Abstract
RNA adopts 3D structures that confer varied functional roles in human biology and dysfunction in disease. Approaches to therapeutically target RNA structures with small molecules are being actively pursued, aided by key advances in the field including the development of computational tools that predict evolutionarily conserved RNA structures, as well as strategies that expand mode of action and facilitate interactions with cellular machinery. Existing RNA-targeted small molecules use a range of mechanisms including directing splicing — by acting as molecular glues with cellular proteins (such as branaplam and the FDA-approved risdiplam), inhibition of translation of undruggable proteins and deactivation of functional structures in noncoding RNAs. Here, we describe strategies to identify, validate and optimize small molecules that target the functional transcriptome, laying out a roadmap to advance these agents into the next decade.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Jones, D., Metzger, H. J., Schatz, A. & Waksman, S. A. Control of Gram-negative bacteria in experimental animals by streptomycin. Science 100, 103–105 (1944).
Waksman, S. A., Reilly, H. C. & Schatz, A. Strain specificity and production of antibiotic substances: V. Strain resistance of bacteria to antibiotic substances, especially to streptomycin. Proc. Natl Acad. Sci. USA 31, 157–164 (1945).
Holley, R. W. et al. Structure of a ribonucleic acid. Science 147, 1462 (1965).
Zaug, A. J. & Cech, T. R. The intervening sequence RNA of Tetrahymena is an enzyme. Science 231, 470–475 (1986).
Stark, B. C., Kole, R., Bowman, E. J., Altman, S. & Altman, S. Ribonuclease P: an enzyme with an essential RNA component. Proc. Natl Acad. Sci. USA 75, 3717–3721 (1978).
Encode Project Consortium. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Tsai, M. C. et al. Long noncoding RNA as modular scaffold of histone modification complexes. Science 329, 689–693 (2010).
He, L. & Hannon, G. J. MicroRNAs: small RNAs with a big role in gene regulation. Nat. Rev. Genet. 5, 522–531 (2004).
Bennett, C. F. Therapeutic antisense oligonucleotides are coming of age. Annu. Rev. Med. 70, 307–321 (2019).
Luther, D. C., Lee, Y. W., Nagaraj, H., Scaletti, F. & Rotello, V. M. Delivery approaches for CRISPR/Cas9 therapeutics in vivo: advances and challenges. Expert Opin. Drug Deliv. 15, 905–913 (2018).
Howe, J. A. et al. Selective small-molecule inhibition of an RNA structural element. Nature 526, 672–677 (2015). Describes the discovery of ribocil, a selective chemical modulator of bacterial riboflavin riboswitches.
Dibrov, S. M. et al. Hepatitis C virus translation inhibitors targeting the internal ribosomal entry site. J. Med. Chem. 57, 1694–1707 (2014).
Zhang, P. et al. Translation of the intrinsically disordered protein alpha-synuclein is inhibited by a small molecule targeting its structured mRNA. Proc. Natl Acad. Sci. USA 117, 1457–1467 (2020). Describes the design of a small molecule that inhibits the translation of the undruggable, intrinsically disordered protein α-synuclein by targeting its mRNA.
Ratni, H. et al. Discovery of risdiplam, a selective survival of motor neuron-2 (SMN2) gene splicing modifier for the treatment of spinal muscular atrophy (SMA). J. Med. Chem. 61, 6501–6517 (2018). Discovery of the first FDA-approved small molecule (risdiplam) for the treatment of SMA by directing the alternative splicing of SMN2 to produce a functionally equivalent SMN1 protein.
Palacino, J. et al. SMN2 splice modulators enhance U1-pre-mRNA association and rescue SMA mice. Nat. Chem. Biol. 12, 304–304 (2016). The discovery of the SMN2 splicing modulator branaplam for the treatment of SMA.
Becquart, C. et al. Exploring heterocycle-spermine conjugates as modulators of oncogenic microRNAs biogenesis. ACS Omega 3, 16500–16508 (2018). Design of a heterocycle–spermine conjugate that inhibits the biogenesis of oncogenic miRNAs.
Turner, D. H. & Mathews, D. H. NNDB: the nearest neighbor parameter database for predicting stability of nucleic acid secondary structure. Nucleic Acids Res. 38, D280–D282 (2010).
Tinoco, I. Jr, Uhlenbeck, O. C. & Levine, M. D. Estimation of secondary structure in ribonucleic acids. Nature 230, 362–367 (1971).
Freier, S. M. et al. Improved free-energy parameters for predictions of RNA duplex stability. Proc. Natl Acad. Sci. USA 83, 9373–9377 (1986).
Zuker, M. mFold web server for nucleic acid folding and hybridization prediction. Nucleic Acids Res. 31, 3406–3415 (2003).
Lorenz, R. et al. ViennaRNA Package 2.0. Algorithms Mol. Biol. 6, 26 (2011).
Mathews, D. H. et al. Incorporating chemical modification constraints into a dynamic programming algorithm for prediction of RNA secondary structure. Proc. Natl Acad. Sci. USA 101, 7287–7292 (2004).
Peattie, D. A. Direct chemical method for sequencing RNA. Proc. Natl Acad. Sci. USA 76, 1760–1764 (1979).
Bevilacqua, P. C., Ritchey, L. E., Su, Z. & Assmann, S. M. Genome-wide analysis of RNA secondary structure. Annu. Rev. Genet. 50, 235–266 (2016).
Sun, L. et al. RNA structure maps across mammalian cellular compartments. Nat. Struct. Mol. Biol. 26, 322–330 (2019).
Marinus, T., Fessler, A. B., Ogle, C. A. & Incarnato, D. A novel SHAPE reagent enables the analysis of RNA structure in living cells with unprecedented accuracy. Nucleic Acids Res. 49, e34 (2021).
Deigan, K. E., Li, T. W., Mathews, D. H. & Weeks, K. M. Accurate SHAPE-directed RNA structure determination. Proc. Natl Acad. Sci. USA 106, 97–102 (2009).
Wells, S. E., Hughes, J. M., Igel, A. H. & Ares, M. Jr. Use of dimethyl sulfate to probe RNA structure in vivo. Methods Enzymol. 318, 479–493 (2000).
Spitale, R. C., Flynn, R. A., Torre, E. A., Kool, E. T. & Chang, H. Y. RNA structural analysis by evolving SHAPE chemistry. Wiley Interdiscip. Rev. RNA 5, 867–881 (2014).
Eddy, S. R. Computational analysis of conserved RNA secondary structure in transcriptomes and genomes. Annu. Rev. Biophys. 43, 433–456 (2014).
Eddy, S. R. & Durbin, R. RNA sequence analysis using covariance models. Nucleic Acids Res. 22, 2079–2088 (1994).
Gutell, R. R., Lee, J. C. & Cannone, J. J. The accuracy of ribosomal RNA comparative structure models. Curr. Opin. Struct. Biol. 12, 301–310 (2002).
Woese, C. R., Gutell, R., Gupta, R. & Noller, H. F. Detailed analysis of the higher-order structure of 16S-like ribosomal ribonucleic acids. Microbiol. Rev. 47, 621–669 (1983).
Mathews, D. H. & Turner, D. H. Dynalign: an algorithm for finding the secondary structure common to two RNA sequences. J. Mol. Biol. 317, 191–203 (2002).
Havgaard, J. H., Lyngsø, R. B. & Gorodkin, J. The FOLDALIGN web server for pairwise structural RNA alignment and mutual motif search. Nucleic Acids Res. 33, W650–W653 (2005).
Hofacker, I. L., Fekete, M. & Stadler, P. F. Secondary structure prediction for aligned RNA sequences. J. Mol. Biol. 319, 1059–1066 (2002).
Watkins, A. M., Rangan, R. & Das, R. FARFAR2: improved de novo Rosetta prediction of complex global RNA folds. Structure 28, 963–976 (2020).
Parisien, M. & Major, F. The MC-Fold and MC-Sym pipeline infers RNA structure from sequence data. Nature 452, 51–55 (2008).
Sharma, S., Ding, F. & Dokholyan, N. V. iFoldRNA: three-dimensional RNA structure prediction and folding. Bioinformatics 24, 1951–1952 (2008).
