Abstract
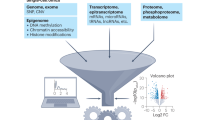
Recent advances in single-cell technologies have enabled high-throughput molecular profiling of cells across modalities and locations. Single-cell transcriptomics data can now be complemented by chromatin accessibility, surface protein expression, adaptive immune receptor repertoire profiling and spatial information. The increasing availability of single-cell data across modalities has motivated the development of novel computational methods to help analysts derive biological insights. As the field grows, it becomes increasingly difficult to navigate the vast landscape of tools and analysis steps. Here, we summarize independent benchmarking studies of unimodal and multimodal single-cell analysis across modalities to suggest comprehensive best-practice workflows for the most common analysis steps. Where independent benchmarks are not available, we review and contrast popular methods. Our article serves as an entry point for novices in the field of single-cell (multi-)omic analysis and guides advanced users to the most recent best practices.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Zappia, L. & Theis, F. J. Over 1000 tools reveal trends in the single-cell RNA-seq analysis landscape. Genome Biol. 22, 301 (2021).
Amezquita, R. A. et al. Orchestrating single-cell analysis with bioconductor. Nat. Methods 17, 137–145 (2020).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587.e29 (2021).
Wolf, F. A., Angerer, P. & Theis, F. J. SCANPY: large-scale single-cell gene expression data analysis. Genome Biol. 19, 15 (2018).
Luecken, M. D. & Theis, F. J. Current best practices in single-cell RNA-seq analysis: a tutorial. Mol. Syst. Biol. 15, e8746 (2019). This scRNA-seq-focused review and workflow template served as inspiration for the current benchmark-based Expert Recommendation.
Kharchenko, P. V. The triumphs and limitations of computational methods for scRNA-seq. Nat. Methods 18, 723–732 (2021). The article highlights limitations of computational methods, which analysts should be aware of.
Sikkema, L. et al. An integrated cell atlas of the human lung in health and disease. bioRxiv https://doi.org/10.1101/2022.03.10.483747 (2022).
Eraslan, G. et al. Single-nucleus cross-tissue molecular reference maps toward understanding disease gene function. Science 376, eabl4290 (2022).
Baek, S. & Lee, I. Single-cell ATAC sequencing analysis: from data preprocessing to hypothesis generation. Comput. Struct. Biotechnol. J. 18, 1429–1439 (2020). This article serves as an introduction to the scATAC-seq analysis workflow.
Stoeckius, M. et al. Simultaneous epitope and transcriptome measurement in single cells. Nat. Methods 14, 865–868 (2017). This article introduces CITE-seq, which is one of the two essential assays for surface protein measurements.
Han, A., Glanville, J., Hansmann, L. & Davis, M. M. Linking T-cell receptor sequence to functional phenotype at the single-cell level. Nat. Biotechnol. 32, 684–692 (2014).
Larsson, L., Frisén, J. & Lundeberg, J. Spatially resolved transcriptomics adds a new dimension to genomics. Nat. Methods 18, 15–18 (2021).
Rai, V. et al. Single-cell ATAC-seq in human pancreatic islets and deep learning upscaling of rare cells reveals cell-specific type 2 diabetes regulatory signatures. Mol. Metab. 32, 109–121 (2020).
Unterman, A. et al. Single-cell multi-omics reveals dyssynchrony of the innate and adaptive immune system in progressive COVID-19. Nat. Commun. 13, 440 (2022).
Gangaev, A. et al. Identification and characterization of a SARS-CoV-2 specific CD8+ T cell response with immunodominant features. Nat. Commun. 12, 2593 (2021).
Dhainaut, M. et al. Spatial CRISPR genomics identifies regulators of the tumor microenvironment. Cell 185, 1223–1239.e20 (2022).
Stuart, T. & Satija, R. Integrative single-cell analysis. Nat. Rev. Genet. 20, 257–272 (2019). A review of the advent of multimodal single-cell data with a focus on the experimental assays and data integration.
Mereu, E. et al. Benchmarking single-cell RNA-sequencing protocols for cell atlas projects. Nat. Biotechnol. 38, 747–755 (2020).
Lafzi, A., Moutinho, C., Picelli, S. & Heyn, H. Tutorial: guidelines for the experimental design of single-cell RNA sequencing studies. Nat. Protoc. 13, 2742–2757 (2018).
Germain, P.-L., Sonrel, A. & Robinson, M. D. pipeComp, a general framework for the evaluation of computational pipelines, reveals performant single cell RNA-seq preprocessing tools. Genome Biol. 21, 227 (2020).
Young, M. D. & Behjati, S. SoupX removes ambient RNA contamination from droplet-based single-cell RNA sequencing data. Gigascience 9, giaa151 (2020).
Fleming, S. J. et al. Unsupervised removal of systematic background noise from droplet-based single-cell experiments using CellBender. bioRxiv https://doi.org/10.1101/791699 (2022).
Yang, S. et al. Decontamination of ambient RNA in single-cell RNA-seq with DecontX. Genome Biol. 21, 57 (2020).
Germain, P.-L., Lun, A., Garcia Meixide, C., Macnair, W. & Robinson, M. D. Doublet identification in single-cell sequencing data using scDblFinder. F1000Res. 10, 979 (2021).
Xi, N. M. & Li, J. J. Protocol for executing and benchmarking eight computational doublet-detection methods in single-cell RNA sequencing data analysis. Star. Protoc. 2, 100699 (2021).
Xi, N. M. & Li, J. J. Benchmarking computational doublet-detection methods for single-cell RNA sequencing data. Cell Syst. 12, 176–194.e6 (2021).
Neavin, D. et al. Demuxafy: improvement in droplet assignment by integrating multiple single-cell demultiplexing and doublet detection methods. bioRxiv https://doi.org/10.1101/2022.03.07.483367 (2022).
Vallejos, C. A., Risso, D., Scialdone, A., Dudoit, S. & Marioni, J. C. Normalizing single-cell RNA sequencing data: challenges and opportunities. Nat. Methods 14, 565–571 (2017).
Ahlmann-Eltze, C. & Huber, W. Comparison of transformations for single-cell RNA-seq data. bioRxiv https://doi.org/10.1101/2021.06.24.449781 (2022).
Lun, A. T. L., Bach, K. & Marioni, J. C. Pooling across cells to normalize single-cell RNA sequencing data with many zero counts. Genome Biol. 17, 75 (2016).
Lause, J., Berens, P. & Kobak, D. Analytic Pearson residuals for normalization of single-cell RNA-seq UMI data. Genome Biol. 22, 258 (2021).
Ahlmann-Eltze, C. & Huber, W. Comparison of transformations for single-cell RNA-seq data. bioRxiv https://doi.org/10.1101/2021.06.24.449781 (2022).
Sina Booeshaghi, A., Hallgrímsdóttir, I. B., Gálvez-Merchán, Á. & Pachter, L. Depth normalization for single-cell genomics count data. bioRxiv https://doi.org/10.1101/2022.05.06.490859 (2022).
