Abstract
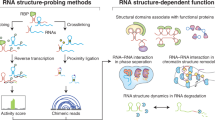
RNA is a key regulator of almost every cellular process, and the structures adopted by RNA molecules are thought to be central to their functions. The recent fast-paced evolution of high-throughput sequencing-based RNA structure mapping methods has enabled the rapid in vivo structural interrogation of entire cellular transcriptomes. Collectively, these studies are shedding new light on the long underestimated complexity of the structural organization of the transcriptome — the RNA structurome. Moreover, recent analyses are challenging the view that the RNA structurome is a static entity by revealing how RNA molecules establish intricate networks of alternative intramolecular and intermolecular interactions and that these ensembles of RNA structures are dynamically regulated to finely tune RNA functions in living cells. This new understanding of how RNA can shape cell phenotypes has important implications for the development of RNA-targeted therapeutic strategies.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Leppek, K., Das, R. & Barna, M. Functional 5′ UTR mRNA structures in eukaryotic translation regulation and how to find them. Nat. Rev. Mol. Cell Biol. 19, 158–174 (2018).
Mayr, C. Regulation by 3′-untranslated regions. Annu. Rev. Genet. 51, 171–194 (2017).
Frankish, A. et al. GENCODE 2021. Nucleic Acids Res. 49, D916–D923 (2021).
Fu, X.-D. Non-coding RNA: a new frontier in regulatory biology. Natl Sci. Rev. 1, 190–204 (2014).
Mustoe, A. M., Brooks, C. L. & Al-Hashimi, H. M. Hierarchy of RNA functional dynamics. Annu. Rev. Biochem. 83, 441–466 (2014).
Ganser, L. R., Kelly, M. L., Herschlag, D. & Al-Hashimi, H. M. The roles of structural dynamics in the cellular functions of RNAs. Nat. Rev. Mol. Cell Biol. 20, 474–489 (2019).
Kortmann, J. & Narberhaus, F. Bacterial RNA thermometers: molecular zippers and switches. Nat. Rev. Microbiol. 10, 255–265 (2012).
Serganov, A. & Nudler, E. A decade of riboswitches. Cell 152, 17–24 (2013).
Kubota, M., Tran, C. & Spitale, R. C. Progress and challenges for chemical probing of RNA structure inside living cells. Nat. Chem. Biol. 11, 933–941 (2015).
Strobel, E. J., Yu, A. M. & Lucks, J. B. High-throughput determination of RNA structures. Nat. Rev. Genet. 19, 615–634 (2018).
Kwok, C. K., Tang, Y., Assmann, S. M. & Bevilacqua, P. C. The RNA structurome: transcriptome-wide structure probing with next-generation sequencing. Trends Biochem. Sci. 40, 221–232 (2015).
Wells, S. E., Hughes, J. M., Igel, A. H. & Ares, M. Use of dimethyl sulfate to probe RNA structure in vivo. Methods Enzymol. 318, 479–493 (2000).
Mustoe, A. M., Lama, N. N., Irving, P. S., Olson, S. W. & Weeks, K. M. RNA base-pairing complexity in living cells visualized by correlated chemical probing. Proc. Natl Acad. Sci. USA 116, 24574–24582 (2019).
Mitchell, D. et al. Glyoxals as in vivo RNA structural probes of guanine base-pairing. RNA 24, 114–124 (2018).
Wang, P. Y., Sexton, A. N., Culligan, W. J. & Simon, M. D. Carbodiimide reagents for the chemical probing of RNA structure in cells. RNA 25, 135–146 (2019).
Mitchell, D. et al. In vivo RNA structural probing of uracil and guanine base-pairing by 1-ethyl-3-(3-dimethylaminopropyl)carbodiimide (EDC). RNA 25, 147–157 (2019).
Merino, E. J., Wilkinson, K. A., Coughlan, J. L. & Weeks, K. M. RNA structure analysis at single nucleotide resolution by selective 2′-hydroxyl acylation and primer extension (SHAPE). J. Am. Chem. Soc. 127, 4223–4231 (2005).
McGinnis, J. L., Dunkle, J. A., Cate, J. H. D. & Weeks, K. M. The mechanisms of RNA SHAPE chemistry. J. Am. Chem. Soc. 134, 6617–6624 (2012).
Xiao, L., Fang, L. & Kool, E. T. Acylation probing of “generic” RNA libraries reveals critical influence of loop constraints on reactivity. Cell Chem. Biol. 29, 1341–1352.e8 (2022).