Capriotti, E., Norambuena, T., Marti-Renom, M. A. & Melo, F. All-atom knowledge-based potential for RNA structure prediction and assessment. Bioinformatics 27, 1086–1093 (2011).
Townshend, R. J. L. et al. Geometric deep learning of RNA structure. Science 373, 1047–1051 (2021).
Furtig, B., Richter, C., Wohnert, J. & Schwalbe, H. NMR spectroscopy of RNA. Chembiochem 4, 936–962 (2003).
Lietzke, S. E., Barnes, C. L. & Kundrot, C. E. Crystallization and structure determination of RNA. Curr. Opin. Struct. Biol. 5, 645–649 (1995).
Spahn, C. M. & Penczek, P. A. Exploring conformational modes of macromolecular assemblies by multiparticle cryo-EM. Curr. Opin. Struct. Biol. 19, 623–631 (2009).
Zhang, K. et al. Cryo-EM structure of a 40 kDa SAM-IV riboswitch RNA at 3.7 Å resolution. Nat. Commun. 10, 5511 (2019).
Kappel, K. et al. Accelerated cryo-EM-guided determination of three-dimensional RNA-only structures. Nat. Methods 17, 699–707 (2020).
Ding, Y., Chan, C. Y. & Lawrence, C. E. Sfold web server for statistical folding and rational design of nucleic acids. Nucleic Acids Res. 32, W135–W141 (2004).
Do, C. B., Woods, D. A. & Batzoglou, S. CONTRAfold: RNA secondary structure prediction without physics-based models. Bioinformatics 22, e90–e98 (2006).
Rivas, E., Clements, J. & Eddy, S. R. A statistical test for conserved RNA structure shows lack of evidence for structure in lncRNAs. Nat. Methods 14, 45–48 (2017). A statistical method that evaluates the functional relevence of a given RNA structure based on its phylogenetic conservation.
Nawrocki, E. P. et al. Rfam 12.0: updates to the RNA families database. Nucleic Acids Res. 43, D130–D137 (2015).
Fang, R., Moss, W. N., Rutenberg-Schoenberg, M. & Simon, M. D. Probing Xist RNA structure in cells using targeted structure-Seq. PLoS Genet. 11, e1005668 (2015).
Somarowthu, S. et al. HOTAIR forms an intricate and modular secondary structure. Mol. Cell 58, 353–361 (2015).
Novikova, I. V., Hennelly, S. P. & Sanbonmatsu, K. Y. Structural architecture of the human long non-coding RNA, steroid receptor RNA activator. Nucleic Acids Res. 40, 5034–5051 (2012).
Rivas, E., Clements, J. & Eddy, S. R. Estimating the power of sequence covariation for detecting conserved RNA structure. Bioinformatics 36, 3072–3076 (2020).
Rouskin, S., Zubradt, M., Washietl, S., Kellis, M. & Weissman, J. S. Genome-wide probing of RNA structure reveals active unfolding of mRNA structures in vivo. Nature 505, 701–705 (2014).
Ding, Y. et al. In vivo genome-wide profiling of RNA secondary structure reveals novel regulatory features. Nature 505, 696–700 (2014).
Wan, Y. et al. Landscape and variation of RNA secondary structure across the human transcriptome. Nature 505, 706–709 (2014).
Sobczak, K. & Krzyzosiak, W. J. CAG repeats containing CAA interruptions form branched hairpin structures in spinocerebellar ataxia type 2 transcripts. J. Biol. Chem. 280, 3898–3910 (2005).
Busan, S. & Weeks, K. M. Role of context in RNA structure: flanking sequences reconfigure CAG motif folding in huntingtin exon 1 transcripts. Biochemistry 52, 8219–8225 (2013).
Kole, R., Krainer, A. R. & Altman, S. RNA therapeutics: beyond RNA interference and antisense oligonucleotides. Nat. Rev. Drug Discov. 11, 125–140 (2012).
Sudarsan, N., Barrick, J. E. & Breaker, R. R. Metabolite-binding RNA domains are present in the genes of eukaryotes. RNA 9, 644–647 (2003).
Andrews, R. J., Roche, J. & Moss, W. N. ScanFold: an approach for genome-wide discovery of local RNA structural elements-applications to Zika virus and HIV. PeerJ 6, e6136 (2018).
Moss, W. N., Priore, S. F. & Turner, D. H. Identification of potential conserved RNA secondary structure throughout influenza A coding regions. RNA 17, 991–1011 (2011).
O’Leary, C. et al. RNA structural analysis of the MYC mRNA reveals conserved motifs that affect gene expression. PLoS ONE 14, e0213758 (2019).
Disney, M. D. Targeting RNA with small molecules to capture opportunities at the Intersection of chemistry, biology, and medicine. J. Am. Chem. Soc. 141, 6776–6790 (2019).Describes the sequence-based design of RNA structure-specific small molecules, including the 2DCS selection platform as well as target validation techiniques.
Naryshkin, N. A. et al. Motor neuron disease. SMN2 splicing modifiers improve motor function and longevity in mice with spinal muscular atrophy. Science 345, 688–693 (2014).
Parsons, J. et al. Conformational inhibition of the hepatitis C virus internal ribosome entry site RNA. Nat. Chem. Biol. 5, 823–825 (2009).
Carnevali, M., Parsons, J., Wyles, D. L. & Hermann, T. A modular approach to synthetic RNA binders of the hepatitis C virus internal ribosome entry site. Chembiochem 11, 1364–1367 (2010).
Chen, C. Z. et al. Two high-throughput screening assays for aberrant RNA-protein interactions in myotonic dystrophy type 1. Anal. Bioanal. Chem. 402, 1889–1898 (2012).
Chen, J. L. et al. Design, optimization, and study of small molecules that target tau pre-mRNA and affect splicing. J. Am. Chem. Soc. 142, 8706–8727 (2020).
Corbett, P. T. et al. Dynamic combinatorial chemistry. Chem. Rev. 106, 3652–3711 (2006).
Motoyaji, T. Revolution of small molecule drug discovery by affinity selection-mass spectrometry technology. Chem. Pharm. Bull. 68, 191–193 (2020).
Kiernan, U. A., Nedelkov, D., Niederkofler, E. E., Tubbs, K. A. & Nelson, R. W. High-throughput affinity mass spectrometry. Methods Mol. Biol. 328, 141–150 (2006).
Annis, D. A. et al. An affinity selection–mass spectrometry method for the identification of small molecule ligands from self-encoded combinatorial libraries. Int. J. Mass. Spectrom. 238, 77–83 (2004).
Rizvi, N. F. et al. Discovery of selective RNA-binding small molecules by affinity-selection mass spectrometry. ACS Chem. Biol. 13, 820–831 (2018).
Rizvi, N. F. & Nickbarg, E. B. RNA-ALIS: methodology for screening soluble RNAs as small molecule targets using ALIS affinity-selection mass spectrometry. Methods 167, 28–38 (2019).
Stover, J. S., Shi, J., Jin, W., Vogt, P. K. & Boger, D. L. Discovery of inhibitors of aberrant gene transcription from libraries of DNA binding molecules: inhibition of LEF-1-mediated gene transcription and oncogenic transformation. J. Am. Chem. Soc. 131, 3342–3348 (2009).
Tran, T. & Disney, M. D. Identifying the preferred RNA motifs and chemotypes that interact by probing millions of combinations. Nat. Commun. 3, 1125 (2012).
Asare-Okai, P. N. & Chow, C. S. A modified fluorescent intercalator displacement assay for RNA ligand discovery. Anal. Biochem. 408, 269–276 (2011). Demonstrates the successful application of ALIS to RNA targets.
Wang, Z. F. et al. The hairpin form of r(G4C2)exp in c9ALS/FTD is repeat-associated non-ATG translated and a target for bioactive small molecules. Cell Chem. Biol. 26, 179–190 (2019).
Zhang, J., Umemoto, S. & Nakatani, K. Fluorescent indicator displacement assay for ligand-RNA interactions. J. Am. Chem. Soc. 132, 3660–3661 (2010).