Vieth, B., Parekh, S., Ziegenhain, C., Enard, W. & Hellmann, I. A systematic evaluation of single cell RNA-seq analysis pipelines. Nat. Commun. 10, 4667 (2019).
Luecken, M. D. et al. Benchmarking atlas-level data integration in single-cell genomics. Nat. Methods 19, 41–50 (2022).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Tran, H. T. N. et al. A benchmark of batch-effect correction methods for single-cell RNA sequencing data. Genome Biol. 21, 12 (2020).
Chazarra-Gil, R., van Dongen, S., Kiselev, V. Y. & Hemberg, M. Flexible comparison of batch correction methods for single-cell RNA-seq using BatchBench. Nucleic Acids Res. 49, e42 (2021).
Xu, C. et al. Probabilistic harmonization and annotation of single-cell transcriptomics data with deep generative models. Mol. Syst. Biol. 17, e9620 (2021).
Lopez, R., Regier, J., Cole, M. B., Jordan, M. I. & Yosef, N. Deep generative modeling for single-cell transcriptomics. Nat. Methods 15, 1053–1058 (2018).
Lotfollahi, M., Wolf, F. A. & Theis, F. J. scGen predicts single-cell perturbation responses. Nat. Methods 16, 715–721 (2019).
Hie, B., Bryson, B. & Berger, B. Efficient integration of heterogeneous single-cell transcriptomes using Scanorama. Nat. Biotechnol. 37, 685–691 (2019).
Chervov, A. & Zinovyev, A. Computational challenges of cell cycle analysis using single cell transcriptomics. arXiv https://doi.org/10.48550/arXiv.2208.05229 (2022).
Hafemeister, C. & Satija, R. Normalization and variance stabilization of single-cell RNA-seq data using regularized negative binomial regression. Genome Biol. 20, 296 (2019).
Zheng, S. C. et al. Universal prediction of cell-cycle position using transfer learning. Genome Biol. 23, 41 (2022).
Townes, F. W., Hicks, S. C., Aryee, M. J. & Irizarry, R. A. Feature selection and dimension reduction for single-cell RNA-seq based on a multinomial model. Genome Biol. 20, 295 (2019).
Moon, K. R. et al. Visualizing structure and transitions in high-dimensional biological data. Nat. Biotechnol. 37, 1482–1492 (2019).
Chari, T., Banerjee, J. & Pachter, L. The specious art of single-cell genomics. bioRxiv https://doi.org/10.1101/2021.08.25.457696 (2022).
Duò, A., Robinson, M. D. & Soneson, C. A systematic performance evaluation of clustering methods for single-cell RNA-seq data. F1000Res. 7, 1141 (2018).
Freytag, S., Tian, L., Lönnstedt, I., Ng, M. & Bahlo, M. Comparison of clustering tools in R for medium-sized 10x Genomics single-cell RNA-sequencing data. F1000Res. 7, 1297 (2018).
Traag, V. A., Waltman, L. & van Eck, N. J. From Louvain to Leiden: guaranteeing well-connected communities. Sci. Rep. 9, 5233 (2019).
Clarke, Z. A. et al. Tutorial: guidelines for annotating single-cell transcriptomic maps using automated and manual methods. Nat. Protoc. 16, 2749–2764 (2021).
Abdelaal, T. et al. A comparison of automatic cell identification methods for single-cell RNA sequencing data. Genome Biol. 20, 194 (2019).
Pasquini, G., Rojo Arias, J. E., Schäfer, P. & Busskamp, V. Automated methods for cell type annotation on scRNA-seq data. Comput. Struct. Biotechnol. J. 19, 961–969 (2021).
Domínguez Conde, C. et al. Cross-tissue immune cell analysis reveals tissue-specific features in humans. Science 376, eabl5197 (2022).
Fu, R. et al. clustifyr: an R package for automated single-cell RNA sequencing cluster classification. F1000Research 9, 223 (2020).
Lotfollahi, M., Naghipourfar, M. & Luecken, M. D. Mapping single-cell data to reference atlases by transfer learning. Nat Biotechnol 40, 121–130 (2022).
Kang, J. B. et al. Efficient and precise single-cell reference atlas mapping with Symphony. Nat. Commun. 12, 5890 (2021).
Pullin, J. M. & McCarthy, D. J. A comparison of marker gene selection methods for single-cell RNA sequencing data. bioRxiv https://doi.org/10.1101/2022.05.09.490241 (2022).
Zhang, J. M., Kamath, G. M. & Tse, D. N. Valid post-clustering differential analysis for single-cell RNA-seq. Cell Syst. 9, 383–392.e6 (2019).
Street, K. et al. Slingshot: cell lineage and pseudotime inference for single-cell transcriptomics. BMC Genomics 19, 477 (2018).
Wolf, F. A. et al. PAGA: graph abstraction reconciles clustering with trajectory inference through a topology preserving map of single cells. Genome Biol. 20, 59 (2019).
Grün, D. et al. De novo prediction of stem cell identity using single-cell transcriptome data. Cell Stem Cell 19, 266–277 (2016).
Saelens, W., Cannoodt, R., Todorov, H. & Saeys, Y. A comparison of single-cell trajectory inference methods: towards more accurate and robust tools. Nat. Biotechnol. 37, 547–554 (2019).
La Manno, G. et al. RNA velocity of single cells. Nature 560, 494–498 (2018).
Bergen, V., Lange, M., Peidli, S., Wolf, F. A. & Theis, F. J. Generalizing RNA velocity to transient cell states through dynamical modeling. Nat. Biotechnol. 38, 1408–1414 (2020).
Lange, M. et al. CellRank for directed single-cell fate mapping. Nat. Methods 19, 159–170 (2022).
Weiler, P., Van den Berge, K., Street, K. & Tiberi, S. A guide to trajectory inference and RNA velocity. Methods Mol. Biol. 2584, 269–292 (2023).
Bergen, V., Soldatov, R. A., Kharchenko, P. V. & Theis, F. J. RNA velocity-current challenges and future perspectives. Mol. Syst. Biol. 17, e10282 (2021).
Gorin, G., Fang, M., Chari, T. & Pachter, L. RNA velocity unraveled. PLoS Comput. Biol. 18, e1010492 (2022).
Jones, M. G. et al. Inference of single-cell phylogenies from lineage tracing data using Cassiopeia. Genome Biol. 21, 92 (2020).
Sokal, R. & Michener, C. A statistical method for evaluating systematic relationships. Univ. Kans., Sci. Bull. 38, 1409–1438 (1958).
Saitou, N. & Nei, M. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 4, 406–425 (1987).
Gong, W. et al. Benchmarked approaches for reconstruction of in vitro cell lineages and in silico models of C. elegans and M. musculus developmental trees. Cell Syst. 12, 810–826.e4 (2021).
Forrow, A. & Schiebinger, G. LineageOT is a unified framework for lineage tracing and trajectory inference. Nat. Commun. 12, 4940 (2021).
McKenna, A. & Gagnon, J. A. Recording development with single cell dynamic lineage tracing. Development 146, dev169730 (2019).
