affinity and vessel permeability. PLoS Comput. Biol. 14, e1006460 (2018).
Moradi Kashkooli, F. & Soltani, M. Evaluation of solid tumor response to sequential treatment cycles via a new computational hybrid approach. Sci. Rep. 11, 21475 (2021).
Ocaña-Tienda, B. & Pérez-García, V. M. Mathematical modeling of brain metastases growth and response to therapies: a review. Math. Biosci. 373, 109207 (2024).
Cogno, N., Axenie, C., Bauer, R. & Vavourakis, V. Agent-based modeling in cancer biomedicine: applications and tools for calibration and validation. Cancer Biol. Ther. 25, 2344600 (2024).
Dewhirst, M. W. & Secomb, T. W. Transport of drugs from blood vessels to tumour tissue. Nat. Rev. Cancer 17, 738–750 (2017).
Butner, J. D. et al. Mathematical modeling of cancer immunotherapy for personalized clinical translation. Nat. Comput. Sci. 2, 785–796 (2022).
Luchini, C., Pea, A. & Scarpa, A. Artificial intelligence in oncology: current applications and future perspectives. Br. J. Cancer 126, 4–9 (2022).
Rösler, W. et al. An overview and a roadmap for artificial intelligence in hematology and oncology. J. Cancer Res. Clin. Oncol. 149, 7997–8006 (2023).
Lorenzo, G. et al. Patient-specific, mechanistic models of tumor growth incorporating artificial intelligence and big data. Annu. Rev. Biomed. Eng. 26, 529–560 (2024).
Lotter, W. et al. Artificial intelligence in oncology: current landscape, challenges, and future directions. Cancer Discov. 14, 711–726 (2024).
Gerlowski, L. E. & Jain, R. K. Physiologically based pharmacokinetic modeling: principles and applications. J. Pharm. Sci. 72, 1103–1127 (1983). This comprehensive review article summarized how pharmacokinetic models can describe the distribution and disposition of drugs in living systems, helping the design of preclinical and clinical studies.
Weissbrod, J. M., Jain, R. K. & Sirotnak, F. M. Pharmacokinetics of methotrexate in leukemia cells: effect of dose and mode of injection. J. Pharmacokinet. Biopharm. 6, 487–503 (1978).
Jayachandran, P., Knox, S. J., Garcia-Cremades, M. & Savić, R. M. Clinical pharmacokinetics of oral sodium selenite and dosing implications in the treatment of patients with metastatic cancer. Drugs R D 21, 169–178 (2021).
Jain, R. K., Wei, J. & Gullino, P. M. Pharmacokinetics of methotrexate in solid tumors. J. Pharmacokinet. Biopharm. 7, 181–194 (1979).
McKenna, M. T. et al. A predictive mathematical modeling approach for the study of doxorubicin treatment in triple negative breast cancer. Sci. Rep. 7, 5725 (2017).
Ciccolini, J. et al. Pharmacokinetics and pharmacodynamics-based mathematical modeling identifies an optimal protocol for metronomic chemotherapy. Cancer Res. 77, 4723–4733 (2017).
Clewell, R. A. & Clewell, H. J. Development and specification of physiologically based pharmacokinetic models for use in risk assessment. Regul. Toxicol. Pharmacol. 50, 129–143 (2008).
Zang, X. & Kagan, L. Physiologically-based modeling and interspecies prediction of paclitaxel pharmacokinetics. J. Pharmacokinet. Pharmacodyn. 45, 577–592 (2018).
Dumontet, C., Reichert, J. M., Senter, P. D., Lambert, J. M. & Beck, A. Antibody–drug conjugates come of age in oncology. Nat. Rev. Drug Discov. 22, 641–661 (2023).
Klein, C., Brinkmann, U., Reichert, J. M. & Kontermann, R. E. The present and future of bispecific antibodies for cancer therapy. Nat. Rev. Drug Discov. 23, 301–319 (2024).
Drago, J. Z., Modi, S. & Chandarlapaty, S. Unlocking the potential of antibody–drug conjugates for cancer therapy. Nat. Rev. Clin. Oncol. 18, 327–344 (2021).
Ray, C. M. P., Yang, H., Spangler, J. B. & Mac Gabhann, F. Mechanistic computational modeling of monospecific and bispecific antibodies targeting interleukin-6/8 receptors. PLoS Comput. Biol. 20, e1012157 (2024).
Yang, H. et al. Engineered bispecific antibodies targeting the interleukin-6 and -8 receptors potently inhibit cancer cell migration and tumor metastasis. Mol. Ther. 30, 3430–3449 (2022).