Wicks, S. L. & Hargrove, A. E. Fluorescent indicator displacement assays to identify and characterize small molecule interactions with RNA. Methods 167, 3–14 (2019).
Jones, A. C. & Neely, R. K. 2-Aminopurine as a fluorescent probe of DNA conformation and the DNA-enzyme interface. Q. Rev. Biophys. 48, 244–279 (2015).
Jean, J. M. & Hall, K. B. 2-Aminopurine fluorescence quenching and lifetimes: role of base stacking. Proc. Natl Acad. Sci. USA 98, 37–41 (2001).
Soulière, M. F., Haller, A., Rieder, R. & Micura, R. A powerful approach for the selection of 2-aminopurine substitution sites to investigate RNA folding. J. Am. Chem. Soc. 133, 16161–16167 (2011).
Soulière, M. F. & Micura, R. Use of SHAPE to select 2AP substitution sites for RNA-ligand interactions and dynamics studies. Methods Mol. Biol. 1103, 227–239 (2014).
Froeyen, M. & Herdewijn, P. RNA as a target for drug design, the example of Tat-TAR interaction. Curr. Top. Med. Chem. 2, 1123–1145 (2002).
Pascale, L. et al. Deciphering structure-activity relationships in a series of Tat/TAR inhibitors. J. Biomol. Struct. Dyn. 34, 2327–2338 (2016).
Yang, M. Discoveries of Tat-TAR interaction inhibitors for HIV-1. Curr. Drug Targets Infect. Disord. 5, 433–444 (2005).
Kumar, A. et al. Chemical correction of pre-mRNA splicing defects associated with sequestration of muscleblind-like 1 protein by expanded r(CAG)-containing transcripts. ACS Chem. Biol. 7, 496–505 (2012).
MacBeath, G., Koehler, A. N. & Schreiber, S. L. Printing small molecules as microarrays and detecting protein- ligand interactions en masse. J. Am. Chem. Soc. 121, 7967–7968 (1999).
Fazio, F., Bryan, M. C., Blixt, O., Paulson, J. C. & Wong, C. H. Synthesis of sugar arrays in microtiter plate. J. Am. Chem. Soc. 124, 14397–14402 (2002).
Peng, B., Thorsell, A. G., Karlberg, T., Schuler, H. & Yao, S. Q. Small molecule microarray based discovery of PARP14 inhibitors. Angew. Chem. Int. Ed. Engl. 56, 248–253 (2017).
Wang, Z. et al. Microarray based screening of peptide nano probes for HER2 positive tumor. Anal. Chem. 87, 8367–8372 (2015).
Disney, M. D. & Seeberger, P. H. Aminoglycoside microarrays to explore interactions of antibiotics with RNAs and proteins. Chemistry 10, 3308–3314 (2004).
Disney, M. D. & Barrett, O. J. An aminoglycoside microarray platform for directly monitoring and studying antibiotic resistance. Biochemistry 46, 11223–11230 (2007).
Abulwerdi, F. A. et al. Selective small-molecule targeting of a triple helix encoded by the long noncoding RNA, MALAT1. ACS Chem. Biol. 14, 223–235 (2019).
Connelly, C. M., Abulwerdi, F. A. & Schneekloth, J. S. Jr. Discovery of RNA binding small molecules using small molecule microarrays. Methods Mol. Biol. 1518, 157–175 (2017). A detailed description of a small molecule microarray method for the identification of small-molecule RNA binders.
Connelly, C. M., Boer, R. E., Moon, M. H., Gareiss, P. & Schneekloth, J. S. Jr Discovery of inhibitors of microRNA-21 processing using small molecule microarrays. ACS Chem. Biol. 12, 435–443 (2017).
Connelly, C. M. et al. Synthetic ligands for PreQ(1) riboswitches provide structural and mechanistic insights into targeting RNA tertiary structure. Nat. Commun. 10, 1501 (2019). An example of how structural studies can enhance the understanding of ligand binding and function, in the context of PreQ1 riboswitch.
Sztuba-Solinska, J. et al. Identification of biologically active, HIV TAR RNA-binding small molecules using small molecule microarrays. J. Am. Chem. Soc. 136, 8402–8410 (2014). A successful application of small-molecule microarray for the discovery of a HIV-1 TAR RNA binder.
Tran, B. et al. Parallel discovery strategies provide a basis for riboswitch ligand design. Cell Chem. Biol. 27, 1241–1249.e1244 (2020).
Labuda, L. P., Pushechnikov, A. & Disney, M. D. Small molecule microarrays of RNA-focused peptoids help identify inhibitors of a pathogenic group I intron. ACS Chem. Biol. 4, 299–307 (2009).
Velagapudi, S. P. et al. Approved anti-cancer drugs target oncogenic non-coding RNAs. Cell Chem. Biol. 25, 1086–1094.e1087 (2018).
Sreeramulu, S. et al. Exploring the druggability of conserved RNA regulatory elements in the SARS-CoV-2 genome. Angew. Chem. Int. Ed. Engl. 60, 19191–19200 (2021).
Parker, C. G. et al. Ligand and target discovery by fragment-based screening in human cells. Cell 168, 527–541 e529 (2017).
Wang, Y. et al. Expedited mapping of the ligandable proteome using fully functionalized enantiomeric probe pairs. Nat. Chem. 11, 1113–1123 (2019).
Suresh, B. M. et al. A general fragment-based approach to identify and optimize bioactive ligands targeting RNA. Proc. Natl Acad. Sci. USA 117, 33197–33203 (2020). The first application of fully functionalized fragments to the discovery of RNA ligands.
Benhamou, R. I. et al. DNA-encoded library-versus-RNA-encoded library selection enables design of an oncogenic non-coding RNA inhibitor. Proc. Natl Acad. Sci. USA 119, e2114971119 (2022).
Litovchick, A. et al. Novel nucleic acid binding small molecules discovered using DNA-encoded chemistry. Molecules 24, 2026 (2019).
Disney, M. D., Velagapudi, S. P., Li, Y., Costales, M. G. & Childs-Disney, J. L. Identifying and validating small molecules interacting with RNA (SMIRNAs). Methods Enzymol. 623, 45–66 (2019).
Disney, M. D. et al. Two-dimensional combinatorial screening identifies specific aminoglycoside-RNA internal loop partners. J. Am. Chem. Soc. 130, 11185–11194 (2008).
Costales, M. G., Childs-Disney, J. L., Haniff, H. S. & Disney, M. D. How we think about targeting RNA with small molecules. J. Med. Chem. 63, 8880–8900 (2020).
Velagapudi, S. P. et al. Defining RNA-small molecule affinity landscapes enables design of a small molecule inhibitor of an oncogenic noncoding RNA. ACS Cent. Sci. 3, 205–216 (2017).
Thomas, J. R. & Hergenrother, P. J. Targeting RNA with small molecules. Chem. Rev. 108, 1171–1224 (2008).
Kuntz, I. D. Structure-based strategies for drug design and discovery. Science 257, 1078–1082 (1992).
Sledz, P. & Caflisch, A. Protein structure-based drug design: from docking to molecular dynamics. Curr. Opin. Struct. Biol. 48, 93–102 (2018).
Yadav, M., Dhagat, S. & Eswari, J. S. Structure based drug design and molecular docking studies of anticancer molecules paclitaxel, etoposide and topotecan using novel ligands. Curr. Drug Discov. Technol. 17, 183–190 (2020).
Batool, M., Ahmad, B. & Choi, S. A structure-based drug discovery paradigm. Int. J. Mol. Sci. 20, 2783 (2019).
Ganser, L. R. et al. Probing RNA conformational equilibria within the functional cellular context. Cell Rep. 30, 2472–2480 (2020). An NMR-based method that assesses the energetic properties of transient RNA conformations in vitro.
Ganser, L. R. et al. High-performance virtual screening by targeting a high-resolution RNA dynamic ensemble. Nat. Struct. Mol. Biol. 25, 425–434 (2018).
Shi, H. et al. Rapid and accurate determination of atomistic RNA dynamic ensemble models using NMR and structure prediction. Nat. Commun. 11, 5531 (2020).