Wang, S.-W., Herriges, M. J., Hurley, K., Kotton, D. N. & Klein, A. M. CoSpar identifies early cell fate biases from single-cell transcriptomic and lineage information. Nat. Biotechnol. 40, 1066–1074 (2022).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Finak, G. et al. MAST: a flexible statistical framework for assessing transcriptional changes and characterizing heterogeneity in single-cell RNA sequencing data. Genome Biol. 16, 278 (2015).
Wang, T., Li, B., Nelson, C. E. & Nabavi, S. Comparative analysis of differential gene expression analysis tools for single-cell RNA sequencing data. BMC Bioinformatics 20, 40 (2019).
Das, S., Rai, A., Merchant, M. L., Cave, M. C. & Rai, S. N. A comprehensive survey of statistical approaches for differential expression analysis in single-cell RNA sequencing studies. Genes 12, 1947 (2021).
Soneson, C. & Robinson, M. D. Bias, robustness and scalability in single-cell differential expression analysis. Nat. Methods 15, 255–261 (2018).
Squair, J. W. et al. Confronting false discoveries in single-cell differential expression. Nat. Commun. 12, 5692 (2021).
Zimmerman, K. D., Espeland, M. A. & Langefeld, C. D. A practical solution to pseudoreplication bias in single-cell studies. Nat. Commun. 12, 738 (2021).
Junttila, S., Smolander, J. & Elo, L. L. Benchmarking methods for detecting differential states between conditions from multi-subject single-cell RNA-seq data. Brief. Bioinform. 23, bbac286 (2022).
Law, C. W. et al. A guide to creating design matrices for gene expression experiments. F1000Res. 9, 1444 (2020).
Thurman, A. L., Ratcliff, J. A., Chimenti, M. S. & Pezzulo, A. A. Differential gene expression analysis for multi-subject single cell RNA sequencing studies with aggregateBioVar. Bioinformatics 37, 3243–3251 (2021).
Lähnemann, D. et al. Eleven grand challenges in single-cell data science. Genome Biol. 21, 31 (2020).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. Ser. B Stat. Methodol. 57, 289–300 (1995).
Liberzon, A. et al. Molecular signatures database (MSigDB) 3.0. Bioinformatics 27, 1739–1740 (2011).
Ashburner, M. et al. Gene ontology: tool for the unification of biology. Nat. Genet. 25, 25–29 (2000).
Kanehisa, M., Furumichi, M., Tanabe, M., Sato, Y. & Morishima, K. KEGG: new perspectives on genomes, pathways, diseases and drugs. Nucleic Acids Res. 45, D353–D361 (2017).
Gillespie, M. et al. The reactome pathway knowledgebase 2022. Nucleic Acids Res. 50, D687–D692 (2022).
Schubert et al. Perturbation-response genes reveal signaling footprints in cancer gene expression. Nat. Commun. 9, 20 (2018).
Garcia-Alonso, L., Holland, C. H., Ibrahim, M. M., Turei, D. & Saez-Rodriguez, J. Benchmark and integration of resources for the estimation of human transcription factor activities. Genome Res. 29, 1363–1375 (2019).
Korotkevich, G. et al. Fast gene set enrichment analysis. bioRxiv https://doi.org/10.1101/060012 (2021).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Hänzelmann, S., Castelo, R. & Guinney, J. GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinformatics 14, 7 (2013).
Holland, C. H. et al. Robustness and applicability of transcription factor and pathway analysis tools on single-cell RNA-seq data. Genome Biol. 21, 36 (2020).
Badia-i-Mompel, P. et al. decoupleR: ensemble of computational methods to infer biological activities from omics data. Bioinform. Adv. 2, vbac016 (2022).
Barkas, N., Pethukov, V., Kharchenko, P. and Biederstedt, E. pagoda2: Single Cell Analysis and Differential Expression, https://github.com/kharchenkolab/pagoda2 (2021).
Zhang, Y. et al. Benchmarking algorithms for pathway activity transformation of single-cell RNA-seq data. Comput. Struct. Biotechnol. J. 18, 2953–2961 (2020).
Pijuan-Sala, B. et al. A single-cell molecular map of mouse gastrulation and early organogenesis. Nature 566, 490–495 (2019).
Smillie, C. S. et al. Intra- and inter-cellular rewiring of the human colon during ulcerative colitis. Cell 178, 714–730.e22 (2019).
Büttner, M., Ostner, J., Müller, C. L., Theis, F. J. & Schubert, B. scCODA is a Bayesian model for compositional single-cell data analysis. Nat. Commun. 12, 6876 (2021).
Cao, Y. et al. scDC: single cell differential composition analysis. BMC Bioinformatics 20(Suppl. 19), 721 (2019).
Ostner, J., Carcy, S. & Müller, C. L. tascCODA: Bayesian tree-aggregated analysis of compositional amplicon and single-cell data. Front. Genet. 12, 766405 (2021).
Zhao, J. et al. Detection of differentially abundant cell subpopulations in scRNA-seq data. Proc. Natl Acad. Sci. USA 118, e2100293118 (2021).
Dann, E., Henderson, N. C., Teichmann, S. A., Morgan, M. D. & Marioni, J. C. Differential abundance testing on single-cell data using k-nearest neighbor graphs. Nat. Biotechnol. 40, 245–253 (2022).
Srivatsan, S. R. et al. Massively multiplex chemical transcriptomics at single-cell resolution. Science 367, 45–51 (2020).
Dixit, A. et al. Perturb-seq: dissecting molecular circuits with scalable single-cell RNA profiling of pooled genetic screens. Cell 167, 1853–1866.e17 (2016).
Datlinger, P. et al. Ultra-high-throughput single-cell RNA sequencing and perturbation screening with combinatorial fluidic indexing. Nat. Methods 18, 635–642 (2021).
Papalexi, E. et al. Characterizing the molecular regulation of inhibitory immune checkpoints with multimodal single-cell screens. Nat. Genet. 53, 322–331 (2021).
Replogle, J. M. et al. Mapping information-rich genotype–phenotype landscapes with genome-scale Perturb-seq. Cell 185, 2559–2575.e28 (2022).
Wessels, H.-H. et al. Efficient combinatorial targeting of RNA transcripts in single cells with Cas13 RNA Perturb-seq. Nat. Methods 20, 86–94 (2023).
Ji, Y., Lotfollahi, M., Wolf, F. A. & Theis, F. J. Machine learning for perturbational single-cell omics. Cell Syst. 12, 522–537 (2021).
Duan, B. et al. Model-based understanding of single-cell CRISPR screening. Nat. Commun. 10, 2233 (2019).
Squair, J. W., Skinnider, M. A., Gautier, M., Foster, L. J. & Courtine, G. Prioritization of cell types responsive to biological perturbations in single-cell data with Augur. Nat. Protoc. 16, 3836–3873 (2021).
Skinnider, M. A. et al. Cell type prioritization in single-cell data. Nat. Biotechnol. 39, 30–34 (2021).
Burkhardt, D. B. et al. Quantifying the effect of experimental perturbations at single-cell resolution. Nat. Biotechnol. 39, 619–629 (2021).