Steen, K.-A., Rice, G. M. & Weeks, K. M. Fingerprinting noncanonical and tertiary RNA structures by differential SHAPE reactivity. J. Am. Chem. Soc. 134, 13160–13163 (2012).
Mortimer, S. A. & Weeks, K. M. Time-resolved RNA SHAPE chemistry. J. Am. Chem. Soc. 130, 16178–16180 (2008).
Busan, S., Weidmann, C. A., Sengupta, A. & Weeks, K. M. Guidelines for SHAPE reagent choice and detection strategy for RNA structure probing studies. Biochemistry 58, 2655–2664 (2019).
Spitale, R. C. et al. RNA SHAPE analysis in living cells. Nat. Chem. Biol. 9, 18–20 (2013).
Spitale, R. C. et al. Structural imprints in vivo decode RNA regulatory mechanisms. Nature 519, 486–490 (2015).
Marinus, T., Fessler, A. B., Ogle, C. A. & Incarnato, D. A novel SHAPE reagent enables the analysis of RNA structure in living cells with unprecedented accuracy. Nucleic Acids Res. 49, e34 (2021).
Ingle, S., Azad, R. N., Jain, S. S. & Tullius, T. D. Chemical probing of RNA with the hydroxyl radical at single-atom resolution. Nucleic Acids Res. 42, 12758–12767 (2014).
Kielpinski, L. J. & Vinther, J. Massive parallel-sequencing-based hydroxyl radical probing of RNA accessibility. Nucleic Acids Res. 42, e70 (2014).
Feng, C. et al. Light-activated chemical probing of nucleobase solvent accessibility inside cells. Nat. Chem. Biol. 14, 276–283 (2018).
Zinshteyn, B. et al. Assaying RNA structure with LASER-Seq. Nucleic Acids Res. 47, 43–55 (2019).
Homan, P. J. et al. Single-molecule correlated chemical probing of RNA. Proc. Natl Acad. Sci. USA 111, 13858–13863 (2014).
Siegfried, N. A., Busan, S., Rice, G. M., Nelson, J. A. E. & Weeks, K. M. RNA motif discovery by SHAPE and mutational profiling (SHAPE-MaP). Nat. Methods 11, 959–965 (2014).
Zubradt, M. et al. DMS-MaPseq for genome-wide or targeted RNA structure probing in vivo. Nat. Methods 14, 75–82 (2017).
Guo, L.-T. et al. Sequencing and structure probing of long RNAs using MarathonRT: a next-generation reverse transcriptase. J. Mol. Biol. 432, 3338–3352 (2020).
Cimino, G. D., Gamper, H. B., Isaacs, S. T. & Hearst, J. E. Psoralens as photoactive probes of nucleic acid structure and function: organic chemistry, photochemistry, and biochemistry. Annu. Rev. Biochem. 54, 1151–1193 (1985).
Nilsen, T. W. Detecting RNA-RNA interactions using psoralen derivatives. Cold Spring Harb. Protoc. 2014, 996–1000 (2014).
Ramani, V., Qiu, R. & Shendure, J. High-throughput determination of RNA structure by proximity ligation. Nat. Biotechnol. 33, 980–984 (2015).
Lu, Z. et al. RNA duplex map in living cells reveals higher order transcriptome structure. Cell 165, 1267–1279 (2016).
Aw, J. G. A. et al. In vivo mapping of eukaryotic RNA interactomes reveals principles of higher-order organization and regulation. Mol. Cell 62, 603–617 (2016).
Sharma, E., Sterne-Weiler, T., O’Hanlon, D. & Blencowe, B. J. Global mapping of human RNA–RNA interactions. Mol. Cell 62, 618–626 (2016).
Nguyen, T. C. et al. Mapping RNA–RNA interactome and RNA structure in vivo by MARIO. Nat. Commun. 7, 12023 (2016).
Ziv, O. et al. COMRADES determines in vivo RNA structures and interactions. Nat. Methods 15, 785–788 (2018).
Velema, W. A., Park, H. S., Kadina, A., Orbai, L. & Kool, E. T. Trapping transient RNA complexes by chemically reversible acylation. Angew. Chem. Int. Ed. Engl. 59, 22017–22022 (2020).
Van Damme, R. et al. Chemical reversible crosslinking enables measurement of RNA 3D distances and alternative conformations in cells. Nat. Commun. 13, 911 (2022).
Christy, T. W. et al. Direct mapping of higher-order RNA interactions by SHAPE-JuMP. Biochemistry 60, 1971–1982 (2021).
Corley, M. et al. Footprinting SHAPE-eCLIP reveals transcriptome-wide hydrogen bonds at RNA–protein interfaces. Mol. Cell 80, 903–914.e8 (2020).