Liu, B., Shi, H. & Al-Hashimi, H. M. Developments in solution-state NMR yield broader and deeper views of the dynamic ensembles of nucleic acids. Curr. Opin. Struct. Biol. 70, 16–25 (2021).
Zhang, Q., Sun, X., Watt, E. D. & Al-Hashimi, H. M. Resolving the motional modes that code for RNA adaptation. Science 311, 653–656 (2006).
Frank, A. T., Stelzer, A. C., Al-Hashimi, H. M. & Andricioaei, I. Constructing RNA dynamical ensembles by combining MD and motionally decoupled NMR RDCs: new insights into RNA dynamics and adaptive ligand recognition. Nucleic Acids Res. 37, 3670–3679 (2009).
Stelzer, A. C. et al. Discovery of selective bioactive small molecules by targeting an RNA dynamic ensemble. Nat. Chem. Biol. 7, 553–559 (2011). An example of using structural docking and virtual screening for the discovery of an HIV TAR RNA inhibitor.
Bush, J. A. et al. Systematically studying the effect of small molecules interacting with RNA in cellular and preclinical models. ACS Chem. Biol. 16, 1111–1127 (2021).
Winkler, W. C., Cohen-Chalamish, S. & Breaker, R. R. An mRNA structure that controls gene expression by binding FMN. Proc. Natl Acad. Sci. USA 99, 15908–15913 (2002).
Sudarsan, N., Cohen-Chalamish, S., Nakamura, S., Emilsson, G. M. & Breaker, R. R. Thiamine pyrophosphate riboswitches are targets for the antimicrobial compound pyrithiamine. Chem. Biol. 12, 1325–1335 (2005).
Lin, A. et al. Off-target toxicity is a common mechanism of action of cancer drugs undergoing clinical trials. Sci. Transl. Med. 11, eaaw8412 (2019).
Osborn, M. F., White, J. D., Haley, M. M. & DeRose, V. J. Platinum-RNA modifications following drug treatment in S. cerevisiae identified by click chemistry and enzymatic mapping. ACS Chem. Biol. 9, 2404–2411 (2014).
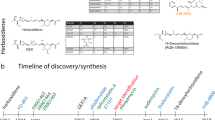
Mortison, J. D. et al. Tetracyclines modify translation by targeting key human rRNA substructures. Cell Chem. Biol. 25, 1506–1518 (2018).
Balaratnam, S. et al. A chemical probe based on the PreQ1 metabolite enables transcriptome-wide mapping of binding sites. Nat. Commun. 12, 5856 (2021).
Guan, L. & Disney, M. D. Covalent small-molecule–RNA complex formation enables cellular profiling of small-molecule–RNA interactions. Angew. Chem. Int. Ed. Engl. 52, 10010–10013 (2013).
Zarrinkar, P. P., Wang, J. & Williamson, J. R. Slow folding kinetics of RNase P RNA. RNA 2, 564–573 (1996).
Zarrinkar, P. P. & Williamson, J. R. Kinetic intermediates in RNA folding. Science 265, 918–924 (1994).
Boger, D. L. & Cai, H. Bleomycin: synthetic and mechanistic studies. Angew. Chem. Int. Ed. Engl. 38, 448–476 (1999).
Ishida, R. & Takahashi, T. Increased DNA chain breakage by combined action of bleomycin and superoxide radical. Biochem. Biophys. Res. Commun. 66, 1432–1438 (1975).
Burger, R. M. Cleavage of nucleic acids by bleomycin. Chem. Rev. 98, 1153–1170 (1998).
Magliozzo, R. S., Peisach, J. & Ciriolo, M. R. Transfer-RNA is cleaved by activated bleomycin. Mol. Pharmacol. 35, 428–432 (1989).
Carter, B. J. et al. Site-specific cleavage of RNA by Fe(II).bleomycin. Proc. Natl Acad. Sci. USA 87, 9373–9377 (1990).
Abraham, A. T., Lin, J. J., Newton, D. L., Rybak, S. & Hecht, S. M. RNA cleavage and inhibition of protein synthesis by bleomycin. Chem. Biol. 10, 45–52 (2003).
Costales, M. G. et al. Small-molecule targeted recruitment of a nuclease to cleave an oncogenic RNA in a mouse model of metastatic cancer. Proc. Natl Acad. Sci. USA 117, 2406–2411 (2020).
Costales, M. G., Matsumoto, Y., Velagapudi, S. P. & Disney, M. D. Small molecule targeted recruitment of a nuclease to RNA. J. Am. Chem. Soc. 140, 6741–6744 (2018).
Bisbal, C., Martinand, C., Silhol, M., Lebleu, B. & Salehzada, T. Cloning and characterization of a RNAse L inhibitor. A new component of the interferon-regulated 2-5A pathway. J. Biol. Chem. 270, 13308–13317 (1995).
Li, X. L. et al. RNase-L-dependent destabilization of interferon-induced mRNAs. A role for the 2-5A system in attenuation of the interferon response. J. Biol. Chem. 275, 8880–8888 (2000).
Zhou, A., Molinaro, R. J., Malathi, K. & Silverman, R. H. Mapping of the human RNASEL promoter and expression in cancer and normal cells. J. Interferon Cytokine Res. 25, 595–603 (2005).
Silverman, R. H. Viral encounters with 2′,5′-oligoadenylate synthetase and RNase L during the interferon antiviral response. J. Virol. 81, 12720–12729 (2007).
Meyer, S. M. et al. Small molecule recognition of disease-relevant RNA structures. Chem. Soc. Rev. 49, 7167–7199 (2020).
Zhang, P. et al. Reprogramming of protein-targeted small-molecule medicines to RNA by ribonuclease recruitment. J. Am. Chem. Soc. 143, 13044–13055 (2021). Shows that known drugs can be reprogrammed to selectively target RNA by its conversion into a ribonuclease targeting chimera (RIBOTAC) degrader.
Lorenz, M. O. Methods of measuring the concentration of wealth. Pub. Am. Stat. Assoc. 9, 209–219 (1905).
Ursu, A. et al. Gini coefficients as a single value metric to define chemical probe selectivity. ACS Chem. Biol. 15, 2031–2040 (2020).
Graczyk, P. P. Gini coefficient: a new way to express selectivity of kinase inhibitors against a family of kinases. J. Med. Chem. 50, 5773–5779 (2007).
Fedorova, O. et al. Small molecules that target group II introns are potent antifungal agents. Nat. Chem. Biol. 14, 1073–1078 (2018). Explores the use of small molecules through high-throughput screening, SAR and lead optimization to target a fungal self-splicing group II intron.
Anastasopoulou, P. et al. Synthesis of triazole-functionalized 2-DOS analogues and their evaluation as A-site binders. Bioorg. Med. Chem. Lett. 24, 1122–1126 (2014).
Iwatani-Yoshihara, M. et al. Discovery of allosteric inhibitors targeting the spliceosomal RNA helicase Brr2. J. Med. Chem. 60, 5759–5771 (2017).
Abulwerdi, F. A. et al. Development of small molecules with a noncanonical binding mode to HIV-1 trans activation response (TAR) RNA. J. Med. Chem. 59, 11148–11160 (2016).
Seth, P. P. et al. SAR by MS: discovery of a new class of RNA-binding small molecules for the hepatitis C virus: internal ribosome entry site IIA subdomain. J. Med. Chem. 48, 7099–7102 (2005).
Vo, D. D. et al. Oncogenic microRNAs biogenesis as a drug target: structure–activity relationship studies on new aminoglycoside conjugates. Chemistry 22, 5350–5362 (2016).
Patwardhan, N. N. et al. Amiloride as a new RNA-binding scaffold with activity against HIV-1 TAR. MedChemComm 8, 1022–1036 (2017).
Mammen, M., Choi, S.-K. & Whitesides, G. M. Polyvalent interactions in biological systems: implications for design and use of multivalent ligands and inhibitors. Angew. Chem. Int. Ed. 37, 2754–2794 (1998).
Gestwicki, J. E., Cairo, C. W., Strong, L. E., Oetjen, K. A. & Kiessling, L. L. Influencing receptor-ligand binding mechanisms with multivalent ligand architecture. J. Am. Chem. Soc. 124, 14922–14933 (2002).