Lotfollahi, M. et al. Learning interpretable cellular responses to complex perturbations in high-throughput screens. bioRxiv https://doi.org/10.1101/2021.04.14.439903 (2021).
Yuan, B. et al. CellBox: interpretable machine learning for perturbation biology with application to the design of cancer combination therapy. Cell Syst. 12, 128–140.e4 (2021).
Dimitrov, D. et al. Comparison of methods and resources for cell-cell communication inference from single-cell RNA-seq data. Nat. Commun. 13, 3224 (2022).
Jin, S. et al. Inference and analysis of cell-cell communication using CellChat. Nat. Commun. 12, 1088 (2021).
Efremova, M., Vento-Tormo, M., Teichmann, S. A. & Vento-Tormo, R. CellPhoneDB: inferring cell–cell communication from combined expression of multi-subunit ligand–receptor complexes. Nat. Protoc. 15, 1484–1506 (2020).
Cabello-Aguilar, S. et al. SingleCellSignalR: inference of intercellular networks from single-cell transcriptomics. Nucleic Acids Res. 48, e55 (2020).
Browaeys, R., Saelens, W. & Saeys, Y. NicheNet: modeling intercellular communication by linking ligands to target genes. Nat. Methods 17, 159–162 (2020).
Hu, Y., Peng, T., Gao, L. & Tan, K. CytoTalk: de novo construction of signal transduction networks using single-cell transcriptomic data. Sci. Adv. 7, eabf1356 (2021).
Isbel, L., Grand, R. S. & Schübeler, D. Generating specificity in genome regulation through transcription factor sensitivity to chromatin. Nat. Rev. Genet. 23, 728–740 (2022).
Cusanovich, D. A. et al. Multiplex single-cell profiling of chromatin accessibility by combinatorial cellular indexing. Science 348, 910–914 (2015).
Buenrostro, J. D. et al. Single-cell chromatin accessibility reveals principles of regulatory variation. Nature 523, 486–490 (2015).
Chen, H. et al. Assessment of computational methods for the analysis of single-cell ATAC-seq data. Genome Biol. 20, 241 (2019).
Fang, R. et al. Comprehensive analysis of single cell ATAC-seq data with SnapATAC. Nat. Commun. 12, 1337 (2021).
Ou, J. et al. ATACseqQC: a Bioconductor package for post-alignment quality assessment of ATAC-seq data. BMC Genomics 19, 169 (2018).
Amemiya, H. M., Kundaje, A. & Boyle, A. P. The ENCODE blacklist: identification of problematic regions of the genome. Sci. Rep. 9, 9354 (2019).
Thibodeau, A. et al. AMULET: a novel read count-based method for effective multiplet detection from single nucleus ATAC-seq data. Genome Biol. 22, 252 (2021).
Granja, J. M. et al. ArchR is a scalable software package for integrative single-cell chromatin accessibility analysis. Nat. Genet. 53, 403–411 (2021).
Bravo González-Blas, C. et al. cisTopic: cis-regulatory topic modeling on single-cell ATAC-seq data. Nat. Methods 16, 397–400 (2019).
Martens, L. D., Fischer, D. S., Theis, F. J. & Gagneur, J. Modeling fragment counts improves single-cell ATAC-seq analysis. bioRxiv https://doi.org/10.1101/2022.05.04.490536 (2022).
Stuart, T., Srivastava, A., Madad, S., Lareau, C. A. & Satija, R. Single-cell chromatin state analysis with Signac. Nat. Methods 18, 1333–1341 (2021).
Ashuach, T., Reidenbach, D. A., Gayoso, A. & Yosef, N. PeakVI: a deep generative model for single-cell chromatin accessibility analysis. Cell Rep. Methods 2, 100182 (2022).
Ashuach, T., Gabitto, M. I., Jordan, M. I. & Yosef, N. MultiVI: deep generative model for the integration of multi-modal data. Preprint at https://doi.org/10.1101/2021.08.20.457057.
Gontarz, P. et al. Comparison of differential accessibility analysis strategies for ATAC-seq data. Sci. Rep. 10, 10150 (2020).
McLean, C. Y. et al. GREAT improves functional interpretation of cis-regulatory regions. Nat. Biotechnol. 28, 495–501 (2010).
Sheffield, N. C. & Bock, C. LOLA: enrichment analysis for genomic region sets and regulatory elements in R and Bioconductor. Bioinformatics 32, 587–589 (2016).
Layer, R. M. et al. GIGGLE: a search engine for large-scale integrated genome analysis. Nat. Methods 15, 123–126 (2018).
Bredikhin, D., Kats, I. & Stegle, O. MUON: multimodal omics analysis framework. Genome Biol. 23, 42 (2022).
Pliner, H. A. et al. Cicero predicts cis-regulatory DNA interactions from single-cell chromatin accessibility data. Mol. Cell 71, 858–871.e8 (2018).
van Dijk, D. et al. Recovering gene interactions from single-cell data using data diffusion. Cell 174, 716–729.e27 (2018).
Schep, A. N., Wu, B., Buenrostro, J. D. & Greenleaf, W. J. chromVAR: inferring transcription-factor-associated accessibility from single-cell epigenomic data. Nat. Methods 14, 975–978 (2017).
Kartha, V. K. et al. Functional inference of gene regulation using single-cell multi-omics. Cell Genom. 2, 100166 (2022).
Cao, J. et al. Joint profiling of chromatin accessibility and gene expression in thousands of single cells. Science 361, 1380–1385 (2018).
Liu, L. et al. Deconvolution of single-cell multi-omics layers reveals regulatory heterogeneity. Nat. Commun. 10, 470 (2019).
Lynch, A.W., Theodoris, C.V., Long, H.W. et al. MIRA: joint regulatory modeling of multimodal expression and chromatin accessibility in single cells. Nat. Methods 19, 1097–1108 (2022).
Fleck, J. S. et al. Inferring and perturbing cell fate regulomes in human brain organoids. Nature https://doi.org/10.1038/s41586-022-05279-8 (2022).
Peterson, V. M. et al. Multiplexed quantification of proteins and transcripts in single cells. Nat. Biotechnol. 35, 936–939 (2017).
Zheng, Y., Jun, S.-H., Tian, Y., Florian, M. & Gottardo, R. Robust normalization and integration of single-cell protein expression across CITE-seq datasets. Preprint at https://doi.org/10.1101/2022.04.29.489989.
Mulè, M. P., Martins, A. J. & Tsang, J. S. Normalizing and denoising protein expression data from droplet-based single cell profiling. Nat. Commun. 13, 2099 (2022).
Gayoso, A. et al. Joint probabilistic modeling of single-cell multi-omic data with totalVI. Nat. Methods 18, 272–282 (2021).
Kim, H. J., Lin, Y., Geddes, T. A., Yang, J. Y. H. & Yang, P. CiteFuse enables multi-modal analysis of CITE-seq data. Bioinformatics 36, 4137–4143 (2020).
Sturm, G. et al. Scirpy: a Scanpy extension for analyzing single-cell T-cell receptor-sequencing data. Bioinformatics 36, 4817–4818 (2020).