Shah, D. K., Haddish-Berhane, N. & Betts, A. Bench to bedside translation of antibody drug conjugates using a multiscale mechanistic PK/PD model: a case study with brentuximab–vedotin. J. Pharmacokinet. Pharmacodyn. 39, 643–659 (2012).
Chudasama, V. L. et al. Semi-mechanistic population pharmacokinetic model of multivalent trastuzumab emtansine in patients with metastatic breast cancer. Clin. Pharmacol. Ther. 92, 520–527 (2012).
Lu, D. et al. Semi-mechanistic multiple-analyte pharmacokinetic model for an antibody–drug-conjugate in cynomolgus monkeys. Pharm. Res. 32, 1907–1919 (2015).
Betts, A. M. et al. Preclinical to clinical translation of antibody–drug conjugates using PK/PD modeling: a retrospective analysis of inotuzumab ozogamicin. AAPS J. 18, 1101–1116 (2016).
Singh, A. & Shah, D. in Innovations for Next-Generation Antibody–Drug Conjugates (ed. Demelin, M.) 73–97 (Springer, 2018).
Scheuher, B. et al. Towards a platform quantitative systems pharmacology (QSP) model for preclinical to clinical translation of antibody drug conjugates (ADCs). J. Pharmacokinet. Pharmacodyn. 51, 429–447 (2023).
Singh, A. P. & Shah, D. K. Application of a PK–PD modeling and simulation-based strategy for clinical translation of antibody–drug conjugates: a case study with trastuzumab emtansine (T-DM1). AAPS J. 19, 1054–1070 (2017).
Nicolò, E. et al. Combining antibody–drug conjugates with immunotherapy in solid tumors: current landscape and future perspectives. Cancer Treat. Rev. 106, 102395 (2022).
Jain, R. K. & Stylianopoulos, T. Delivering nanomedicine to solid tumors. Nat. Rev. Clin. Oncol. 7, 653–664 (2010).
Nirmala, M. J. et al. Cancer nanomedicine: a review of nano-therapeutics and challenges ahead. RSC Adv. 13, 8606–8629 (2023).
Li, M., Al-Jamal, K. T., Kostarelos, K. & Reineke, J. Physiologically based pharmacokinetic modeling of nanoparticles. ACS Nano 4, 6303–6317 (2010).
Kutumova, E. O. et al. Physiologically based pharmacokinetic modeling of nanoparticle biodistribution: a review of existing models, simulation software, and data analysis tools. Int. J. Mol. Sci. 23, 12560 (2022).
Dogra, P. et al. A mathematical model to predict nanomedicine pharmacokinetics and tumor delivery. Comput. Struct. Biotechnol. J. 18, 518–531 (2020).
Patnaik, A. et al. Phase I study of pembrolizumab (MK-3475; anti–PD-1 monoclonal antibody) in patients with advanced solid tumors. Clin. Cancer Res. 21, 4286–4293 (2015).
Elassaiss-Schaap, J. et al. Using model-based ‘Learn and Confirm’ to reveal the pharmacokinetics–pharmacodynamics relationship of pembrolizumab in the KEYNOTE-001 trial: modeling of the PK/PD of Pembro in KEYNOTE-001. CPT Pharmacomet. Syst. Pharmacol. 6, 21–28 (2017).
Ahamadi, M. et al. Model-based characterization of the pharmacokinetics of pembrolizumab: a humanized anti-PD-1 monoclonal antibody in advanced solid tumors: pharmacokinetics of pembro in solid tumors. CPT Pharmacomet. Syst. Pharmacol. 6, 49–57 (2017).
Lala, M. et al. A six-weekly dosing schedule for pembrolizumab in patients with cancer based on evaluation using modelling and simulation. Eur. J. Cancer 131, 68–75 (2020).
Tao, Z. et al. Impact of T cell characteristics on CAR-T cell therapy in hematological malignancies. Blood Cancer J. 14, 213 (2024).
US Food and Drug Administration. FDA approves first cellular therapy to treat patients with unresectable or metastatic melanoma (FDA, 2024).
Chen, T., Wang, M., Chen, Y. & Liu, Y. Current challenges and therapeutic advances of CAR-T cell therapy for solid tumors. Cancer Cell Int. 24, 133 (2024).
Zhu, H., Melder, R. J., Baxter, L. T. & Jain, R. K. Physiologically based kinetic model of effector cell biodistribution in mammals: implications for adoptive immunotherapy. Cancer Res. 56, 3771–3781 (1996). This paper presented the first mathematical model of adoptively transferred immune cells in tumours and normal tissues and suggested ways to improve the improve their delivery of tumours — confirmed later in CAR T cells in clinical trials.