Jahromi, A. H. et al. Developing bivalent ligands to target CUG triplet repeats, the causative agent of myotonic dystrophy type 1. J. Med. Chem. 56, 9471–9481 (2013).
Pushechnikov, A. et al. Rational design of ligands targeting triplet repeating transcripts that cause RNA dominant disease: application to myotonic muscular dystrophy type 1 and spinocerebellar ataxia type 3. J. Am. Chem. Soc. 131, 9767–9779 (2009).
Childs-Disney, J. L., Hoskins, J., Rzuczek, S. G., Thornton, C. A. & Disney, M. D. Rationally designed small molecules targeting the RNA that causes myotonic dystrophy type 1 are potently bioactive. ACS Chem. Biol. 7, 856–862 (2012).
Thadke, S. A. et al. Design of bivalent nucleic acid ligands for recognition of RNA-repeated expansion associated with Huntington’s disease. Biochemistry 57, 2094–2108 (2018).
Velagapudi, S. P. et al. Design of a small molecule against an oncogenic noncoding RNA. Proc. Natl Acad. Sci. USA 113, 5898–5903 (2016).
Le Grice, S. F. Targeting the HIV RNA genome: high-hanging fruit only needs a longer ladder. Curr. Top. Microbiol. Immunol. 389, 147–169 (2015).
Zapp, M. L., Stern, S. & Green, M. R. Small molecules that selectively block RNA binding of HIV-1 rev protein inhibit Rev function and viral production. Cell 74, 969–978 (1993).
Mei, H.-Y. et al. Inhibition of an HIV-1 Tat-derived peptide binding to TAR RNA by aminoglycoside antibiotics. Bioorg. Med. Chem. Lett. 5, 2755–2760 (1995).
Wang, S., Huber, P. W., Cui, M., Czarnik, A. W. & Mei, H. Y. Binding of neomycin to the TAR element of HIV-1 RNA induces dissociation of Tat protein by an allosteric mechanism. Biochemistry 37, 5549–5557 (1998).
Ratmeyer, L. et al. Inhibition of HIV-1 Rev–RRE interaction by diphenylfuran derivatives. Biochemistry 35, 13689–13696 (1996).
Park, W. K. C., Auer, M., Jaksche, H. & Wong, C.-H. Rapid combinatorial synthesis of aminoglycoside antibiotic mimetics: use of a polyethylene glycol-linked amine and a neamine-derived aldehyde in multiple component condensation as a strategy for the discovery of new inhibitors of the HIV RNA Rev responsive element. J. Am. Chem. Soc. 118, 10150–10155 (1996).
Mei, H. Y. et al. Inhibitors of protein-RNA complexation that target the RNA: specific recognition of human immunodeficiency virus type 1 TAR RNA by small organic molecules. Biochemistry 37, 14204–14212 (1998).
Hellen, C. U. & Pestova, T. V. Translation of hepatitis C virus RNA. J. Viral Hepat. 6, 79–87 (1999).
Ji, H., Fraser, C. S., Yu, Y., Leary, J. & Doudna, J. A. Coordinated assembly of human translation initiation complexes by the hepatitis C virus internal ribosome entry site RNA. Proc. Natl Acad. Sci. USA 101, 16990–16995 (2004).
Otto, G. A. & Puglisi, J. D. The pathway of HCV IRES-mediated translation initiation. Cell 119, 369–380 (2004).
Wang, W. et al. Hepatitis C viral IRES inhibition by phenazine and phenazine-like molecules. Bioorg. Med. Chem. Lett. 10, 1151–1154 (2000).
Jefferson, E. A. et al. Biaryl guanidine inhibitors of in vitro HCV-IRES activity. Bioorg. Med. Chem. Lett. 14, 5139–5143 (2004).
Dibrov, S. M. et al. Structure of a hepatitis C virus RNA domain in complex with a translation inhibitor reveals a binding mode reminiscent of riboswitches. Proc. Natl Acad. Sci. USA 109, 5223–5228 (2012).
Zafferani, M. et al. Amilorides inhibit SARS-CoV-2 replication in vitro by targeting RNA structures. Sci. Adv. 7, eabl6096 (2021). Identification of small molecules that inhibit SARS-CoV-2 viral replication by interacting with the 5′UTR of the viral mRNA.
Michel, F. & Dujon, B. Conservation of RNA secondary structures in two intron families including mitochondrial-, chloroplast- and nuclear-encoded members. EMBO J. 2, 33–38 (1983).
Fire, A. et al. Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature 391, 806–811 (1998).
Lee, R. C. & Ambros, V. An extensive class of small RNAs in Caenorhabditis elegans. Science 294, 862–864 (2001).
Bushati, N. & Cohen, S. microRNA functions. Annu. Rev. Cell Dev. Biol. 23, 175–205 (2007).
Warner, K. D., Hajdin, C. E. & Weeks, K. M. Principles for targeting RNA with drug-like small molecules. Nat. Rev. Drug Discov. 17, 547–558 (2018).
Di Giorgio, A. & Duca, M. Synthetic small-molecule RNA ligands: future prospects as therapeutic agents. MedChemComm 10, 1242–1255 (2019).
Disney, M. D. & Angelbello, A. J. Rational design of small molecules targeting oncogenic noncoding RNAs from sequence. Acc. Chem. Res. 49, 2698–2704 (2016).
Pomplun, S., Gates, Z. P., Zhang, G., Quartararo, A. J. & Pentelute, B. L. Discovery of nucleic acid binding molecules from combinatorial biohybrid nucleobase peptide libraries. J. Am. Chem. Soc. 142, 19642–19651 (2020).
Garner, A. L. et al. Tetracyclines as inhibitors of pre-microRNA maturation: a disconnection between RNA binding and inhibition. ACS Med. Chem. Lett. 10, 816–821 (2019).
Di Giorgio, A., Tran, T. P. & Duca, M. Small-molecule approaches toward the targeting of oncogenic miRNAs: roadmap for the discovery of RNA modulators. Future Med. Chem. 8, 803–816 (2016).
Staedel, C. et al. Modulation of oncogenic miRNA biogenesis using functionalized polyamines. Sci. Rep. 8, 1667 (2018).
Maucort, C. et al. Design and implementation of synthetic RNA binders for the inhibition of miR-21 biogenesis. ACS Med. Chem. Lett. 12, 899–906 (2021).
Costales, M. G. et al. A designed small molecule inhibitor of a non-coding RNA sensitizes HER2 negative cancers to herceptin. J. Am. Chem. Soc. 141, 2960–2974 (2019).
Cox, A. D., Fesik, S. W., Kimmelman, A. C., Luo, J. & Der, C. J. Drugging the undruggable RAS: mission possible? Nat. Rev. Drug Discov. 13, 828–851 (2014).
Dang, C. V., Reddy, E. P., Shokat, K. M. & Soucek, L. Drugging the ‘undruggable’ cancer targets. Nat. Rev. Cancer 17, 502–508 (2017).
Zhou, Z. D. & Tan, E. K. Iron regulatory protein (IRP)–iron responsive element (IRE) signaling pathway in human neurodegenerative diseases. Mol. Neurodegener. 12, 75 (2017).
Pantopoulos, K. Iron metabolism and the IRE/IRP regulatory system: an update. Ann. N. Y. Acad. Sci. 1012, 1–13 (2004).
Volz, K. Conservation in the iron responsive element family. Genes 12, 1365 (2021).
Muckenthaler, M. U., Galy, B. & Hentze, M. W. Systemic iron homeostasis and the iron-responsive element/iron-regulatory protein (IRE/IRP) regulatory network. Annu. Rev. Nutr. 28, 197–213 (2008).
Maio, N. et al. Mechanisms of cellular iron sensing, regulation of erythropoiesis and mitochondrial iron utilization. Semin. Hematol. 58, 161–174 (2021).
Shaw, K. T. et al. Phenserine regulates translation of beta-amyloid precursor protein mRNA by a putative interleukin-1 responsive element, a target for drug development. Proc. Natl Acad. Sci. USA 98, 7605–7610 (2001).