Chan, D. et al. Diverse functional elements in RNA predicted transcriptome-wide by orthogonal RNA structure probing. Nucleic Acids Res. 49, 11868–11882 (2021).
Weidmann, C. A., Mustoe, A. M., Jariwala, P. B., Calabrese, J. M. & Weeks, K. M. Analysis of RNA–protein networks with RNP-MaP defines functional hubs on RNA. Nat. Biotechnol. 39, 347–356 (2021).
Li, P. et al. Integrative analysis of Zika virus genome RNA structure reveals critical determinants of viral infectivity. Cell Host Microbe 24, 875–886.e5 (2018).
Huber, R. G. et al. Structure mapping of dengue and Zika viruses reveals functional long-range interactions. Nat. Commun. 10, 1408 (2019).
Ziv, O. et al. The short- and long-range RNA–RNA interactome of SARS-CoV-2. Mol. Cell 80, 1067–1077.e5 (2020).
Yang, S. L. et al. Comprehensive mapping of SARS-CoV-2 interactions in vivo reveals functional virus–host interactions. Nat. Commun. 12, 5113 (2021).
Zhang, Y. et al. In vivo structure and dynamics of the SARS-CoV-2 RNA genome. Nat. Commun. 12, 5695 (2021).
Uroda, T. et al. Conserved pseudoknots in lncRNA MEG3 are essential for stimulation of the p53 pathway. Mol. Cell 75, 982–995.e9 (2019).
Mustoe, A. M. et al. Pervasive regulatory functions of mRNA structure revealed by high-resolution SHAPE probing. Cell 173, 181–195.e18 (2018).
Beaudoin, J.-D. et al. Analyses of mRNA structure dynamics identify embryonic gene regulatory programs. Nat. Struct. Mol. Biol. 25, 677–686 (2018).
Rouskin, S., Zubradt, M., Washietl, S., Kellis, M. & Weissman, J. S. Genome-wide probing of RNA structure reveals active unfolding of mRNA structures in vivo. Nature 505, 701–705 (2014).
Sanford, T. J., Mears, H. V., Fajardo, T., Locker, N. & Sweeney, T. R. Circularization of flavivirus genomic RNA inhibits de novo translation initiation. Nucleic Acids Res. 47, 9789–9802 (2019).
Gabryelska, M. M. et al. Global mapping of RNA homodimers in living cells. Genome Res. 32, 956–967 (2022).
Dethoff, E. A., Chugh, J., Mustoe, A. M. & Al-Hashimi, H. M. Functional complexity and regulation through RNA dynamics. Nature 482, 322–330 (2012).
Herschlag, D. RNA chaperones and the RNA folding problem. J. Biol. Chem. 270, 20871–20874 (1995).
Ding, Y. et al. In vivo genome-wide profiling of RNA secondary structure reveals novel regulatory features. Nature 505, 696–700 (2014).
Incarnato, D., Neri, F., Anselmi, F. & Oliviero, S. Genome-wide profiling of mouse RNA secondary structures reveals key features of the mammalian transcriptome. Genome Biol. 15, 491 (2014).
Watters, K. E., Strobel, E. J., Yu, A. M., Lis, J. T. & Lucks, J. B. Cotranscriptional folding of a riboswitch at nucleotide resolution. Nat. Struct. Mol. Biol. 23, 1124–1131 (2016).
Incarnato, D. et al. In vivo probing of nascent RNA structures reveals principles of cotranscriptional folding. Nucleic Acids Res. 45, 9716–9725 (2017).
Cheng, C. Y., Kladwang, W., Yesselman, J. D. & Das, R. RNA structure inference through chemical mapping after accidental or intentional mutations. Proc. Natl Acad. Sci. USA 114, 9876–9881 (2017).
Byeon, G. W. et al. Functional and structural basis of extreme conservation in vertebrate 5′ untranslated regions. Nat. Genet. 53, 729–741 (2021).
Cordero, P. & Das, R. Rich RNA structure landscapes revealed by mutate-and-map analysis. PLoS Comput. Biol. 11, e1004473 (2015).
Aviran, S. & Incarnato, D. Computational approaches for RNA structure ensemble deconvolution from structure probing data. J. Mol. Biol. https://doi.org/10.1016/j.jmb.2022.167635 (2022).
Li, H. & Aviran, S. Statistical modeling of RNA structure profiling experiments enables parsimonious reconstruction of structure landscapes. Nat. Commun. 9, 606 (2018).