Stephenson, E. et al. Single-cell multi-omics analysis of the immune response in COVID-19. Nat. Med. 27, 904–916 (2021).
Borcherding, N., Bormann, N. L. & Kraus, G. scRepertoire: an R-based toolkit for single-cell immune receptor analysis. F1000Res. 9, 47 (2020).
Bolotin, D. A. et al. MiXCR: software for comprehensive adaptive immunity profiling. Nat. Methods 12, 380–381 (2015).
Song, L. et al. TRUST4: immune repertoire reconstruction from bulk and single-cell RNA-seq data. Nat. Methods 18, 627–630 (2021).
Upadhyay, A. A. et al. BALDR: a computational pipeline for paired heavy and light chain immunoglobulin reconstruction in single-cell RNA-seq data. Genome Med. 10, 20 (2018).
Canzar, S., Neu, K. E., Tang, Q., Wilson, P. C. & Khan, A. A. BASIC: BCR assembly from single cells. Bioinformatics 33, 425–427 (2017).
Lindeman, I. et al. BraCeR: B-cell-receptor reconstruction and clonality inference from single-cell RNA-seq. Nat. Methods 15, 563–565 (2018).
Andreani, T. et al. Benchmarking computational methods for B-cell receptor reconstruction from single-cell RNA-seq data. NAR Genom. Bioinform. 4, lqac049 (2022).
Schuldt, N. J. & Binstadt, B. A. Dual TCR T cells: identity crisis or multitaskers? J. Immunol. 202, 637–644 (2019).
Polonsky, M., Chain, B. & Friedman, N. Clonal expansion under the microscope: studying lymphocyte activation and differentiation using live-cell imaging. Immunol. Cell Biol. 94, 242–249 (2016).
Greiff, V., Miho, E., Menzel, U. & Reddy, S. T. Bioinformatic and statistical analysis of adaptive immune repertoires. Trends Immunol. 36, 738–749 (2015). This article reviews the assumptions and scope of high-throughput immune repertoire data in the context of statistical analysis.
Springer, I., Tickotsky, N. & Louzoun, Y. Contribution of T cell receptor alpha and beta CDR3, MHC typing, V and J genes to peptide binding prediction. Front. Immunol. 12, 664514 (2021).
Setliff, I. et al. High-throughput mapping of B cell receptor sequences to antigen specificity. Cell 179, 1636–1646.e15 (2019).
Zhang, S.-Q. et al. High-throughput determination of the antigen specificities of T cell receptors in single cells. Nat. Biotechnol. https://doi.org/10.1038/nbt.4282 (2018).
Nazarov, V. I. et al. immunarch: bioinformatics analysis of T-cell and B-cell immune repertoires (immunarch, 2022).
Fleri, W. et al. The immune epitope database and analysis resource in epitope discovery and synthetic vaccine design. Front. Immunol. 8, 278 (2017).
Zhang, W. et al. PIRD: pan immune repertoire database. Bioinformatics 36, 897–903 (2020).
Shugay, M. et al. VDJdb: a curated database of T-cell receptor sequences with known antigen specificity. Nucleic Acids Res. 46, D419–D427 (2018).
Glanville, J. et al. Identifying specificity groups in the T cell receptor repertoire. Nature 547, 94–98 (2017).
Dash, P. et al. Quantifiable predictive features define epitope-specific T cell receptor repertoires. Nature 547, 89–93 (2017).
Chronister, W. D. et al. TCRMatch: predicting T-cell receptor specificity based on sequence similarity to previously characterized receptors. Front. Immunol. 12, 640725 (2021).
Zhang, Z., Xiong, D., Wang, X., Liu, H. & Wang, T. Mapping the functional landscape of T cell receptor repertoires by single-T cell transcriptomics. Nat. Methods 18, 92–99 (2021).
An, Y., Drost, F., Theis, F., Schubert, B. & Lotfollahi, M. Jointly learning T-cell receptor and transcriptomic information to decipher the immune response. Preprint at https://doi.org/10.1101/2021.06.24.449733.
Schattgen, S. A. et al. Integrating T cell receptor sequences and transcriptional profiles by clonotype neighbor graph analysis (CoNGA). Nat. Biotechnol. 40, 54–63 (2022).
Zhang, Z. et al. Interpreting the B-cell receptor repertoire with single-cell gene expression using Benisse. Nat. Mach. Intell. 4, 596–604 (2022).
Palla, G., Fischer, D. S., Regev, A. & Theis, F. J. Spatial components of molecular tissue biology. Nat. Biotechnol. 40, 308–318 (2022). This review highlights both the importance of different spatial profiling technologies and computational challenges for single-cell data resolved in space.
Dries, R. et al. Advances in spatial transcriptomic data analysis. Genome Res. 31, 1706–1718 (2021).
Asp, M., Bergenstråhle, J. & Lundeberg, J. Spatially resolved transcriptomes-next generation tools for tissue exploration. Bioessays 42, e1900221 (2020).
Tan, X., Su, A., Tran, M. & Nguyen, Q. SpaCell: integrating tissue morphology and spatial gene expression to predict disease cells. Bioinformatics 36, 2293–2294 (2020).
He, B. et al. Integrating spatial gene expression and breast tumour morphology via deep learning. Nat. Biomed. Eng. 4, 827–834 (2020).
Bergenstråhle, L. et al. Super-resolved spatial transcriptomics by deep data fusion. Nat. Biotechnol. 40, 476–479 (2022).
Velten, B. et al. Identifying temporal and spatial patterns of variation from multimodal data using MEFISTO. Nat. Methods 19, 179–186 (2022).
Liao, J., Lu, X., Shao, X., Zhu, L. & Fan, X. Uncovering an organ’s molecular architecture at single-cell resolution by spatially resolved transcriptomics. Trends Biotechnol. 39, 43–58 (2021).
Deng, Y. et al. Spatial profiling of chromatin accessibility in mouse and human tissues. Nature 609, 375–383 (2022).
Liu, S. et al. Spatial maps of T cell receptors and transcriptomes reveal distinct immune niches and interactions in the adaptive immune response. Immunity 55, 1940–1952.e5 (2022).
Lundberg, E. & Borner, G. H. H. Spatial proteomics: a powerful discovery tool for cell biology. Nat. Rev. Mol. Cell Biol. 20, 285–302 (2019).
Ståhl, P. L. et al. Visualization and analysis of gene expression in tissue sections by spatial transcriptomics. Science 353, 78–82 (2016).
Rodriques, S. G. et al. Slide-seq: a scalable technology for measuring genome-wide expression at high spatial resolution. Science 363, 1463–1467 (2019).
Lubeck, E., Coskun, A. F., Zhiyentayev, T., Ahmad, M. & Cai, L. Single-cell in situ RNA profiling by sequential hybridization. Nat. Methods 11, 360–361 (2014).
Chen, K. H., Boettiger, A. N., Moffitt, J. R., Wang, S. & Zhuang, X. RNA imaging. Spatially resolved, highly multiplexed RNA profiling in single cells. Science 348, aaa6090 (2015).
