Abstract
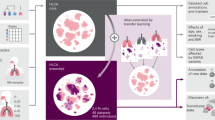
The goal of the Human Liver Cell Atlas (HLiCA) is to create a comprehensive map that defines the normal functions of diverse liver cell types and their spatial relationships over the human lifespan. This project fits within the goals of the Human Cell Atlas to create comprehensive reference maps of all human cells as a basis for both understanding human health and diagnosing, monitoring and treating disease. Through collection of samples from diverse individuals, data integration across technologies and overcoming liver-specific challenges for experimental methods, the HLiCA will map as many cell types and states as possible in healthy human livers from individuals across all ages and many ancestries. Establishing this HLiCA of healthy livers is a critical step to begin to understand perturbations in disease. The HLiCA will be available on an open-access platform to facilitate data sharing and dissemination. We expect that creation of the HLiCA will help to lay the foundation for new research initiatives to advance our understanding of liver disease, improve methods of tissue engineering, and identify novel prognostic biomarkers and therapies to improve patient outcomes. We describe key experimental and computational challenges to overcome in building the atlas and the potential impact of the atlas on disease research.
Key points
-
Building the Human Liver Cell Atlas requires collaborative effort within the liver single-cell genomics community.
-
Characterization of the normal human liver must account for sample-to-sample variability due to age, gender, ancestry, lifestyle, microbiome, environmental factors and experimental approaches, among other factors.
-
Collecting standardized metadata and optimizing data integration is critical to generate a useful and comprehensive cell atlas across multiple laboratories and institutions.
-
The Human Liver Cell Atlas will provide the foundation for understanding disease-specific perturbations and hopefully identify cell-type-specific therapeutic strategies to reduce the global burden of liver disease.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
23 October 2025
A Correction to this paper has been published: https://doi.org/10.1038/s41575-025-01145-w
References
Moon, A. M., Singal, A. G. & Tapper, E. B. Contemporary epidemiology of chronic liver disease and cirrhosis. Clin. Gastroenterol. Hepatol. 18, 2650–2666 (2020).
GBD 2017 Cirrhosis Collaborators The global, regional, and national burden of cirrhosis by cause in 195 countries and territories, 1990-2017: a systematic analysis for the global burden of disease study 2017. Lancet Gastroenterol. Hepatol. 5, 245–266 (2020).
Cheemerla, S. & Balakrishnan, M. Global epidemiology of chronic liver disease. Clin. Liver Dis. 17, 365–370 (2021).
Riazi, K. et al. The prevalence and incidence of NAFLD worldwide: a systematic review and meta-analysis. Lancet Gastroenterol. Hepatol. 7, 851–861 (2022).
Huang, D. Q., Mathurin, P., Cortez-Pinto, H. & Loomba, R. Global epidemiology of alcohol-associated cirrhosis and HCC: trends, projections and risk factors. Nat. Rev. Gastroenterol. Hepatol. 20, 37–49 (2023).
Wong, R. J. & Singal, A. K. Trends in liver disease etiology among adults awaiting liver transplantation in the United States, 2014-2019. JAMA Netw. Open 3, e1920294 (2020).
World Health Organization. Global Progress Report on HIV, Viral Hepatitis and Sexually Transmitted Infections, 2021 (WHO, 2021).
Haga, J. et al. Liver regeneration in donors and adult recipients after living donor liver transplantation. Liver Transpl. 14, 1718–1724 (2008).
Read, A. E. Clinical physiology of the liver. Br. J. Anaesth. 44, 910–917 (1972).
Trefts, E., Gannon, M. & Wasserman, D. H. The liver. Curr. Biol. 27, R1147–R1151 (2017).
Kiernan, F. XXIX. The anatomy and physiology of the liver. Phil. Trans. R. Soc. 123, 711–770 (1833).
Arias, I. M. et al. The Liver: Biology and Pathobiology (Wiley, 2020).
Jungermann, K. & Kietzmann, T. Zonation of parenchymal and nonparenchymal metabolism in liver. Annu. Rev. Nutr. 16, 179–203 (1996).
Paris, J. & Henderson, N. C. Liver zonation, revisited. Hepatology 76, 1219–1230 (2022).
MacParland, S. A. et al. Single cell RNA sequencing of human liver reveals distinct intrahepatic macrophage populations. Nat. Commun. 9, 4383 (2018).