Rogers, J. T. & Cahill, C. M. Iron-responsive-like elements and neurodegenerative ferroptosis. Learn. Mem. 27, 395–413 (2020).
Lumsden, A. L. et al. Dysregulation of neuronal iron homeostasis as an alternative unifying effect of mutations causing familial Alzheimer’s disease. Front. Neurosci. 12, 533 (2018).
Rogers, J. T. et al. An iron-responsive element type II in the 5′-untranslated region of the Alzheimer’s amyloid precursor protein transcript. J. Biol. Chem. 277, 45518–45528 (2002).
Zhang, P. et al. Ferroptosis was more initial in cell death caused by iron overload and its underlying mechanism in Parkinson’s disease. Free Radic. Biol. Med. 152, 227–234 (2020).
Olivares, D., Huang, X., Branden, L., Greig, N. H. & Rogers, J. T. Physiological and pathological role of alpha-synuclein in Parkinson’s disease through iron mediated oxidative stress; the role of a putative iron-responsive element. Int. J. Mol. Sci. 10, 1226–1260 (2009).
Bandyopadhyay, S. et al. Novel 5′ untranslated region directed blockers of iron-regulatory protein-1 dependent amyloid precursor protein translation: implications for down syndrome and Alzheimer’s disease. PLoS ONE 8, e65978 (2013).
Utsuki, T. et al. Identification of novel small molecule inhibitors of amyloid precursor protein synthesis as a route to lower Alzheimer’s disease amyloid-beta peptide. J. Pharmacol. Exp. Ther. 318, 855–862 (2006).
Canzoneri, J. C. & Oyelere, A. K. Interaction of anthracyclines with iron responsive element mRNAs. Nucleic Acids Res. 36, 6825–6834 (2008).
Venti, A. et al. The integrated role of desferrioxamine and phenserine targeted to an iron-responsive element in the APP-mRNA 5’-untranslated region. Ann. N. Y. Acad. Sci. 1035, 34–48 (2004).
Tibodeau, J. D., Fox, P. M., Ropp, P. A., Theil, E. C. & Thorp, H. H. The up-regulation of ferritin expression using a small-molecule ligand to the native mRNA. Proc. Natl Acad. Sci. USA 103, 253–257 (2006).
Cahill, C. M., Aleyadeh, R., Gao, J., Wang, C. & Rogers, J. T. Alpha-synuclein in alcohol use disorder, connections with Parkinson’s disease and potential therapeutic role of 5′ untranslated region-directed small molecules. Biomolecules 10, 1465 (2020).
Stefanis, L. alpha-Synuclein in Parkinson’s disease. Cold Spring Harb. Perspect. Med. 2, a009399 (2012).
Rocha, E. M., De Miranda, B. & Sanders, L. H. Alpha-synuclein: pathology, mitochondrial dysfunction and neuroinflammation in Parkinson’s disease. Neurobiol. Dis. 109, 249–257 (2018).
Junn, E. et al. Repression of alpha-synuclein expression and toxicity by microRNA-7. Proc. Natl Acad. Sci. USA 106, 13052–13057 (2009).
Ule, J. & Blencowe, B. J. Alternative splicing regulatory networks: functions, mechanisms, and evolution. Mol. Cell. 76, 329–345 (2019).
Lee, Y. & Rio, D. C. Mechanisms and regulation of alternative pre-mRNA splicing. Annu. Rev. Biochem. 84, 291–323 (2015).
Bauman, J., Jearawiriyapaisarn, N. & Kole, R. Therapeutic potential of splice-switching oligonucleotides. Oligonucleotides 19, 1–13 (2009).
Sazani, P. & Kole, R. Therapeutic potential of antisense oligonucleotides as modulators of alternative splicing. J. Clin. Invest. 112, 481–486 (2003).
Havens, M. A. & Hastings, M. L. Splice-switching antisense oligonucleotides as therapeutic drugs. Nucleic Acids Res. 44, 6549–6563 (2016).
Scharner, J. et al. Hybridization-mediated off-target effects of splice-switching antisense oligonucleotides. Nucleic Acids Res. 48, 802–816 (2020).
Arendt, T., Stieler, J. T. & Holzer, M. Tau and tauopathies. Brain Res. Bull. 126, 238–292 (2016).
Donahue, C. P., Muratore, C., Wu, J. Y., Kosik, K. S. & Wolfe, M. S. Stabilization of the tau exon 10 stem loop alters pre-mRNA splicing. J. Biol. Chem. 281, 23302–23306 (2006).
Zheng, S., Chen, Y., Donahue, C. P., Wolfe, M. S. & Varani, G. Structural basis for stabilization of the tau pre-mRNA splicing regulatory element by novantrone (mitoxantrone). Chem. Biol. 16, 557–566 (2009).
Lisowiec, J., Magner, D., Kierzek, E., Lenartowicz, E. & Kierzek, R. Structural determinants for alternative splicing regulation of the MAPT pre-mRNA. RNA Biol. 12, 330–342 (2015).
Liu, Y. et al. Mitoxantrone analogues as ligands for a stem–loop structure of tau pre-mRNA. J. Med. Chem. 52, 6523–6526 (2009).
Luo, Y. & Disney, M. D. Bottom-up design of small molecules that stimulate exon 10 skipping in mutant MAPT pre-mRNA. Chembiochem 15, 2041–2044 (2014).
Sakamoto, K. M. et al. Protacs: chimeric molecules that target proteins to the Skp1-Cullin-F box complex for ubiquitination and degradation. Proc. Natl Acad. Sci. USA 98, 8554–8559 (2001).
Zou, Y., Ma, D. & Wang, Y. The PROTAC technology in drug development. Cell Biochem. Funct. 37, 21–30 (2019).
Sakamoto, K. M. Chimeric molecules to target proteins for ubiquitination and degradation. Methods Enzymol. 399, 833–847 (2005).
Ottis, P. & Crews, C. M. Proteolysis-targeting chimeras: induced protein degradation as a therapeutic strategy. ACS Chem. Biol. 12, 892–898 (2017).
Crews, C. M. Inducing protein degradation as a therapeutic strategy. J. Med. Chem. 61, 403–404 (2018).
Chamberlain, P. P. & Hamann, L. G. Development of targeted protein degradation therapeutics. Nat. Chem. Biol. 15, 937–944 (2019).
Costales, M. G., Suresh, B., Vishnu, K. & Disney, M. D. Targeted degradation of a hypoxia-associated non-coding RNA enhances the selectivity of a small molecule interacting with RNA. Cell Chem. Biol. 26, 1180–1186 (2019).
Thakur, C. S. et al. Small-molecule activators of RNase L with broad-spectrum antiviral activity. Proc. Natl Acad. Sci. USA 104, 9585–9590 (2007).
Janes, J. et al. The ReFRAME library as a comprehensive drug repurposing library and its application to the treatment of cryptosporidiosis. Proc. Natl Acad. Sci. USA 115, 10750–10755 (2018).
Grundy, F. J. & Henkin, T. M. The S box regulon: a new global transcription termination control system for methionine and cysteine biosynthesis genes in Gram-positive bacteria. Mol. Microbiol. 30, 737–749 (1998).
Gelfand, M. A conserved RNA structure element involved in the regulation of bacterial riboflavin synthesis genes. Trends Genet. 15, 439–442 (1999).
Mironov, A. S. et al. Sensing small molecules by nascent RNA. Cell 111, 747–756 (2002).
Nahvi, A. et al. Genetic control by a metabolite binding mRNA. Chem. Biol. 9, 1043–1049 (2002).
Winkler, W., Nahvi, A. & Breaker, R. R. Thiamine derivatives bind messenger RNAs directly to regulate bacterial gene expression. Nature 419, 952–956 (2002).
Winkler, W. C. & Breaker, R. R. Regulation of bacterial gene expression by riboswitches. Annu. Rev. Microbiol. 59, 487–517 (2005).
Polaski, J. T., Kletzien, O. A., Drogalis, L. K. & Batey, R. T. A functional genetic screen reveals sequence preferences within a key tertiary interaction in cobalamin riboswitches required for ligand selectivity. Nucleic Acids Res. 46, 9094–9105 (2018).