Ke, R. et al. In situ sequencing for RNA analysis in preserved tissue and cells. Nat. Methods 10, 857–860 (2013).
Crosetto, N., Bienko, M. & van Oudenaarden, A. Spatially resolved transcriptomics and beyond. Nat. Rev. Genet. 16, 57–66 (2015).
Zhang, M. et al. Spatial molecular profiling: platforms, applications and analysis tools. Brief. Bioinform. 22, bbaa145 (2021).
Zhuang, X. Spatially resolved single-cell genomics and transcriptomics by imaging. Nat. Methods 18, 18–22 (2021).
Palla, G. et al. Squidpy: a scalable framework for spatial omics analysis. Nat. Methods 19, 171–178 (2022).
Dries, R. et al. Giotto: a toolbox for integrative analysis and visualization of spatial expression data. Genome Biol. 22, 78 (2021).
Righelli, D. et al. SpatialExperiment: infrastructure for spatially resolved transcriptomics data in R using Bioconductor. Bioinformatics 38, 3128–3131 (2022).
Kleshchevnikov, V. et al. Cell2location maps fine-grained cell types in spatial transcriptomics. Nat. Biotechnol. 40, 661–671 (2022).
Dong, R. & Yuan, G.-C. SpatialDWLS: accurate deconvolution of spatial transcriptomic data. Genome Biol. 22, 145 (2021).
Cable, D. M. et al. Robust decomposition of cell type mixtures in spatial transcriptomics. Nat. Biotechnol. 40, 517–526 (2022).
Li, B. et al. Benchmarking spatial and single-cell transcriptomics integration methods for transcript distribution prediction and cell type deconvolution. Nat. Methods 19, 662–670 (2022).
Yan, L. & Sun, X. Benchmarking and integration of methods for deconvoluting spatial transcriptomic data. Bioinformatics 39, btac805 (2023).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
McQuin, C. et al. CellProfiler 3.0: Next-generation image processing for biology. PLoS Biol. 16, e2005970 (2018).
Berg, S. et al. ilastik: interactive machine learning for (bio)image analysis. Nat. Methods 16, 1226–1232 (2019).
Stringer, C., Wang, T., Michaelos, M. & Pachitariu, M. Cellpose: a generalist algorithm for cellular segmentation. Nat. Methods 18, 100–106 (2021).
Park, J. et al. Cell segmentation-free inference of cell types from in situ transcriptomics data. Nat. Commun. 12, 3545 (2021).
Petukhov, V. et al. Cell segmentation in imaging-based spatial transcriptomics. Nat. Biotechnol. 40, 345–354 (2022).
Kuemmerle, L. B. et al. Probe set selection for targeted spatial transcriptomics. Preprint at https://doi.org/10.1101/2022.08.16.504115.
Biancalani, T. et al. Deep learning and alignment of spatially resolved single-cell transcriptomes with Tangram. Nat. Methods 18, 1352–1362 (2021).
Lopez, R. et al. A joint model of unpaired data from scRNA-seq and spatial transcriptomics for imputing missing gene expression measurements. arXiv https://doi.org/10.48550/arXiv.1905.02269 (2019).
Abdelaal, T., Mourragui, S., Mahfouz, A. & Reinders, M. J. T. SpaGE: spatial gene enhancement using scRNA-seq. Nucleic Acids Res. 48, e107 (2020).
Zhao, E. et al. Spatial transcriptomics at subspot resolution with BayesSpace. Nat. Biotechnol. 39, 1375–1384 (2021).
Pham, D. et al. stLearn: integrating spatial location, tissue morphology and gene expression to find cell types, cell-cell interactions and spatial trajectories within undissociated tissues. bioRxiv https://doi.org/10.1101/2020.05.31.125658 (2020).
Hu, J. et al. SpaGCN: integrating gene expression, spatial location and histology to identify spatial domains and spatially variable genes by graph convolutional network. Nat. Methods 18, 1342–1351 (2021).
Daly, A. C., Geras, K. J. & Bonneau, R. A. A convolutional neural network for common coordinate registration of high-resolution histology images. Bioinformatics 37, 4216–4226 (2021).
Andersson, A. et al. A landmark-based common coordinate framework for spatial transcriptomics data. bioRxiv https://doi.org/10.1101/2021.11.11.468178 (2021).
Rood, J. E. et al. Toward a common coordinate framework for the human body. Cell 179, 1455–1467 (2019).
Sun, S., Zhu, J. & Zhou, X. Statistical analysis of spatial expression patterns for spatially resolved transcriptomic studies. Nat. Methods 17, 193–200 (2020).
Svensson, V., Teichmann, S. A. & Stegle, O. SpatialDE: identification of spatially variable genes. Nat. Methods 15, 343–346 (2018).
Anderson, A. & Lundeberg, J. sepal: identifying transcript profiles with spatial patterns by diffusion-based modeling. Bioinformatics 37, 2644–2650 (2021).
Fischer, D. S., Schaar, A. C. & Theis, F. J. Modeling intercellular communication in tissues using spatial graphs of cells. Nat. Biotechnol. https://doi.org/10.1038/s41587-022-01467-z (2022).
Yuan, Y. & Bar-Joseph, Z. GCNG: graph convolutional networks for inferring gene interaction from spatial transcriptomics data. Genome Biol. 21, 300 (2020).
Tanevski, J., Flores, R. O. R., Gabor, A., Schapiro, D. & Saez-Rodriguez, J. Explainable multiview framework for dissecting spatial relationships from highly multiplexed data. Genome Biol. 23, 97 (2022).
Cang, Z. & Nie, Q. Inferring spatial and signaling relationships between cells from single cell transcriptomic data. Nat. Commun. 11, 2084 (2020).
Arnol, D., Schapiro, D., Bodenmiller, B., Saez-Rodriguez, J. & Stegle, O. Modeling cell-cell interactions from spatial molecular data with spatial variance component analysis. Cell Rep. 29, 202–211.e6 (2019).
Kamimoto, K., Stringa, B., Hoffmann, C.M. et al. Dissecting cell identity via network inference and in silico gene perturbation. Nature 614, 742–751 (2023).
González-Blas, C. B. et al. SCENIC+: single-cell multiomic inference of enhancers and gene regulatory networks. bioRxiv https://doi.org/10.1101/2022.08.19.504505 (2022).
Brunner, A.-D. et al. Ultra-high sensitivity mass spectrometry quantifies single-cell proteome changes upon perturbation. Mol. Syst. Biol. 18, e10798 (2022).
Del Rossi, N., Chen, J. G., Yuan, G.-C. & Dries, R. Analyzing spatial transcriptomics data using Giotto. Curr. Protoc. 2, e405 (2022).
Argelaguet, R., Cuomo, A. S. E., Stegle, O. & Marioni, J. C. Computational principles and challenges in single-cell data integration. Nat. Biotechnol. 39, 1202–1215 (2021).
Mimitou, E. P. et al. Scalable, multimodal profiling of chromatin accessibility, gene expression and protein levels in single cells. Nat. Biotechnol. 39, 1246–1258 (2021). A review on multimodal data integration that further categorizes methods into different types of data integration task as for example used here.