Choi, B. D. et al. Intraventricular CARv3-TEAM-E T cells in recurrent glioblastoma. N. Engl. J. Med. 390, 1290–1298 (2024).
Nikitich, A., Helmlinger, G., Peskov, K. & Bocharov, G. Mathematical modeling of endogenous and exogenously administered T cell recirculation in mouse and its application to pharmacokinetic studies of cell therapies. Front. Immunol. 15, 1357706 (2024).
Adhikarla, V. et al. Designing combination therapies for cancer treatment: application of a mathematical framework combining CAR T-cell immunotherapy and targeted radionuclide therapy. Front. Immunol. 15, 1358478 (2024). This mathematical model showed how combination treatment with CAR T cell therapy and targeted radionuclide therapy can be optimized.
Jafarnejad, M. et al. A computational model of neoadjuvant PD-1 inhibition in non-small cell lung cancer. AAPS J. 21, 79 (2019).
Arulraj, T. et al. Virtual patient analysis identifies strategies to improve the performance of predictive biomarkers for PD-1 blockade. Proc. Natl Acad. Sci. USA 121, e2410911121 (2024).
Ma, H. et al. Combination therapy with T cell engager and PD-L1 blockade enhances the antitumor potency of T cells as predicted by a QSP model. J. Immunother. Cancer 8, e001141 (2020).
Milberg, O. et al. A QSP model for predicting clinical responses to monotherapy, combination and sequential therapy following CTLA-4, PD-1, and PD-L1 checkpoint blockade. Sci. Rep. 9, 11286 (2019).
Wang, H. et al. Conducting a virtual clinical trial in HER2-negative breast cancer using a quantitative systems pharmacology model with an epigenetic modulator and immune checkpoint inhibitors. Front. Bioeng. Biotechnol. 8, 141 (2020).
Hardiansyah, D. & Ng, C. M. Quantitative systems pharmacology model of chimeric antigen receptor T‐cell therapy. Clin. Transl. Sci. 12, 343–349 (2019).
Huitema, A. D. R. et al. Relationship between exposure and toxicity in high-dose chemotherapy with cyclophosphamide, thiotepa and carboplatin. Ann. Oncol. 13, 374–384 (2002).
Harkos, C., Svensson, S. F., Emblem, K. E. & Stylianopoulos, T. Inducing biomechanical heterogeneity in brain tumor modeling by MR elastography: effects on tumor growth, vascular density and delivery of therapeutics. Cancers 14, 884 (2022). This paper demonstrated that integrating patient-specific biomechanical properties with magnetic resonance elastography can improve mathematical models of glioblastoma, enabling more realistic simulations of tumour growth, vascular changes and drug delivery.
Jain, R. K. & Wei, J. Dynamics of drug transport in solid tumors: distributed parameter model. J. Bioeng. 1, 313–329 (1977). The first distributed parameter mathematical model to describe the non-uniform distribution of methotrexate — a widely prescribed cancer drug — in solid tumours grown in rats, and identified poor and heterogeneous blood supply as a challenge in drug delivery to tumours.
Baxter, L. T. & Jain, R. K. Transport of fluid and macromolecules in tumors. I. Role of interstitial pressure and convection. Microvasc. Res. 37, 77–104 (1989).
Baxter, L. T. & Jain, R. K. Transport of fluid and macromolecules in tumors: III. Role of binding and metabolism. Microvasc. Res. 41, 5–23 (1991).
Baxter, L. T. & Jain, R. K. Transport of fluid and macromolecules in tumors. II. Role of heterogeneous perfusion and lymphatics. Microvasc. Res. 40, 246–263 (1990).
Baxter, L. T. & Jain, R. K. Transport of fluid and macromolecules in tumors. IV. A microscopic model of the perivascular distribution. Microvasc. Res. 41, 252–272 (1991).
Boucher, Y., Baxter, L. T. & Jain, R. K. Interstitial pressure gradients in tissue-isolated and subcutaneous tumors: implications for therapy. Cancer Res. 50, 4478–4484 (1990).
Boucher, Y., Salehi, H., Witwer, B., Harsh, G. & Jain, R. Interstitial fluid pressure in intracranial tumours in patients and in rodents. Br. J. Cancer 75, 829–836 (1997).
Goel, S. et al. Normalization of the vasculature for treatment of cancer and other diseases. Physiol. Rev. 91, 1071–1121 (2011).