Spasic, A., Assmann, S. M., Bevilacqua, P. C. & Mathews, D. H. Modeling RNA secondary structure folding ensembles using SHAPE mapping data. Nucleic Acids Res. 46, 314–323 (2018).
Zhou, J. et al. IRIS: a method for predicting in vivo RNA secondary structures using PARIS data. Quant. Biol. 8, 369–381 (2020).
McCaskill, J. S. The equilibrium partition function and base pair binding probabilities for RNA secondary structure. Biopolymers 29, 1105–1119 (1990).
Tomezsko, P. J. et al. Determination of RNA structural diversity and its role in HIV-1 RNA splicing. Nature 582, 438–442 (2020).
Morandi, E. et al. Genome-scale deconvolution of RNA structure ensembles. Nat. Methods 18, 249–252 (2021).
Olson, S. W. et al. Discovery of a large-scale, cell-state-responsive allosteric switch in the 7SK RNA using DANCE-MaP. Mol. Cell 82, 1708–1723.e10 (2022).
Wu, M. T.-P. & D’Souza, V. Alternate RNA structures. Cold Spring Harb. Perspect. Biol. 12, a032425 (2020).
Sherpa, C., Rausch, J. W., Le Grice, S. F. J., Hammarskjold, M.-L. & Rekosh, D. The HIV-1 Rev response element (RRE) adopts alternative conformations that promote different rates of virus replication. Nucleic Acids Res. 43, 4676–4686 (2015).
Lan, T. C. T. et al. Secondary structural ensembles of the SARS-CoV-2 RNA genome in infected cells. Nat. Commun. 13, 1128 (2022).
Manfredonia, I. & Incarnato, D. Structure and regulation of coronavirus genomes: state-of-the-art and novel insights from SARS-CoV-2 studies. Biochemical Soc. Trans. 49, 341–352 (2020).
Plant, E. P. et al. A three-stemmed mRNA pseudoknot in the SARS coronavirus frameshift signal. PLoS Biol. 3, e172 (2005).
Rangan, R. et al. De novo 3D models of SARS-CoV-2 RNA elements from consensus experimental secondary structures. Nucleic Acids Res. 49, 3092–3108 (2021).
Manfredonia, I. et al. Genome-wide mapping of SARS-CoV-2 RNA structures identifies therapeutically-relevant elements. Nucleic Acids Res. 48, 12436–12452 (2020).
Huston, N. C. et al. Comprehensive in vivo secondary structure of the SARS-CoV-2 genome reveals novel regulatory motifs and mechanisms. Mol. Cell 81, 584–598.e5 (2021).
Schlick, T. et al. To knot or not to knot: multiple conformations of the SARS-CoV-2 frameshifting RNA element. J. Am. Chem. Soc. 143, 11404–11422 (2021).
Park, S.-J., Kim, Y.-G. & Park, H.-J. Identification of RNA pseudoknot-binding ligand that inhibits the –1 ribosomal frameshifting of SARS-coronavirus by structure-based virtual screening. J. Am. Chem. Soc. 133, 10094–10100 (2011).
Sun, Y. et al. Restriction of SARS-CoV-2 replication by targeting programmed −1 ribosomal frameshifting. Proc. Natl Acad. Sci. USA 118, e2023051118 (2021).
Fujinaga, K. P-TEFb as a promising therapeutic target. Molecules 25, E838 (2020).
Hsue, B. & Masters, P. S. A bulged stem-loop structure in the 3’ untranslated region of the genome of the coronavirus mouse hepatitis virus is essential for replication. J. Virol. 71, 7567–7578 (1997).
Robertson, M. P. et al. The structure of a rigorously conserved RNA element within the SARS virus genome. PLOS Biol. 3, e5 (2004).
Guo, A.-X., Cui, J.-J., Wang, L.-Y. & Yin, J.-Y. The role of CSDE1 in translational reprogramming and human diseases. Cell Commun. Signal. 18, 14 (2020).
Hajdin, C. E. et al. Accurate SHAPE-directed RNA secondary structure modeling, including pseudoknots. Proc. Natl Acad. Sci. USA 110, 5498–5503 (2013).
Batey, R. T. Riboswitches: still a lot of undiscovered country. RNA 21, 560–563 (2015).
Fazal, F. M. et al. Atlas of subcellular RNA localization revealed by APEX-seq. Cell 178, 473–490.e26 (2019).
Singha, M., Spitalny, L., Nguyen, K., Vandewalle, A. & Spitale, R. C. Chemical methods for measuring RNA expression with metabolic labeling. Wiley Interdiscip. Rev. RNA 12, e1650 (2021).