Ramachandran, P. et al. Resolving the fibrotic niche of human liver cirrhosis at single-cell level. Nature 575, 512–518 (2019).
Aizarani, N. et al. A human liver cell atlas reveals heterogeneity and epithelial progenitors. Nature 572, 199–204 (2019).
Sharma, A. et al. Onco-fetal reprogramming of endothelial cells drives immunosuppressive macrophages in hepatocellular carcinoma. Cell 183, 377–394.e21 (2020).
Andrews, T. S. et al. Single-cell, single-nucleus, and spatial RNA sequencing of the human liver identifies cholangiocyte and mesenchymal heterogeneity. Hepatol. Commun. 6, 821–840 (2022).
Parlakgul, G. et al. Regulation of liver subcellular architecture controls metabolic homeostasis. Nature 603, 736–742 (2022).
Walsh, C. L. et al. Imaging intact human organs with local resolution of cellular structures using hierarchical phase-contrast tomography. Nat. Methods 18, 1532–1541 (2021).
Consortium, M. I. Functional connectomics spanning multiple areas of mouse visual cortex. Nature 640, 435–447 (2025).
Watson, B. R. et al. Spatial transcriptomics of healthy and fibrotic human liver at single-cell resolution. Nat. Commun. 16, 319 (2025).
Waxman, D. J. & O’Connor, C. Growth hormone regulation of sex-dependent liver gene expression. Mol. Endocrinol. 20, 2613–2629 (2006).
Jaffe, C. A. et al. Regulatory mechanisms of growth hormone secretion are sexually dimorphic. J. Clin. Invest. 102, 153–164 (1998).
Kaplan, M. M. & Gershwin, M. E. Primary biliary cirrhosis. N. Engl. J. Med. 353, 1261–1273 (2005).
Lin, L. et al. The burden and trends of primary liver cancer caused by specific etiologies from 1990 to 2017 at the global, regional, national, age, and sex level results from the Global Burden of Disease Study 2017. Liver Cancer 9, 563–582 (2020).
Targher, G., Byrne, C. D. & Tilg, H. MASLD: a systemic metabolic disorder with cardiovascular and malignant complications. Gut 73, 691–702 (2024).
Niriella, M. A. et al. Prevalence and associated factors for non-alcoholic fatty liver disease among adults in the south Asian region: a meta-analysis. Lancet Reg. Health Southeast Asia 15, 100220 (2023).
Vendhan, R. et al. Comparison of characteristics between nonobese and overweight/obese subjects with nonalcoholic fatty liver disease in a South Indian population. Diabetes Technol. Ther. 16, 48–55 (2014).
Peiseler, M. et al. Immune mechanisms linking metabolic injury to inflammation and fibrosis in fatty liver disease — novel insights into cellular communication circuits. J. Hepatol. 77, 1136–1160 (2022).
Lombardi, R., Iuculano, F., Pallini, G., Fargion, S. & Fracanzani, A. L. Nutrients, genetic factors, and their interaction in non-alcoholic fatty liver disease and cardiovascular disease. Int. J. Mol. Sci. 21, 8761 (2020).
Lakshminarayanan, B. & Davenport, M. Biliary atresia: a comprehensive review. J. Autoimmun. 73, 1–9 (2016).
Mann, D. A. Epigenetics in liver disease. Hepatology 60, 1418–1425 (2014).
Bolshette, N., Ibrahim, H., Reinke, H. & Asher, G. Circadian regulation of liver function: from molecular mechanisms to disease pathophysiology. Nat. Rev. Gastroenterol. Hepatol. 20, 695–707 (2023).
Duane, W. C., Levitt, D. G., Mueller, S. M. & Behrens, J. C. Regulation of bile acid synthesis in man. Presence of a diurnal rhythm. J. Clin. Invest. 72, 1930–1936 (1983).
March, S. et al. Autonomous circadian rhythms in the human hepatocyte regulate hepatic drug metabolism and inflammatory responses. Sci. Adv. 10, eadm9281 (2024).
Narmada, B. C. et al. Single cell landscape of functionally cured chronic hepatitis B patients reveals activation of innate and altered CD4-CTL-driven adaptive immunity. J. Hepatol. 81, 42–61 (2024).