Nia, H. T., Munn, L. L. & Jain, R. K. Physical traits of cancer. Science 370, eaaz0868 (2020).
Helmlinger, G., Netti, P. A., Lichtenbeld, H. C., Melder, R. J. & Jain, R. K. Solid stress inhibits the growth of multicellular tumor spheroids. Nat. Biotechnol. 15, 778–783 (1997).
Netti, P. A., Baxter, L. T., Boucher, Y., Skalak, R. & Jain, R. K. Time-dependent behavior of interstitial fluid pressure in solid tumors: implications for drug delivery. Cancer Res. 55, 5451–5458 (1995).
Skalak, R., Zargaryan, S., Jain, R. K., Netti, P. A. & Hoger, A. Compatibility and the genesis of residual stress by volumetric growth. J. Math. Biol. 34, 889–914 (1996).
Swartz, M. A. et al. Mechanics of interstitial–lymphatic fluid transport: theoretical foundation and experimental validation. J. Biomech. 32, 1297–1307 (1999).
Roose, T., Netti, P. A., Munn, L. L., Boucher, Y. & Jain, R. K. Solid stress generated by spheroid growth estimated using a linear poroelasticity model. Microvasc. Res. 66, 204–212 (2003).
Stylianopoulos, T. et al. Coevolution of solid stress and interstitial fluid pressure in tumors during progression: implications for vascular collapse. Cancer Res. 73, 3833–3841 (2013). The first mathematical model to calculate the increase in both solid stress and interstitial fluid pressure and their implications in drug delivery.
Mpekris, F., Angeli, S., Pirentis, A. P. & Stylianopoulos, T. Stress-mediated progression of solid tumors: effect of mechanical stress on tissue oxygenation, cancer cell proliferation, and drug delivery. Biomech. Model. Mechanobiol. 14, 1391–1402 (2015).
Mpekris, F., Baish, J. W., Stylianopoulos, T. & Jain, R. K. Role of vascular normalization in benefit from metronomic chemotherapy. Proc. Natl Acad. Sci. USA 114, 1994–1999 (2017).
Seano, G. et al. Solid stress in brain tumours causes neuronal loss and neurological dysfunction and can be reversed by lithium. Nat. Biomed. Eng. 3, 230–245 (2019).
Nia, H. T. et al. Solid stress estimation via intraoperative 3D navigation in patients with brain tumors. Preprint at medRxiv https://doi.org/10.1101/2024.11.28.24318104 (2024).
Papageorgis, P. et al. Tranilast-induced stress alleviation in solid tumors improves the efficacy of chemo- and nanotherapeutics in a size-independent manner. Sci. Rep. 7, 46140 (2017).
Voutouri, C. et al. Ultrasound stiffness and perfusion markers correlate with tumor volume responses to immunotherapy. Acta Biomater. 167, 121–134 (2023).
Angeli, S., Emblem, K. E., Due-Tonnessen, P. & Stylianopoulos, T. Towards patient-specific modeling of brain tumor growth and formation of secondary nodes guided by DTI-MRI. Neuroimage Clin. 20, 664–673 (2018).
Swanson, K. R., Alvord, E. C. & Murray, J. D. Virtual brain tumours (gliomas) enhance the reality of medical imaging and highlight inadequacies of current therapy. Br. J. Cancer 86, 14–18 (2002).
Hadjigeorgiou, A. G. & Stylianopoulos, T. Evaluation of growth-induced, mechanical stress in solid tumors and spatial association with extracellular matrix content. Biomech. Model. Mechanobiol. 22, 1625–1643 (2023).
Swanson, K. R., Alvord, E. C. & Murray, J. D. A quantitative model for differential motility of gliomas in grey and white matter. Cell Prolif. 33, 317–329 (2000).
Swanson, K. R., Bridge, C., Murray, J. D. & Alvord, E. C. Virtual and real brain tumors: using mathematical modeling to quantify glioma growth and invasion. J. Neurol. Sci. 216, 1–10 (2003).
Jbabdi, S. et al. Simulation of anisotropic growth of low‐grade gliomas using diffusion tensor imaging. Magn. Reson. Med. 54, 616–624 (2005).
Katsamba, I. et al. Biomechanical modelling of spinal tumour anisotropic growth. Proc. R. Soc. A 476, 20190364 (2020).