Lake, B. B. et al. Integrative single-cell analysis of transcriptional and epigenetic states in the human adult brain. Nat. Biotechnol. 36, 70–80 (2018).
Argelaguet, R. et al. MOFA+: a statistical framework for comprehensive integration of multi-modal single-cell data. Genome Biol. 21, 111 (2020).
Demetci, P., Santorella, R., Sandstede, B., Noble, W. S. & Singh, R. SCOT: single-cell multi-omics alignment with optimal transport. J. Comput. Biol. 29, 3–18 (2022).
Cao, K., Bai, X., Hong, Y. & Wan, L. Unsupervised topological alignment for single-cell multi-omics integration. Bioinformatics 36, i48–i56 (2020).
Cao, ZJ., Gao, G. Multi-omics single-cell data integration and regulatory inference with graph-linked embedding. Nat. Biotechnol. 40, 1458–1466 (2022).
Lance, C. et al. Multimodal single cell data integration challenge: results and lessons learned. in Proceedings of the NeurIPS 2021 Competitions and Demonstrations Track (eds Kiela, D., Ciccone, M. & Caputo, B.) vol. 176 162–176 (PMLR, 2022).
Ghazanfar, S., Guibentif, C. & Marioni, J. C. StabMap: mosaic single cell data integration using non-overlapping features. bioRxiv https://doi.org/10.1101/2022.02.24.481823 (2022).
Lotfollahi, M., Litinetskaya, A. & Theis, F. J. Multigrate: single-cell multi-omic data integration. bioRxiv https://doi.org/10.1101/2022.03.16.484643 (2022).
Barshan, E., Ghodsi, A., Azimifar, Z. & Zolghadri Jahromi, M. Supervised principal component analysis: visualization, classification and regression on subspaces and submanifolds. Pattern Recognit. 44, 1357–1371 (2011).
Hao, Y. et al. Dictionary learning for integrative, multimodal, and scalable single-cell analysis. bioRxiv https://doi.org/10.1101/2022.02.24.481684 (2022).
Acknowledgements
The authors acknowledge Y. Chen for editing the single-cell RNA-sequencing discussions, Y. Ji for editing the perturbation modelling discussions, A. McKenna for editing the lineage tracing discussions, C. N. Talavera Lopez for providing helpful suggestions for the adaptive immune receptor repertoire discussion, L. B. Kuemmerle for editing the spatial omics discussions, and all members of the Theis group for reviews and helpful discussion. This work was supported by the German Federal Ministry of Education and Research (BMBF) under grant no. 01IS18053A, by the Bavarian Ministry of Science and the Arts in the framework of the Bavarian Research Association “ForInter” (Interaction of human brain cells), by the Wellcome Trust grant 108413/A/15/D and by the Helmholtz Association’s Initiative and Networking Fund through Helmholtz AI (grant number: ZT-I-PF-5-01). Main author list, individual acknowledgements: F.D. is supported by the Helmholtz Association under the joint research school Munich School for Data Science and by the Joachim Herz Stiftung. F.C. acknowledges support from a German Research Foundation (DFG) (SFB-TRR 338/1 2021-452881907), Bavarian Ministry of Science and the Arts in the framework of the Bavarian Research Association “ForInter” (Interaction of human brain cells) and by the Deutsche Forschungsgemeinschaft. A.C.S., F.C. and L.Z. acknowledge support from the Bavarian Ministry of Science and the Arts in the framework of the Bavarian Research Association “ForInter” (Interaction of human brain cells). C.L. is supported by the Helmholtz Association under the joint research school Munich School for Data Science. Single-cell Best Practices Consortium, individual acknowledgements: G.P. and L.D. are supported by the Joachim Herz Stiftung. G.P. is supported by the Helmholtz Association under the joint research school Munich School for Data Science. R.P. acknowledges funding from US NIH (R01 HG009937) and US National Science Foundation (CCF-1750472, and CNS-1763680). L. Hetzel and L.D.M. are supported by the Helmholtz Association under the joint research school Munich School for Data Science. B.S. acknowledges funding from (DFG, German Research Foundation) Projektnummer 490846870-TRR355/1 TPZ02.
Author information
Authors and Affiliations
Consortia
Contributions
Main author list: A.C.S., L. Heumos and F.J.T. conceived the project. L. Heumos and A.C.S. contributed equally and have the right to list their name first in their curriculum vitae. A.C.S., L. Heumos, C.L. and F.D. wrote the manuscript. L.Z. and M.D.L. provided expertise for the discussion on transcriptomics; C.L. on chromatin accessibility; D.C.S. on surface protein expression; F.D., J.H. and F.C. on adaptive immune receptor repertoire analysis; and A.L. and F.C. on multimodal data integration. F.J.T. and H.B.S. supervised the work. Single-cell Best Practices Consortium: A.F., H.A., I.L.I., L.D., L.S., M.B., M.L., P.W., S.H.-z., Z.P., M.G.J., A.S., H.S., D.H., E.D., J.O., I.V., D.D., R.P., C.L.M., J.S.-R., J.H., P.B.M. and M.N. provided expertise for the discussion on transcriptomics; L.D.M. and I.L.I. on chromatin accessibility; C.R.-S. on surface protein expression; B.S. on adaptive immune receptor repertoire analysis; and G.P., L. Hetzel, J.T. and J.S.-R. on single-cell data resolved in space. M.A. contributed to the figure design. All authors read, edited and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
Main author list: M.D.L. has received speaker’s honoraria from Pfizer and Janssen, and received consulting fees from Chan-Zuckerberg Initiative. F.J.T. consults for Immunai Inc., Singularity Bio B.V., CytoReason Ltd and Omniscope Ltd, and has ownership interest in Dermagnostix GmbH and Cellarity. M.G.J. consults for and has ownership interests in Vevo Therapeutics. L. Heumos has received speaker’s honorarium from Vesalius Therapeutics. Single-Cell Best Practices Consortium: M.G.J. consults for and has ownership interests in Vevo Therapeutics. R.P. is co-founder of Ocean Genomics, Inc. The other authors declare no competing interests.
Peer review
Peer review information
Nature Reviews Genetics thanks Francesca Finotello, Jong-Eun Park and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Single-Cell Best Practices online book: https://sc-best-practices.org
Glossary
- Adaptive immune receptor
-
(AIR). Transmembrane complex of proteins expressed on T and B cells that is key for the recognition of potential hazardous antigens and pathogens invading the body.
- Ambient RNA
-
mRNA counts that originate from other lysed cells in the input solution and do not belong to the cell captured in the droplet itself.
- Antibody-derived tags
-
(ADTs). Antibodies (also known as soluble immunoglobulins) are Y-shaped proteins used by the immune system to identify and neutralize pathogens by recognizing antigens. ADTs are directly conjugated DNA-barcode oligonucleotides that can be used to recover expressed surface proteins.
- Antigens
-
Substances recognized as non-self that induce an immune response and lead to the production of antibodies.