Lotto, J., Stephan, T. L. & Hoodless, P. A. Fetal liver development and implications for liver disease pathogenesis. Nat. Rev. Gastroenterol. Hepatol. 20, 561–581 (2023).
Wesley, B. T. et al. Single-cell atlas of human liver development reveals pathways directing hepatic cell fates. Nat. Cell Biol. 24, 1487–1498 (2022).
Santos Silva, E. et al. Bile acids profile and redox status in healthy infants. Pediatr. Res. 93, 1856–1864 (2023).
Grijalva, J. & Vakili, K. Neonatal liver physiology. Semin. Pediatr. Surg. 22, 185–189 (2013).
Itoh, S. et al. Fetal and neonatal bilirubin metabolism. Front. Pediatr. 10, 1002408 (2022).
Edgar, R. D. et al. Single-cell atlas of human pediatric liver reveals age-related hepatic gene signatures. Hepatol. Commun. 9, e0813 (2025).
Nobili, V., Alisi, A., Newton, K. P. & Schwimmer, J. B. Comparison of the phenotype and approach to pediatric vs adult patients with nonalcoholic fatty liver disease. Gastroenterology 150, 1798–1810 (2016).
Ben-Moshe, S. & Itzkovitz, S. Spatial heterogeneity in the mammalian liver. Nat. Rev. Gastroenterol. Hepatol. 16, 395–410 (2019).
Nobili, V. et al. A protective effect of breastfeeding on the progression of non-alcoholic fatty liver disease. Arch. Dis. Child. 94, 801–805 (2009).
Park, Y. et al. The association between breastfeeding and nonalcoholic fatty liver disease in parous women: a nation-wide cohort study. Hepatology 74, 2988–2997 (2021).
Eom, J. A. et al. Diet-regulating microbiota and host immune system in liver disease. Int. J. Mol. Sci. 22, 6326 (2021).
Yang, X. et al. The gut–liver axis in immune remodeling: new insight into liver diseases. Int. J. Biol. Sci. 16, 2357–2366 (2020).
Schreiber, R. A., Harpavat, S., Hulscher, J. B. F. & Wildhaber, B. E. Biliary atresia in 2021: epidemiology, screening and public policy. J. Clin. Med. 11, 999 (2022).
Wang, J. et al. Liver immune profiling reveals pathogenesis and therapeutics for biliary atresia. Cell 183, 1867–1883.e26 (2020).
Taylor, S. A. et al. Transcriptional profiling of pediatric cholestatic livers identifies three distinct macrophage populations. PLoS ONE 16, e0244743 (2021).
Gebhardt, R. Metabolic zonation of the liver: regulation and implications for liver function. Pharmacol. Ther. 53, 275–354 (1992).
Rappaport, A. M., Borowy, Z. J., Lougheed, W. M. & Lotto, W. N. Subdivision of hexagonal liver lobules into a structural and functional unit; role in hepatic physiology and pathology. Anat. Rec. 119, 11–33 (1954).
Halpern, K. B. et al. Single-cell spatial reconstruction reveals global division of labour in the mammalian liver. Nature 542, 352–356 (2017).
Manco, R. & Itzkovitz, S. Liver zonation. J. Hepatol. 74, 466–468 (2021).
Cheng, X. et al. Glucagon contributes to liver zonation. Proc. Natl Acad. Sci. USA 115, E4111–E4119 (2018).
Halpern, K. B. et al. Paired-cell sequencing enables spatial gene expression mapping of liver endothelial cells. Nat. Biotechnol. 36, 962–970 (2018).
Dobie, R. et al. Single-cell transcriptomics uncovers zonation of function in the mesenchyme during liver fibrosis. Cell Rep. 29, 1832–1847.e8 (2019).
Hu, S. et al. Single-cell spatial transcriptomics reveals a dynamic control of metabolic zonation and liver regeneration by endothelial cell Wnt2 and Wnt9b. Cell Rep. Med. 3, 100754 (2022).
Zhao, L. et al. Tissue repair in the mouse liver following acute carbon tetrachloride depends on injury-induced Wnt/β-catenin signaling. Hepatology 69, 2623–2635 (2019).
Planas-Paz, L. et al. The RSPO-LGR4/5-ZNRF3/RNF43 module controls liver zonation and size. Nat. Cell Biol. 18, 467–479 (2016).
Gola, A. et al. Commensal-driven immune zonation of the liver promotes host defence. Nature 589, 131–136 (2021).