Yang, M. et al. In vivo single-molecule analysis reveals COOLAIR RNA structural diversity. Nature https://doi.org/10.1038/s41586-022-05135-9 (2022).
Sengupta, A., Rice, G. M. & Weeks, K. M. Single-molecule correlated chemical probing reveals large-scale structural communication in the ribosome and the mechanism of the antibiotic spectinomycin in living cells. PLOS Biol. 17, e3000393 (2019).
Zeller, M. J. et al. SHAPE-enabled fragment-based ligand discovery for RNA. Proc. Natl Acad. Sci. USA 119, e2122660119 (2022).
Fang, L. et al. Pervasive transcriptome interactions of protein-targeted drugs. Preprint at https://doi.org/10.1101/2022.07.18.500496 (2022).
Bushhouse, D. Z., Choi, E. K., Hertz, L. M. & Lucks, J. B. How does RNA fold dynamically? J. Mol. Biol. 167665 https://doi.org/10.1016/j.jmb.2022.167665 (2022).
Strobel, E. J., Cheng, L., Berman, K. E., Carlson, P. D. & Lucks, J. B. A ligand-gated strand displacement mechanism for ZTP riboswitch transcription control. Nat. Chem. Biol. 15, 1067–1076 (2019).
Cheng, L. et al. Cotranscriptional RNA strand exchange underlies the gene regulation mechanism in a purine-sensing transcriptional riboswitch. Nucleic Acids Res. https://doi.org/10.1093/nar/gkac102 (2022).
Saldi, T., Riemondy, K., Erickson, B. & Bentley, D. L. Alternative RNA structures formed during transcription depend on elongation rate and modify RNA processing. Mol. Cell 81, 1789–1801.e5 (2021).
Fu, Y., Dominissini, D., Rechavi, G. & He, C. Gene expression regulation mediated through reversible m6A RNA methylation. Nat. Rev. Genet. 15, 293–306 (2014).
Roost, C. et al. Structure and thermodynamics of N6-methyladenosine in RNA: a spring-loaded base modification. J. Am. Chem. Soc. 137, 2107–2115 (2015).
Liu, N. et al. N6-Methyladenosine-dependent RNA structural switches regulate RNA–protein interactions. Nature 518, 560–564 (2015).
Aw, J. G. A. et al. Determination of isoform-specific RNA structure with nanopore long reads. Nat. Biotechnol. 39, 336–346 (2021).
Sun, L. et al. RNA structure maps across mammalian cellular compartments. Nat. Struct. Mol. Biol. 26, 322–330 (2019).
Liu, Z. et al. In vivo nuclear RNA structurome reveals RNA-structure regulation of mRNA processing in plants. Genome Biol. 22, 11 (2021).
Ray, P. S. et al. A stress-responsive RNA switch regulates VEGF expression. Nature 457, 915–919 (2009).
Acknowledgements
This work was supported by funding from the Groningen Biomolecular Sciences and Biotechnology Institute (GBB, University of Groningen) to D.I.
Author information
Authors and Affiliations
Contributions
The authors contributed equally to all aspects of the article.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Reviews Genetics thanks Y. Ding and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Glossary
- Boltzmann distribution
-
A probability distribution describing the probability that a system will be in a certain state (in this case, a certain RNA conformation) as a function of the state’s energy and of the system’s temperature.
- Covariation
-
In an RNA multiple sequence alignment, two covarying positions are those for which the sequence changes but their ability to base-pair is preserved.
- Hydrogen abstraction
-
Removal of an atom or group from a molecule by a radical.
- Pseudoknot
-
A non-nested structural RNA motif formed upon base-pairing between the loop of a secondary structure element (such as a stem-loop (SL)) and any complementary region along the RNA.
- RNA structurome
-
The full range of RNA structures formed by the transcriptome of an organism.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Spitale, R.C., Incarnato, D. Probing the dynamic RNA structurome and its functions. Nat Rev Genet 24, 178–196 (2023). https://doi.org/10.1038/s41576-022-00546-w
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41576-022-00546-w
This article is cited by
-
Cryo-specimen preparation and imaging of highly-structured and dynamic large non-coding RNAs
BMC Methods (2025)
-
Decoding the interactions and functions of non-coding RNA with artificial intelligence
Nature Reviews Molecular Cell Biology (2025)
-
Functions and therapeutic applications of pseudouridylation
Nature Reviews Molecular Cell Biology (2025)
-
Mapping of HOCl-oxidized RNA identifies abasic sites as major damage and oxidation product of oxo8G
Nature Communications (2025)
-
RNA sample optimization for cryo-EM analysis
Nature Protocols (2025)