- Barcodes
-
Unique known nucleic acid sequences of fixed length used to label individual cells to enable tracking through space and time.
- Batch effects
-
Confounding effects that result from technical differences in data generation across different batches, such as samples obtained through different experimental set-ups or from different laboratories.
- CDR3
-
Whereas complementarity-determining region 1 (CDR1) and CDR2 are encoded in the germline V genes, CDR3 loops are assembled from V(D)J segments, giving rise to the variability of adaptive immune receptors.
- Cell fate
-
A cell’s final cell type that is established by corresponding, specific transcriptional programmes.
- Cell–cell communication
-
Interactions of cells through secreted ligands and plasma membrane receptors, secreted enzymes, extracellular matrix proteins or cell–cell adhesion proteins and gap junctions.
- Cell-type deconvolution
-
Decomposing the cell-type composition of individual barcode regions based on a reference data set to obtain abundances or proportions of individual cells within a barcode region.
- Cell segmentation
-
Processing of microscopic image domains into segments that represent individual cells.
- Chain pairing
-
Assignment of cells to V(D)J chain types such as orphans, single pair, extra VJ/VDJ or multichains.
- Cis-regulatory elements
-
(CREs). Regions of non-coding DNA — such as promoters, enhancers and silencers — that control the transcription of nearby genes.
- Clonotype
-
Collection of T or B cells that descended from an antecedent cell, have the same adaptive immune receptors and henceforth recognize the same epitopes.
- Compositional data
-
Comprises multi-dimensional data points (for example, cell-type composition) in which each component (or part) carries only proportional or relative abundance information about some whole.
- Confounding sources of variation
-
Technical artefacts that arise from library preparation and sequencing, and biological confounders such as cell cycle status, which cause systematic bias and may distort biological findings.
- Differential gene expression
-
(DGE). The inference of statistically significant differences in expression between groups such as healthy and diseased.
- Epitopes
-
The parts of antigens that are recognized by antibodies, B cells or T cells to potentially stimulate immune responses.
- Gene set enrichment
-
Grouping genes with shared characteristics together and testing for over-representation.
- Graph neural networks
-
A deep-learning approach to do inference on input data represented in the form of a graph. For example, in spatial transcriptomics, cells are typically represented as nodes in graphs obtained through spatial proximity.
- Highly variable genes
-
A measure to identify genes that vary in terms of gene expression across all cells present in the data set.
- K nearest-neighbours graph
-
(KNN graph). A computational data structure in which cells are represented as nodes in a graph. Based on distance metrics such as the Euclidean distance on a principal-component reduced expression, cells are connected to their K most similar cells. K is commonly set to be between 5 and 100 depending on the data set.
- Latent semantic indexing
-
(LSI). A dimension reduction method that uses term frequency inverse document frequency transformation (TFIDF) followed by singular value decomposition (SVD).
- Lineage tracing
-
Tracking physiological or pathological changes by exogenous or endogenous cell markers such as DNA mutations.
- Major histocompatibility complex
-
(MHC). Surface proteins that display or ‘present’ small peptides (epitopes) on the cell surface for T and B cells to potentially react to. Presented endogenous self-antigens prevent the immune system from targeting its own cells, whereas presented pathogen-derived peptides alarm nearby immune cells.
- Nucleosome signal
-
The ratio of long fragments resulting from one or multiple histones bound between the Tn5 transposition sites and short nucleosome-free fragments; the ratio is small in high-quality single-cell assay for transposase-accessible chromatin sequencing (scATAC-seq) data.
- Optimal transport
-
Mathematical framework to estimate the optimal transport plan of mass between two (discrete) distributions.
- Phase portrait
-
For any given gene, the phase portrait visualizes splicing kinetics as a parametric curve (with time as a parameter).
- Pseudobulks
-
Aggregated cells within a biological replicate whereby the data from every single cell is combined via sum or mean of counts into a single pseudo-sample to resemble a bulk RNA experiment.
- Pseudoreplication
-
Also known as subsampling. Pseudoreplication occurs when replicates are not statistically independent, but are treated as if they were, such as cell samples from a single individual.
- Reference mapping
-
The process of leveraging and transferring information from a reference data set to a query.
- RNA velocity
-
Ratios of spliced mRNA, unspliced mRNA and mRNA degradation. Positive ratios (velocities) indicate recent increases in unspliced transcripts followed by upregulation of spliced transcripts. Negative velocities indicate downregulation. Examining velocities across genes can provide insight into future states of individual cells.
- Scaling
-
Normalization of gene expression levels that scales gene counts to zero mean and unit variance.
- Somatic hypermutation
-
Mechanism of B cell receptors to allow the immune system to adapt its response to unseen threats. Somatic hypermutation is triggered when B cells engage antigens, which results in the introduction of point mutations in the variable regions of the V(D)J genes. Cells harbouring mutagenized antibodies with a high affinity for the antigen proliferate preferentially (known as affinity maturation).
- Spatially variable genes
-
(SVGs). Genes with variable expression levels between individual locations in the spatial transcriptomics data set.
- Spectratyping
-
Measuring the heterogeneity of complementarity-determining region 3 (CDR3) regions by their length diversity across different cell types or conditions.
- Trajectory inference
-
Also known as pseudotime analysis. Ordering of cells along a trajectory based on gene expression similarity.
- Transcription factor motif
-
(TF motif). DNA sequence pattern that is specifically recognized by a sequence-specific TF. It is commonly represented as a logo diagram representing the most informative DNA positions by height.
- Variational autoencoders
-
A generative artificial neural network architecture that allows for statistical inference. Input data are sampled from a parameterized distribution (prior), and an encoder and decoder are trained jointly to minimize the reconstruction error between the updated prior probability (posterior) and its parametric approximation (variational posterior).
- V(D)J recombination
-
Somatic recombination in developing lymphocytes whereby variable (V), diversity (D) and joining (J) segments are randomly selected and joined to form the V region of a full-length receptor.
- V(D)J sequencing
-
Determination of protein sequence of the adaptive immune receptor (AIR) for both chains, from which the variable (V), diversity (D), joining (J) and constant (C) sequences are determined in addition to the complementarity-determining region (CDR) sequences.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Heumos, L., Schaar, A.C., Lance, C. et al. Best practices for single-cell analysis across modalities. Nat Rev Genet 24, 550–572 (2023). https://doi.org/10.1038/s41576-023-00586-w
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41576-023-00586-w
This article is cited by
-
Aligned cross-modal integration and regulatory heterogeneity characterization of single-cell multiomic data with deep contrastive learning
Genome Medicine (2026)
-
Mutation-informed gene pairs to predict melanoma metastasis
Cell Communication and Signaling (2026)
-
omnideconv: a unifying framework for using and benchmarking single-cell-informed deconvolution of bulk RNA-seq data
Genome Biology (2026)
-
Srnc: semi-supervised learning for robust novel cell-type identification in single cell RNA sequencing data
BMC Bioinformatics (2026)
-
Single-cell profiling defines the cellular landscape of the urinary bladder: a scoping review
European Journal of Medical Research (2026)