Sakai, M. et al. Liver-derived signals sequentially reprogram myeloid enhancers to initiate and maintain Kupffer cell identity. Immunity 51, 655–670.e8 (2019).
Bonnardel, J. et al. Stellate cells, hepatocytes, and endothelial cells imprint the Kupffer cell identity on monocytes colonizing the liver macrophage niche. Immunity 51, 638–654.e9 (2019).
Guilliams, M. et al. Spatial proteogenomics reveals distinct and evolutionarily conserved hepatic macrophage niches. Cell 185, 379–396.e38 (2022).
Remmerie, A. et al. Osteopontin expression identifies a subset of recruited macrophages distinct from Kupffer cells in the fatty liver. Immunity 53, 641–657.e14 (2020).
Seidman, J. S. et al. Niche-specific reprogramming of epigenetic landscapes drives myeloid cell diversity in nonalcoholic steatohepatitis. Immunity 52, 1057–1074.e7 (2020).
Balmer, M. L. et al. The liver may act as a firewall mediating mutualism between the host and its gut commensal microbiota. Sci. Transl. Med. 6, 237ra266 (2014).
Brazovskaja, A. et al. Cell atlas of the regenerating human liver after portal vein embolization. Nat. Commun. 15, 5827 (2024).
Matchett, K. P. et al. Multimodal decoding of human liver regeneration. Nature 630, 158–165 (2024).
Gribben, C. et al. Acquisition of epithelial plasticity in human chronic liver disease. Nature 630, 166–173 (2024).
Chung, B. K., Ogaard, J., Reims, H. M., Karlsen, T. H. & Melum, E. Spatial transcriptomics identifies enriched gene expression and cell types in human liver fibrosis. Hepatol. Commun. 6, 2538–2550 (2022).
Buonomo, E. L. et al. Liver stromal cells restrict macrophage maturation and stromal IL-6 limits the differentiation of cirrhosis-linked macrophages. J. Hepatol. 76, 1127–1137 (2022).
Filliol, A. et al. Opposing roles of hepatic stellate cell subpopulations in hepatocarcinogenesis. Nature 610, 356–365 (2022).
Kajiyama, K., Maeda, T., Takenaka, K., Sugimachi, K. & Tsuneyoshi, M. The significance of stromal desmoplasia in intrahepatic cholangiocarcinoma: a special reference of ‘scirrhous-type’ and ‘nonscirrhous-type’ growth. Am. J. Surg. Pathol. 23, 892–902 (1999).
Sirica, A. E. The role of cancer-associated myofibroblasts in intrahepatic cholangiocarcinoma. Nat. Rev. Gastroenterol. Hepatol. 9, 44–54 (2011).
Affo, S. et al. Promotion of cholangiocarcinoma growth by diverse cancer-associated fibroblast subpopulations. Cancer Cell 39, 866–882.e11 (2021).
Jagadeesh, K. A. et al. Identifying disease-critical cell types and cellular processes by integrating single-cell RNA-sequencing and human genetics. Nat. Genet. 54, 1479–1492 (2022).
Zhang, Q. et al. Landscape and dynamics of single immune cells in hepatocellular carcinoma. Cell 179, 829–845.e20 (2019).
Korsunsky, I. et al. Cross-tissue, single-cell stromal atlas identifies shared pathological fibroblast phenotypes in four chronic inflammatory diseases. Med 3, 481–518.e14 (2022).
Muhl, L. et al. Single-cell analysis uncovers fibroblast heterogeneity and criteria for fibroblast and mural cell identification and discrimination. Nat. Commun. 11, 3953 (2020).
Genshaft, A. S. et al. Single-cell RNA sequencing of liver fine-needle aspirates captures immune diversity in the blood and liver in chronic hepatitis B patients. Hepatology 78, 1525–1541 (2023).
Habib, N. et al. Massively parallel single-nucleus RNA-seq with DroNc-seq. Nat. Methods 14, 955–958 (2017).
Stoeckius, M. et al. Simultaneous epitope and transcriptome measurement in single cells. Nat. Methods 14, 865–868 (2017).
Atif, J., Thoeni, C., Bader, G. D., McGilvray, I. D. & MacParland, S. A. Unraveling the complexity of liver disease one cell at a time. Semin. Liver Dis. 42, 250–270 (2022).