Kolokotroni, E. et al. A multidisciplinary hyper-modeling scheme in personalized in silico oncology: coupling cell kinetics with metabolism, signaling networks, and biomechanics as plug-in component models of a cancer digital twin. J. Pers. Med. 14, 475 (2024). This paper developed a modeling scheme for personalized simulations for Wilms tumour and non-small-cell lung cancer that predicts treatment responses and promising future applications in clinical decision support and in silico trials.
Bodei, L., Herrmann, K., Schöder, H., Scott, A. M. & Lewis, J. S. Radiotheranostics in oncology: current challenges and emerging opportunities. Nat. Rev. Clin. Oncol. 19, 534–550 (2022).
Piranfar, A. et al. Radiopharmaceutical transport in solid tumors via a 3-dimensional image-based spatiotemporal model. npj Syst. Biol. Appl. 10, 39 (2024). This paper presented a model based on clinical imaging that simulates the pharmacokinetics of 177Lu-PSMA in prostate tumours, offering insights for personalizing treatment for metastatic prostate cancer.
Walker, C. R. & Angelo, M. Insights and opportunity costs in applying spatial biology to study the tumor microenvironment. Cancer Discov. 14, 707–710 (2024).
Lai, X. & Friedman, A. Combination therapy of cancer with cancer vaccine and immune checkpoint inhibitors: a mathematical model. PLoS ONE 12, e0178479 (2017).
Friedman, A. & Hao, W. The role of exosomes in pancreatic cancer microenvironment. Bull. Math. Biol. 80, 1111–1133 (2018).
Mpekris, F. et al. Combining microenvironment normalization strategies to improve cancer immunotherapy. Proc. Natl Acad. Sci. USA 117, 3728–3737 (2020).
Harkos, C. & Stylianopoulos, T. Investigating the synergistic effects of immunotherapy and normalization treatment in modulating tumor microenvironment and enhancing treatment efficacy. J. Theor. Biol. 583, 111768 (2024).
Harkos, C., Stylianopoulos, T. & Jain, R. K. Mathematical modeling of intratumoral immunotherapy yields strategies to improve the treatment outcomes. PLoS Comput. Biol. 19, e1011740 (2023). This paper presented a mechanistic mathematical model for optimizing intratumoral immunotherapy that revealed how the tumour microenvironment and conjugated cytokine properties influence treatment outcome.
Anderson, A. R. A. & Chaplain, M. A. J. Continuous and discrete mathematical models of tumor-induced angiogenesis. Bull. Math. Biol. 60, 857–899 (1998). The first discrete model of tumour angiogenesis.
Anderson, A. R. A., Chaplain, M. A. J. & McDougall, S. in Modeling Tumor Vasculature: Molecular, Cellular, and Tissue Level Aspects and Implications (ed. Jackson, T. L. L.) 105–133 (Springer, 2012).
Valentim, C. A., Rabi, J. A. & David, S. A. Cellular-automaton model for tumor growth dynamics: virtualization of different scenarios. Comput. Biol. Med. 153, 106481 (2023).
Chauhan, V. P. et al. Normalization of tumour blood vessels improves the delivery of nanomedicines in a size-dependent manner. Nat. Nanotechnol. 7, 383–388 (2012).
Stylianopoulos, T., Soteriou, K., Fukumura, D. & Jain, R. K. Cationic nanoparticles have superior transvascular flux into solid tumors: insights from a mathematical model. Ann. Biomed. Eng. 41, 68–77 (2013).
Stylianopoulos, T., Economides, E.-A., Baish, J. W., Fukumura, D. & Jain, R. K. Towards optimal design of cancer nanomedicines: multi-stage nanoparticles for the treatment of solid tumors. Ann. Biomed. Eng. 43, 2291–2300 (2015).
Rejniak, K. A. et al. The role of tumor tissue architecture in treatment penetration and efficacy: an integrative study. Front. Oncol. 3, 111 (2013).
Kingsley, J. L., Costello, J. R., Raghunand, N. & Rejniak, K. A. Bridging cell-scale simulations and radiologic images to explain short-time intratumoral oxygen fluctuations. PLoS Comput. Biol. 17, e1009206 (2021).
Shields, J. D. et al. Autologous chemotaxis as a mechanism of tumor cell homing to lymphatics via interstitial flow and autocrine CCR7 signaling. Cancer Cell 11, 526–538 (2007).
Ben-Ami, Y., Pitt-Francis, J. M., Maini, P. K. & Byrne, H. M. Using a probabilistic approach to derive a two-phase model of flow-induced cell migration. Biophys. J. 123, 799–813 (2024).