Batey, R. T., Gilbert, S. D. & Montange, R. K. Structure of a natural guanine-responsive riboswitch complexed with the metabolite hypoxanthine. Nature 432, 411–415 (2004).
Batey, R. T., Rambo, R. P. & Doudna, J. A. Tertiary motifs in RNA structure and folding. Angew. Chem. Int. Ed. Engl. 38, 2326–2343 (1999).
Breaker, R. R. Riboswitches and the RNA world. Cold Spring Harb. Perspect. Biol. 4, a003566 (2012).
Garst, A. D., Edwards, A. L. & Batey, R. T. Riboswitches: structures and mechanisms. Cold Spring Harb. Perspect. Biol. 3, a003533 (2011).
Sherlock, M. E. & Breaker, R. R. Former orphan riboswitches reveal unexplored areas of bacterial metabolism, signaling, and gene control processes. RNA 26, 675–693 (2020).
Wang, J. X. & Breaker, R. R. Riboswitches that sense S-adenosylmethionine and S-adenosylhomocysteine. Biochem. Cell. Biol. 86, 157–168 (2008).
Balibar, C. J. et al. Validation and development of an Escherichia coli riboflavin pathway phenotypic screen hit as a small-molecule ligand of the flavin mononucleotide riboswitch. Methods Mol. Biol. 1787, 19–40 (2018).
Serganov, A., Huang, L. & Patel, D. J. Coenzyme recognition and gene regulation by a flavin mononucleotide riboswitch. Nature 458, 233–237 (2009).
Vicens, Q., Mondragón, E. & Batey, R. T. Molecular sensing by the aptamer domain of the FMN riboswitch: a general model for ligand binding by conformational selection. Nucleic Acids Res. 39, 8586–8598 (2011). A structural analysis of FMN binding to the FMN riboswitch using SHAPE mapping and X-ray crystallography reveals detailed conformational changes upon ligand binding.
Blount, K. F. et al. Novel riboswitch-binding flavin analog that protects mice against Clostridium difficile infection without inhibiting cecal flora. Antimicrob. Agents Chemother. 59, 5736–5746 (2015).
Vicens, Q. et al. Structure–activity relationship of flavin analogues that target the flavin mononucleotide riboswitch. ACS Chem. Biol. 13, 2908–2919 (2018).
Cho, S. & Dreyfuss, G. A degron created by SMN2 exon 7 skipping is a principal contributor to spinal muscular atrophy severity. Genes Dev. 24, 438–442 (2010).
Ratni, H. et al. Specific correction of alternative survival motor neuron 2 splicing by small molecules: discovery of a potential novel medicine to treat spinal muscular atrophy. J. Med. Chem. 59, 6086–6100 (2016).
Campagne, S. et al. Structural basis of a small molecule targeting RNA for a specific splicing correction. Nat. Chem. Biol. 15, 1191–1198 (2019).
Sivaramakrishnan, M. et al. Binding to SMN2 pre-mRNA–protein complex elicits specificity for small molecule splicing modifiers. Nat. Commun. 8, 1476 (2017).
Wang, J., Schultz, P. G. & Johnson, K. A. Mechanistic studies of a small-molecule modulator of SMN2 splicing. Proc. Natl Acad. Sci. USA 115, E4604–E4612 (2018).
Keller, C. G. et al. An orally available, brain penetrant, small molecule lowers huntingtin levels by enhancing pseudoexon inclusion. Nat. Commun. 13, 1150 (2022).
Chaidir, C., Edrada-Ebel, R., Ebel, R., Bohnenstengel, F. & Nugroho, B. W. Chemistry and biological activity of rocaglamide derivatives and related compounds in Aglaia species (Meliaceae). Curr. Org. Chem. 5, 923–938 (2001).
Kim, S. et al. Silvestrol, a potential anticancer rocaglate derivative from Aglaia foveolata, induces apoptosis in LNCaP cells through the mitochondrial/apoptosome pathway without activation of executioner caspase-3 or -7. Anticancer Res. 27, 2175–2183 (2007).
Schulz, G., Victoria, C., Kirschning, A. & Steinmann, E. Rocaglamide and silvestrol: a long story from anti-tumor to anti-coronavirus compounds. Nat. Prod. Rep. 38, 18–23 (2021).
Iwasaki, S., Floor, S. N. & Ingolia, N. T. Rocaglates convert DEAD-box protein eIF4A into a sequence-selective translational repressor. Nature 534, 558–561 (2016). A mechanistic study showing that the anti-tumour drug rocaglamide A functions by clamping elF4A to polypurine sequences and blocking ribosomal scanning in the 5′UTR.
Ernst, J. T. et al. Design of development candidate eFT226, a first in class inhibitor of eukaryotic initiation factor 4A RNA helicase. J. Med. Chem. 63, 5879–5955 (2020).
Nilewski, C. et al. 1-Aminomethyl SAR in a novel series of flavagline-inspired eIF4A inhibitors: effects of amine substitution on cell potency and in vitro PK properties. Bioorg. Med. Chem. Lett. 47, 128111 (2021).
Nilewski, C. et al. Strategic diastereoselective C1 functionalization in the aza-rocaglamide scaffold toward natural product-inspired eIF4A inhibitors. Org. Lett. 22, 6257–6261 (2020).
Kruger, K. et al. Self-splicing RNA: autoexcision and autocyclization of the ribosomal RNA intervening sequence of Tetrahymena. Cell 31, 147–157 (1982).
Guerrier-Takada, C., Gardiner, K., Marsh, T., Pace, N. & Altman, S. The RNA moiety of ribonuclease P is the catalytic subunit of the enzyme. Cell 35, 849–857 (1983).
Cech, T. R. & Bass, B. L. Biological catalysis by RNA. Annu. Rev. Biochem. 55, 599–629 (1986).
Peattie, D. A., Douthwaite, S., Garrett, R. A. & Noller, H. F. A “bulged” double helix in a RNA–protein contact site. Proc. Natl Acad. Sci. USA 78, 7331–7335 (1981).
Moazed, D. & Noller, H. F. Interaction of antibiotics with functional sites in 16S ribosomal RNA. Nature 327, 389–394 (1987).
Velagapudi, S. P., Li, Y. & Disney, M. D. A cross-linking approach to map small molecule–RNA binding sites in cells. Bioorg. Med. Chem. Lett. 29, 1532–1536 (2019).
Lipinski, C. A., Lombardo, F., Dominy, B. W. & Feeney, P. J. Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv. Drug Deliv. Rev. 46, 3–26 (2001).
Davila-Calderon, J. et al. IRES-targeting small molecule inhibits enterovirus 71 replication via allosteric stabilization of a ternary complex. Nat. Commun. 11, 4775 (2020).
Liu, T. & Pyle, A. M. Discovery of highly reactive self-splicing group II introns within the mitochondrial genomes of human pathogenic fungi. Nucleic Acids Res. 49, 12422–12432 (2021).
Angelbello, A. J. et al. Precise small- molecule cleavage of an r(CUG) repeat expansion in a myotonic dystrophy mouse model. Proc. Natl Acad. Sci. USA 116, 7799–7804 (2019).
Donahue, C. P., Ni, J., Rozners, E., Glicksman, M. A. & Wolfe, M. S. Identification of tau stem loop RNA stabilizers. J. Biomol. Screen. 12, 789–799 (2007).
Angelbello, A. J et al. A small molecule that binds an RNA repeat expansion stimulates its decay via the exosome complex. Cell Chem. Biol. 28, 34–45 (2021). Demonstrates that RNA-targeting small molecules can interface the target with natural decay pathways, in particular the exosome.
Murchie, A. I. et al. Structure-based drug design targeting an inactive RNA conformation: exploiting the flexibility of HIV-1 TAR RNA. J. Mol. Biol. 336, 625–638 (2004).
Foloppe, N. et al. A structure-based strategy to identify new molecular scaffolds targeting the bacterial ribosomal A-site. Bioorg. Med. Chem. 12, 935–947 (2004).