Cheng, M. et al. Spatially resolved transcriptomics: a comprehensive review of their technological advances, applications, and challenges. J. Genet. Genom. 50, 625–640 (2023).
Luecken, M. D. et al. Benchmarking atlas-level data integration in single-cell genomics. Nat. Methods 19, 41–50 (2022).
Young, M. D. & Behjati, S. SoupX removes ambient RNA contamination from droplet-based single-cell RNA sequencing data. Gigascience 9, giaa151 (2020).
Lahnemann, D. et al. Eleven grand challenges in single-cell data science. Genome Biol. 21, 31 (2020).
Dann, E., Henderson, N. C., Teichmann, S. A., Morgan, M. D. & Marioni, J. C. Differential abundance testing on single-cell data using k-nearest neighbor graphs. Nat. Biotechnol. 40, 245–253 (2022).
Koussounadis, A., Langdon, S. P., Um, I. H., Harrison, D. J. & Smith, V. A. Relationship between differentially expressed mRNA and mRNA–protein correlations in a xenograft model system. Sci. Rep. 5, 10775 (2015).
Repapi, E., Agarwal, D., Napolitani, G., Sims, D. & Taylor, S. Integration of single-cell RNA-Seq and CyTOF data characterises heterogeneity of rare cell subpopulations. F1000Res. 11, 560 (2023).
Black, S. et al. CODEX multiplexed tissue imaging with DNA-conjugated antibodies. Nat. Protoc. 16, 3802–3835 (2021).
Clarke, G. M. et al. A novel, automated technology for multiplex biomarker imaging and application to breast cancer. Histopathology 64, 242–255 (2014).
Gerdes, M. J. et al. Highly multiplexed single-cell analysis of formalin-fixed, paraffin-embedded cancer tissue. Proc. Natl Acad. Sci. USA 110, 11982–11987 (2013).
Xie, R. et al. Spatially resolved gene expression prediction from histology images via bi-modal contrastive learning. In Adv. Neural Inf. Process. Syst. 36 (NeurIPS, 2023).
Lunger, I., Fawaz, M. & Rieger, M. A. Single-cell analyses to reveal hematopoietic stem cell fate decisions. FEBS Lett. 591, 2195–2212 (2017).
Wagner, D. E. & Klein, A. M. Lineage tracing meets single-cell omics: opportunities and challenges. Nat. Rev. Genet. 21, 410–427 (2020).
Brand, M. & Morrissey, E. Single-cell fate decisions of bipotential hematopoietic progenitors. Curr. Opin. Hematol. 27, 232–240 (2020).
Cunningham, R. P. & Porat-Shliom, N. Liver zonation — revisiting old questions with new technologies. Front. Physiol. 12, 732929 (2021).
Wolock, S. L., Lopez, R. & Klein, A. M. Scrublet: computational identification of cell doublets in single-cell transcriptomic data. Cell Syst. 8, 281–291.e9 (2019).
Kudryavtsev, B. N., Kudryavtseva, M. V., Sakuta, G. A. & Stein, G. I. Human hepatocyte polyploidization kinetics in the course of life cycle. Virchows Arch. B Cell Pathol. Incl. Mol. Pathol. 64, 387–393 (1993).
Kiselev, V. Y., Andrews, T. S. & Hemberg, M. Challenges in unsupervised clustering of single-cell RNA-seq data. Nat. Rev. Genet. 20, 273–282 (2019).
Zhang, J. M., Kamath, G. M. & Tse, D. N. Valid post-clustering differential analysis for single-cell RNA-Seq. Cell Syst. 9, 383–392.e6 (2019).
Kim, M. & Costello, J. DNA methylation: an epigenetic mark of cellular memory. Exp. Mol. Med. 49, e322 (2017).
Kim, M. S. et al. A draft map of the human proteome. Nature 509, 575–581 (2014).
Hu, C. CellMarker 2.0: an updated database of manually curated cell markers in human/mouse and web tools based on scRNA-seq data. Nucleic Acids Res. 51, D870–D876 (2023).
Osumi-Sutherland, D. et al. Cell type ontologies of the human cell atlas. Nat. Cell Biol. 23, 1129–1135 (2021).
Wang, S. et al. An autocrine signaling circuit in hepatic stellate cells underlies advanced fibrosis in nonalcoholic steatohepatitis. Sci. Transl. Med. 15, eadd3949 (2023).