Kecman, V. in Real World Applications of Computational Intelligence Vol. 179 (eds Gh. Negoita, M. & Reusch, B.) 49–103 (Springer, 2005). This paper described an AI-powered analyser for whole-slide images of tumour-infiltrating lymphocytes that identified three immune phenotypes that correlate with tumour response and survival in advanced non-small-cell lung cancer.
Voutouri, C. et al. A convolutional attention model for predicting response to chemo-immunotherapy from ultrasound elastography in mouse tumor models. Commun. Med. 4, 203 (2024).
Englezos, D., Voutouri, C. & Stylianopoulos, T. Machine learning analysis reveals tumor stiffness and hypoperfusion as biomarkers predictive of cancer treatment efficacy. Transl. Oncol. 44, 101944 (2024).
Rumelhart, D. E., Hinton, G. E. & Williams, R. J. Learning representations by back-propagating errors. Nature 323, 533–536 (1986).
Esteva, A. et al. Dermatologist-level classification of skin cancer with deep neural networks. Nature 542, 115–118 (2017).
Malibari, A. A. et al. Optimal deep neural network-driven computer aided diagnosis model for skin cancer. Comput. Electr. Eng. 103, 108318 (2022).
Krakowski, I. et al. Human-AI interaction in skin cancer diagnosis: a systematic review and meta-analysis. npj Digital Med. 7, 78 (2024).
Setio, A. A. A. et al. Pulmonary nodule detection in CT images: false positive reduction using multi-view convolutional networks. IEEE Trans. Med. Imaging 35, 1160–1169 (2016).
Ma, L., Li, G., Feng, X., Fan, Q. & Liu, L. TiCNet: transformer in convolutional neural network for pulmonary nodule detection on CT images. J. Imaging Inform. Med. 37, 196–208 (2024).
Wakili, M. A. et al. Classification of breast cancer histopathological images using DenseNet and transfer learning. Comput. Intell. Neurosci. 2022, 1–31 (2022).
Falou, O., Sannachi, L., Haque, M., Czarnota, G. J. & Kolios, M. C. Transfer learning of pre-treatment quantitative ultrasound multi-parametric images for the prediction of breast cancer response to neoadjuvant chemotherapy. Sci. Rep. 14, 2340 (2024).
Stashko, C. et al. A convolutional neural network STIFMap reveals associations between stromal stiffness and EMT in breast cancer. Nat. Commun. 14, 3561 (2023).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Ekins, S. et al. Exploiting machine learning for end-to-end drug discovery and development. Nat. Mater. 18, 435–441 (2019).
Stokes, J. M. et al. A deep learning approach to antibiotic discovery. Cell 180, 688–702.e13 (2020).
Mongia, M., Guler, M. & Mohimani, H. An interpretable machine learning approach to identify mechanism of action of antibiotics. Sci. Rep. 12, 10342 (2022).
You, Y. et al. Artificial intelligence in cancer target identification and drug discovery. Signal Transduct. Target. Ther. 7, 156 (2022).
Campana, P. A. et al. Cancer drug sensitivity estimation using modular deep Graph Neural Networks. NAR Genom. Bioinform. 6, lqae043 (2024).
Stärk, H., Ganea, O.-E., Pattanaik, L., Barzilay, R. & Jaakkola, T. EquiBind: geometric deep learning for drug binding structure prediction. Preprint at arXiv https://doi.org/10.48550/ARXIV.2202.05146 (2022).
Boehm, K. M. et al. Multimodal data integration using machine learning improves risk stratification of high-grade serous ovarian cancer. Nat. Cancer 3, 723–733 (2022).
Cheerla, A. & Gevaert, O. Deep learning with multimodal representation for pancancer prognosis prediction. Bioinformatics 35, i446–i454 (2019).
Carrillo-Perez, F. et al. Machine-learning-based late fusion on multi-omics and multi-scale data for non-small-cell lung cancer diagnosis. J. Pers. Med. 12, 601 (2022).
Vaswani, A. et al. in Advances in Neural Information Processing Systems Vol. 30 (eds Guyon, I. et al.) (Curran Associates, 2017).
Wang, L. et al. Transformer-based deep learning model for the diagnosis of suspected lung cancer in primary care based on electronic health record data. eBioMedicine 110, 105442 (2024).
Wagner, S. J. et al. Transformer-based biomarker prediction from colorectal cancer histology: a large-scale multicentric study. Cancer Cell 41, 1650–1661.e4 (2023).
Cai, Z. et al. DeePathNet: a transformer-based deep learning model integrating multi-omic data with cancer pathways. Cancer Res. Commun. 4, 3151–3164 (2024).