Morgan, B. S., Forte, J. E., Culver, R. N., Zhang, Y. & Hargrove, A. E. Discovery of key physicochemical, structural, and spatial properties of RNA-targeted bioactive ligands. Angew. Chem. Int. Ed. Engl. 56, 13498–13502 (2017).
Childs-Disney, J. L. et al. A massively parallel selection of small molecule–RNA motif binding partners informs design of an antiviral from sequence. Chem 4, 2384–2404 (2018).
Bickerton, G. R., Paolini, G. V., Besnard, J., Muresan, S. & Hopkins, A. L. Quantifying the chemical beauty of drugs. Nat. Chem. 4, 90–98 (2012).
Bhullar, K. S. et al. Kinase-targeted cancer therapies: progress, challenges and future directions. Mol. Cancer 17, 48 (2018).
Paulson, H. Repeat expansion diseases. Handb. Clin. Neurol. 147, 105–123 (2018).
Bernat, V. & Disney, M. D. RNA structures as mediators of neurological diseases and as drug targets. Neuron 87, 28–46 (2015).
de Mezer, M., Wojciechowska, M., Napierala, M., Sobczak, K. & Krzyzosiak, W. J. Mutant CAG repeats of huntingtin transcript fold into hairpins, form nuclear foci and are targets for RNA interference. Nucleic Acids Res. 39, 3852–3863 (2011).
Sobczak, K., de Mezer, M., Michlewski, G., Krol, J. & Krzyzosiak, W. J. RNA structure of trinucleotide repeats associated with human neurological diseases. Nucleic Acids Res. 31, 5469–5482 (2003).
Tian, B. et al. Expanded CUG repeat RNAs form hairpins that activate the double-stranded RNA-dependent protein kinase PKR. RNA 6, 79–87 (2000).
Jog, S. P. et al. RNA splicing is responsive to MBNL1 dose. PLoS ONE 7, e48825 (2012).
Rzuczek, S. G. et al. Precise small-molecule recognition of a toxic CUG RNA repeat expansion. Nat. Chem. Biol. 13, 188–193 (2017).
Wagner-Griffin, S. et al. A druglike small molecule that targets r(CCUG) repeats in myotonic dystrophy type 2 facilitates degradation by RNA quality control pathways. J. Med. Chem. 64, 8474–8485 (2021).
Bush, J. A. et al. Ribonuclease recruitment using a small molecule reduced c9ALS/FTD r(G4C2) repeat expansion in vitro and in vivo ALS models. Sci. Transl. Med. 13, eabd5991 (2021).
Angelbello, A. J. & Disney, M. D. A toxic RNA templates the synthesis of its own fluorogenic inhibitor by using a bio-orthogonal tetrazine ligation in cells and tissues. ACS Chem. Biol. 15, 1820–1825 (2020).
Burma, S., Chen, B. P., Murphy, M., Kurimasa, A. & Chen, D. J. ATM phosphorylates histone H2AX in response to DNA double-strand breaks. J. Biol. Chem. 276, 42462–42467 (2001).
Li, Y. & Disney, M. D. Precise small molecule degradation of a noncoding RNA identifies cellular binding sites and modulates an oncogenic phenotype. ACS Chem. Biol. 13, 3065–3071 (2018).
Liu, X. et al. Targeted degradation of the oncogenic microRNA 17-92 cluster by structure-targeting ligands. J. Am. Chem. Soc. 142, 6970–6982 (2020).
Benhamou, R. I. et al. Structure-specific cleavage of an RNA repeat expansion with a dimeric small molecule is advantageous over sequence-specific recognition by an oligonucleotide. ACS Chem. Biol. 15, 485–493 (2020).
Acknowledgements
This work was funded by the NIH (R35 NS116846, R01 CA249180, UG3 NS116921 and P01 NS099114 to M.D.D. and R01 GM073850 to R.T.B.) and the Department of Defense (W81XWH-19-1-0719 and W81XWH-20-1-0727 to M.D.D).
Author information
Authors and Affiliations
Contributions
All authors contributed to the writing of the manuscript and preparation of figures.
Corresponding authors
Ethics declarations
Competing interests
M.D.D. is a founder of Expansion Therapeutics. R.T.B. serves on the Scientific Advisory Board of Expansion Therapeutics and MeiraGTx. The remaining authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Glossary
- Antisense oligonucleotides
-
(ASOs). Single-stranded DNA or RNA oligonucleotides, including combinations thereof that are chemically modified, that are complementary to the sequence of target RNA and can induce RNase H-mediated degradation or sterically block the ribosome.
- Riboswitch
-
Region of RNA structure located in the 5′ leader of bacterial RNAs that undergoes conformational switching upon small-molecule binding to regulate translation.
- Internal ribosome entry site
-
(IRES). RNA structural element typically in the 5′ untranslated region that can recruit ribosomes and initiate cap-independent translation.
- MicroRNA
-
(miRNA). Short noncoding RNA that has important roles in mediating gene expression by guiding Argonaute proteins to their mRNA target via base pairing to the 3′ untranslated region. miRNAs are formed stepwise, first from primary miRNAs to precursor miRNAs by the nuclease Drosha; then from precursor to mature (functional) miRNAs by the nuclease Dicer.
- Pharmacophore
-
3D arrangement of a molecule or molecular group that confers bioactivity via interactions with the compound’s target.
- Transcriptome
-
A landscape of all RNAs transcribed from a genome.
- Splicing modulators
-
Small molecules or proteins that are capable of directing RNA splicing by inducing inclusion or exclusion of exons.
- Functional selectivity
-
A metric that compares the effect of a compound on the biological activity of the desired target versus its effect on other targets.
- On- and off-targets
-
An on-target is the biomolecule that a compound is designed to modulate the function of; an off-target is unintendedly modulated by the small molecule.
- RNA sequencing
-
(RNA-seq). A high-throughput method that analyses gene expression transcriptome-wide.
- Phenotypic screening
-
A method that screens small molecules on the basis of a desired phenotypic outcome, without knowledge of its actual target.
- Structure–activity relationship
-
(SAR). Approach that aims to correlate the chemical structure of a compound with its biological activity.
- Ribonuclease-targeting chimeras
-
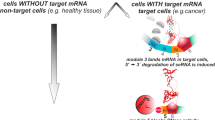
(RIBOTACs). Small-molecule chimeras that activate and recruit endogenous ribonucleases to an RNA target to trigger its degradation.
- Gini coefficient
-
A statistical measurement that quantifies the selectivity of a small molecule, ranging from 0 to 1, where 0 indicates that the compound inhibits every drug target studied equally and 1 indicates a compound selective for one drug target.
- Intrinsically disordered proteins
-
(IDPs). Proteins that lack well-defined 3D structures and are thus considered undruggable.
- Iron-responsive elements
-
(IREs). Small stem–loop structures present in 5′ or 3′ untranslated regions (UTRs) that bind to iron regulatory proteins (IRPs). IRP binding to IREs in the 5′UTR prevents ribosome docking and blocks RNA translation whereas binding to the 3′UTR stabilizes the transcript and upregulates translation.
- Proteolysis-targeting chimeras
-
(PROTACs). Chimeric molecules comprising a protein-binding module and a E3 ubiquitin ligase-recognition module, which tags the targeted protein for selective degradation by the proteasome.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Childs-Disney, J.L., Yang, X., Gibaut, Q.M.R. et al. Targeting RNA structures with small molecules. Nat Rev Drug Discov 21, 736–762 (2022). https://doi.org/10.1038/s41573-022-00521-4
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41573-022-00521-4
This article is cited by
-
Predicting small molecule–RNA interactions without RNA tertiary structures
Nature Biotechnology (2026)
-
Deciphering the regulatory landscape of enhancer RNAs in health and disease
Signal Transduction and Targeted Therapy (2026)
-
Chemical and ribosomal synthesis of atropisomeric and macrocyclic peptides with embedded quinolines
Nature Chemistry (2026)
-
Bridging technical innovation and computational advances in studies of RNA–protein assemblies
Nature Reviews Genetics (2026)
-
Oral splicing modulator branaplam in Huntington’s disease: a phase 2 randomized controlled trial
Nature Medicine (2026)