Fred, R. G. et al. Single-cell transcriptome and cell type-specific molecular pathways of human non-alcoholic steatohepatitis. Sci. Rep. 12, 13484 (2022).
Poch, T. et al. Single-cell atlas of hepatic T cells reveals expansion of liver-resident naive-like CD4+ T cells in primary sclerosing cholangitis. J. Hepatol. 75, 414–423 (2021).
Ye, C. et al. Single-cell and spatial transcriptomics reveal the fibrosis-related immune landscape of biliary atresia. Clin. Transl. Med. 12, e1070 (2022).
Song, G. et al. Single-cell transcriptomic analysis suggests two molecularly subtypes of intrahepatic cholangiocarcinoma. Nat. Commun. 13, 1642 (2022).
Huang, H. et al. Single-cell transcriptomics uncovers cellular architecture and developmental trajectories in hepatoblastoma. Hepatology 77, 1911–1928 (2023).
Wang, X. et al. Comparative analysis of cell lineage differentiation during hepatogenesis in humans and mice at the single-cell transcriptome level. Cell Res. 30, 1109–1126 (2020).
Massalha, H. et al. A single cell atlas of the human liver tumor microenvironment. Mol. Syst. Biol. 16, e9682 (2020).
Chalasani, N. et al. Relationship of steatosis grade and zonal location to histological features of steatohepatitis in adult patients with non-alcoholic fatty liver disease. J. Hepatol. 48, 829–834 (2008).
Schwen, L. O. et al. Zonated quantification of steatosis in an entire mouse liver. Comput. Biol. Med. 73, 108–118 (2016).
Alexandrov, T. Spatial metabolomics: from a niche field towards a driver of innovation. Nat. Metab. 5, 1443–1445 (2023).
Deng, Y. et al. Spatial profiling of chromatin accessibility in mouse and human tissues. Nature 609, 375–383 (2022).
Xu, H. et al. SPACEL: deep learning-based characterization of spatial transcriptome architectures. Nat. Commun. 14, 7603 (2023).
Almagro-Pérez, C. et al. AI-driven 3D spatial transcriptomics. Preprint at https://arxiv.org/abs/2502.17761 (2025).
Lotfollahi, M. et al. Mapping single-cell data to reference atlases by transfer learning. Nat. Biotechnol. 40, 121–130 (2022).
Xu, C. et al. Probabilistic harmonization and annotation of single-cell transcriptomics data with deep generative models. Mol. Syst. Biol. 17, e9620 (2021).
Brbic, M. et al. MARS: discovering novel cell types across heterogeneous single-cell experiments. Nat. Methods 17, 1200–1206 (2020).
Song, H. et al. Single-cell analysis of hepatoblastoma identifies tumor signatures that predict chemotherapy susceptibility using patient-specific tumor spheroids. Nat. Commun. 13, 4878 (2022).
CZI Cell Science Program et al. CZ CELLxGENE discover: a single-cell data platform for scalable exploration, analysis and modeling of aggregated data. Nucleic Acids Res. 53, D886–D900 (2025).