Raissi, M., Perdikaris, P. & Karniadakis, G. E. Physics-informed neural networks: a deep learning framework for solving forward and inverse problems involving nonlinear partial differential equations. J. Comput. Phys. 378, 686–707 (2019).
Sarabian, M., Babaee, H. & Laksari, K. Physics-informed neural networks for brain hemodynamic predictions using medical imaging. IEEE Trans. Med. Imaging 41, 2285–2303 (2022).
Sel, K., Mohammadi, A., Pettigrew, R. I. & Jafari, R. Physics-informed neural networks for modeling physiological time series for cuffless blood pressure estimation. npj Digit. Med. 6, 110 (2023).
Ragoza, M. & Batmanghelich, K. in Medical Image Computing and Computer Assisted Intervention — MICCAI 2023 Vol. 14229 (eds Greenspan, H. et al.) 333–343 (Springer, 2023).
Rodrigues, J. A. Using physics-informed neural networks (PINNs) for tumor cell growth modeling. Mathematics 12, 1195 (2024).
Mukhmetov, O. et al. Physics-informed neural network for fast prediction of temperature distributions in cancerous breasts as a potential efficient portable AI-based diagnostic tool. Comput. Methods Prog. Biomed. 242, 107834 (2023).
Metzcar, J., Jutzeler, C. R., Macklin, P., Köhn-Luque, A. & Brüningk, S. C. A review of mechanistic learning in mathematical oncology. Front. Immunol. 15, 1363144 (2024).
Gu, Y. & Ng, M. K. Deep neural networks for solving large linear systems arising from high-dimensional problems. SIAM J. Sci. Comput. 45, A2356–A2381 (2023).
Ghaffarizadeh, A., Heiland, R., Friedman, S. H., Mumenthaler, S. M. & Macklin, P. PhysiCell: an open source physics-based cell simulator for 3-D multicellular systems. PLoS Comput. Biol. 14, e1005991 (2018).
Fernandes, M. R., Aggarwal, P., Costa, R. G. F., Cole, A. M. & Trinchieri, G. Targeting the gut microbiota for cancer therapy. Nat. Rev. Cancer 22, 703–722 (2022).
Park, E. M. et al. Targeting the gut and tumor microbiota in cancer. Nat. Med. 28, 690–703 (2022).
Battaglia, T. W. et al. A pan-cancer analysis of the microbiome in metastatic cancer. Cell 187, 2324–2335.e19 (2024).
Davar, D. et al. Fecal microbiota transplant overcomes resistance to anti-PD-1 therapy in melanoma patients. Science 371, 595–602 (2021).
Baruch, E. N. et al. Fecal microbiota transplant promotes response in immunotherapy-refractory melanoma patients. Science 371, 602–609 (2021).
Liang, H. et al. Predicting cancer immunotherapy response from gut microbiomes using machine learning models. Oncotarget 13, 876–889 (2022).
Marino, S., Baxter, N. T., Huffnagle, G. B., Petrosino, J. F. & Schloss, P. D. Mathematical modeling of primary succession of murine intestinal microbiota. Proc. Natl Acad. Sci. USA 111, 439–444 (2014).
Bucci, V. et al. MDSINE: microbial dynamical systems inference engine for microbiome time-series analyses. Genome Biol. 17, 121 (2016).
Rolig, A. S., Parthasarathy, R., Burns, A. R., Bohannan, B. J. M. & Guillemin, K. Individual members of the microbiota disproportionately modulate host innate immune responses. Cell Host Microbe 18, 613–620 (2015).
Stein, R. R. et al. Computer-guided design of optimal microbial consortia for immune system modulation. eLife 7, e30916 (2018).
Hadjigeorgiou, A.G. et al. Dissecting the impact of the gut microbiome on cancer immunotherapy. Preprint at Research Square https://doi.org/10.21203/rs.3.rs-3647386/v1 (2023).
Meng, X. et al. The application of large language models in medicine: a scoping review. iScience 27, 109713 (2024).
Rao, A. et al. Evaluating GPT as an adjunct for radiologic decision making: GPT-4 versus GPT-3.5 in a breast imaging pilot. J. Am. Coll. Radiol. 20, 990–997 (2023).
Rao, A. et al. Assessing the utility of ChatGPT throughout the entire clinical workflow: development and usability study. J. Med. Internet Res. 25, e48659 (2023).
Lawson McLean, A., Wu, Y., Lawson McLean, A. C. & Hristidis, V. Large language models as decision aids in neuro-oncology: a review of shared decision-making applications. J. Cancer Res. Clin. Oncol. 150, 139 (2024).