Acknowledgements
The authors thank F. Hamade, biomedical illustrator, for creating the initial draft of the figures. They thank the HCA executive office, data portal and Lattice staff for outstanding project support. The HLiCA project is supported by funding from the Chan Zuckerberg Initiative (to S. A. Taylor, G.D.B., S.M., A.C.M., A.G.C., R.D., D.G., M.G., N.C.H., S.S.H., S.I., G.M.L., I.M., K.R.M., G.Q., A. Regev, A. Ricciuto, C.L.S., M.M.T., S. A. Teichmann, L.V., B.W. and M.Z.). Additional support was provided by the NIH National Institute of Diabetes and Digestive and Kidney Diseases (NIDDK) K08 grant DK121937 (to S. A. Taylor); NSERC Discovery grant RGPIN-2023-03419 and CIHR grant RN482632-481311 (to T.A.); CIHR grant PJT 180542 (to G.D.B.); CZI Seed Network grant CZIF2019-002429, Singapore National Medical Research Council (NMRC) grants NMRC/TCR/14-NUHS/2015 and NMRC/OFLCG19May-0038, and National Research Foundation grant (NRF-CRP) CRP26-2021-0005 (to R.D.); the Bundesministerium für Bildung und Forschung (BMBF) CureFib – 01EJ2201C (to D.G.); Ghent University BOF18-GOA-024 and BOF24/GOA/035 and FWO project grants 3G000519 and G013823N (to M.G.); Wellcome Trust Senior Research Fellowship in Clinical Science (ref. 219542/Z/19/Z) and the Medical Research Council (N.C.H.); R01DK132751 and R01DK120765 (S.S.H.); Helen and Martin Kimmel Award for Innovative Investigation, the Yad Abraham Research Center for Cancer Diagnostics and Therapy, the Moross Integrated Cancer Center, the Minerva Stiftung grant, a Weizmann-Sheba grant, the Israel Science Foundation grants no. 908/21 and 3663/21, the European Research Council (ERC) under the European Union’s Horizon 2020 Research and Innovation Programme grant no. 768956 and a grant from the Ministry of Innovation, Science & Technology, Israel (to S.I.); Massachusetts Life Science Center (MLSC) and NIH grants R01AA030770 and K08 DK115883 (to Z.G.J); the ERC (MyeFattyLiver no. 851908) and FWO project grants 3G000519, G075923N and G0A9Z24N (to C.L.S.); CSL Centenary Fellowship and National Health and Medical Research Council (NHMRC) Ideas Grant 2021/GNT2010795 (to A.S.); NIDDK R21 grant DK127275-01 (to M.M.T.); Advanced grant NewChol (to L.V.); NHLBI U54 HL 165440-01 grant (to I.S.V.); Hector Foundation, Dieter Morszeck Foundation and the German Research Foundation (SFB 1479) (to M.R.).
Author information
Authors and Affiliations
Contributions
The authors contributed equally to writing and reviewing/editing the manuscript before submission. S. A. Taylor, G.D.B., S.M. and A.C.M. researched data for the article and contributed substantially to discussion of the content.
Corresponding authors
Ethics declarations
Competing interests
S. A. Taylor serves as a consultant for Ipsen Pharmaceutical. A.C.M. receives funding from GSK and Boehringer Ingelheim for unrelated projects. A.J.G. receives research funding from Aligos Therapeutics, Bluejay Therapeutics, GSK, Roche and Vir Biotechnology, and performs scientific advisory services for Aligos Therapeutics, Arbutus Biopharma, Assembly Biosciences, Bluejay Therapeutics, Gilead Sciences, GSK, Janssen Pharmaceuticals, Roche, Vir Biotechnology, Virion Therapeutics and VBI for unrelated projects. D.G. serves as a consultant for Gordian Biotechnology. M.G. receives funding from the Sanofi iTech Award programme for an unrelated project. N.C.H. has received research funding from AbbVie, Pfizer, Gilead and Boehringer-Ingelheim, and is an adviser or consultant for Astra-Zeneca, GSK and MSD. S.S.H serves as a consultant for ARNATAR Therapeutics. Z.G.J. serves on the advisory board of Olix Pharmaceuticals, and received grants from Pfizer and Gilead Sciences. A. Ricciuto receives a stipend for academic lectures from Janssen and Organon. C.L.S. receives funding from Novo Nordisk for unrelated projects. M.M.T has served as a consultant for Merck for an unrelated project. L.V. is a shareholder of Definigen, BiliTech and Bit.bio. I.S.V. consults for Guidepoint Global, Cowen and Mosaic. B.W. serves as a consultant for Alnylam Pharmaceuticals, Disc Medicine, Mitsubishi Tanabe Pharma and Recordati Rare Diseases. G.D.B. advises Adela Bio and BioRender. The other authors declare no competing interests.
Peer review
Peer review information
Nature Reviews Gastroenterology & Hepatology thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Cell Annotation Platform: https://celltype.info
CellMarker: http://117.50.127.228/CellMarker
CELL×GENE: https://cellxgene.cziscience.com
HCA community: https://www.humancellatlas.org/join-the-hca
Human Cell Atlas: https://www.humancellatlas.org
Human Liver Cell Atlas: https://www.humancellatlas.org/biological-networks/liver-biological-network
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Taylor, S.A., Bader, G.D., MacParland, S. et al. Towards a reference cell atlas of liver diversity over the human lifespan. Nat Rev Gastroenterol Hepatol 23, 97–109 (2026). https://doi.org/10.1038/s41575-025-01114-3
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41575-025-01114-3