Longwell, J. B. et al. Performance of large language models on medical oncology examination questions. JAMA Netw. Open 7, e2417641 (2024).
Acknowledgements
We would like to thank J. Baish, L.L. Munn, F. Yuan and W. Lotter for their helpful comments during the preparation of this article. This work was supported through NIH grant R35-CA197743, and in part through grants R01CA259253, R01-CA269672, R01-NS118929, U01-CA224348 and U01-CA261842 and by the National Foundation for Cancer Research, Harvard Ludwig Cancer Center, Nile Albright Research Foundation and Jane’s Trust Foundation (to R.K.J.). Also, this work has received funding from the European Research Council (ERC) under the European Union’s Horizon 2020 research and innovation programme (grant agreement no. 863955 to T.S.).
Author information
Authors and Affiliations
Contributions
C.H., A.G.H., C.V., A.S.K., T.S. and R.K.J. researched data for the article. C.H., A.G.H., C.V., A.S.K., T.S. and R.K.J. contributed substantially to discussion of the content. All authors wrote the article.
Corresponding authors
Ethics declarations
Competing interests
R.K.J. received consultant fees from DynamiCure, SPARC and SynDevRx; owns equity in Accurius, Enlight, SynDevRx; and served on the Boards of Trustees of Tekla Healthcare Investors, Tekla Life Sciences Investors, Tekla Healthcare Opportunities Fund and Tekla World Healthcare Fund; and received grants from Boehringer Ingelheim and Sanofi. Neither any reagent nor any funding from these organizations was used in this study. Other co-authors have no conflict of interests to declare.
Peer review
Peer review information
Nature Reviews Cancer thanks Georgios Lolas, Aleksander Popel who co-reviewed with Theinmozhi Arulraj and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Glossary
- Anastomosis
-
The process of joining of two blood vessels.
- Autologous chemotaxis
-
The process whereby a cell can receive directional cues while at the same time be the source of such cues.
- Convection
-
The bulk motion of agents driven by the fluid velocity, which is proportional to fluid pressure gradient.
- Finite element modelling
-
A computational numerical method used to solve partial differential equations by discretizing a structure into smaller parts called finite elements.
- Hypoperfusion
-
Insufficient blood supply to tissues.
- Immune checkpoint inhibitors
-
Therapeutic antibodies that inhibit regulators of the immune system, immune checkpoint proteins, that when stimulated suppress immune cell.
- Interstitial fluid pressure
-
The pressure exerted by the fluid present in the spaces between cells in a tissue.
- Pharmacodynamics
-
The study of the effect of the drug at the site of action.
- Pharmacokinetics
-
The study of how drugs are absorbed, distributed, metabolized and eliminated in the body.
- Random-walk model
-
A stochastic model that simulates the behaviour of cells, including cancer and endothelial cells, by capturing probabilistic processes such as cell migration, proliferation and the formation of new blood vessels.
- Solid stresses
-
The mechanical forces (per unit area), in the context of tumour biology, refers to the mechanical stresses generated during tumour growth due to the deformation of the structural components of the tissue.
- Targeted radionucleotide therapy
-
A systemic treatment that uses radiolabeled drugs (for example, antibodies, antibody fragments, proteins and peptides) that are cancer specific to deliver cytotoxic doses of radiation to kill cancer cells.
- Tensotaxis
-
The migration of cells in response to mechanical stress.
- Vascular normalization
-
Restoring normal structure and function of blood vessels in tumours (for example, with judicious use of anti-angiogenic agents).
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Harkos, C., Hadjigeorgiou, A.G., Voutouri, C. et al. Using mathematical modelling and AI to improve delivery and efficacy of therapies in cancer. Nat Rev Cancer 25, 324–340 (2025). https://doi.org/10.1038/s41568-025-00796-w
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41568-025-00796-w
This article is cited by
-
Design parameter effects on controlled drug delivery through implantable hydrogels
Drug Delivery and Translational Research (2026)
-
Antibody–drug conjugates in cancer therapy: current landscape, challenges, and future directions
Molecular Cancer (2025)
-
AI-engineered multifunctional nanoplatforms: synergistically bridging precision diagnosis and intelligent therapy in next-generation oncology
Journal of Nanobiotechnology (2025)
-
Artificial intelligence based quantification of T lymphocyte infiltrate predicts prognosis in high grade breast cancer using deep learning and statistical validation
Discover Oncology (2025)
-
Tackling tumor hypoxia: advances in breaking the oncogenic HIF-1α–p300/CBP alliance
Investigational New Drugs (2025)