Abstract
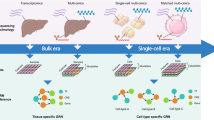
Gene regulatory networks (GRNs) explain how the genome controls cellular behaviour and tissue morphogenesis, serving to connect molecular mechanism to functional output. Single-cell technologies now provide descriptions of these networks with unprecedented detail, but this advance has also revealed gene regulatory systems that are too complex for our existing conceptual frameworks. GRNs, which should provide mechanistic explanations, are increasingly reduced to statistical correlations — ‘hairballs’ that fail to capture molecular causation. Here, we explore why this dilemma exists and propose a path forward. We argue that methods in ‘representation learning’ can be used to model GRNs, without needing to capture every molecular detail. For this framework, we advocate three linked principles: models must be inherently mechanistic, with structures grounded in cellular and evolutionary biology; molecular principles and constraints must be used to reduce the solution space for learning GRN models; and more sophisticated forms of experimental perturbation and synthetic biological engineering are needed to train models and test predictions. By reimagining GRNs through these principles, we can bridge the gap from data abundance to new conceptual understanding.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Nüsslein-Volhard, C., Wieschaus, E. & Kluding, H. Mutations affecting the pattern of the larval cuticle Indrosophila melanogaster: I. Zygotic loci on the second chromosome. Wilhelm Roux Arch. Dev. Biol. 193, 267–282 (1984).
Jürgens, G., Wieschaus, E., Nüsslein-Volhard, C. & Kluding, H. Mutations affecting the pattern of the larval cuticle Indrosophila melanogaster: II. Zygotic loci on the third chromosome. Wilhelm Roux Arch. Dev. Biol. 193, 283–295 (1984).
Wieschaus, E., Nüsslein-Volhard, C. & Jürgens, G. Mutations affecting the pattern of the larval cuticle Indrosophila melanogaster: III. Zygotic loci on the X-chromosome and fourth chromosome. Wilhelm Roux Arch. Dev. Biol. 193, 296–307 (1984).
Rood, J. E., Hupalowska, A. & Regev, A. Toward a foundation model of causal cell and tissue biology with a perturbation cell and tissue atlas. Cell 187, 4520–4545 (2024).
DiFrisco, J. & Jaeger, J. Genetic causation in complex regulatory systems: an integrative dynamic perspective. BioEssays 42, e1900226 (2020).
Davidson, E. H. in Genomic Regulatory Systems (ed. Davidson, E. H.) Ch. 1 (Academic, 2001).
Davidson, E. H. & Peter, I. S. in Genomic Control Process (eds Davidson, E. H. & Peter, I. S.) Ch. 2 (Academic, 2015).
Davidson, E. H. & Erwin, D. H. Gene regulatory networks and the evolution of animal body plans. Science 311, 796–800 (2006). This paper has established the foundational GRN concept within an evolutionary context and hierarchical organization.
Davidson, E. H. & Erwin, D. H. Gene regulatory networks and the evolution of animal body plans. Science 311, 796–800 (2006).
Perez-Carrasco, R. et al. Combining a toggle switch and a repressilator within the AC-DC circuit generates distinct dynamical behaviors. Cell Syst. 6, 521–530.e3 (2018).
Pourquié, O. & Goldbeter, A. Segmentation clock: insights from computational models. Curr. Biol. 13, R632–R634 (2003).
Hirata, H. et al. Oscillatory expression of the bHLH factor Hes1 regulated by a negative feedback loop. Science 298, 840–843 (2002).
Jaeger, J. The gap gene network. Cell. Mol. Life Sci. 68, 243–74 (2011).
Sick, S., Reinker, S., Timmer, J. & Schlake, T. WNT and DKK determine hair follicle spacing through a reaction-diffusion mechanism. Science 314, 1447–1450 (2006).
Briscoe, J. & Small, S. Morphogen rules: design principles of gradient-mediated embryo patterning. Development 142, 3996–4009 (2015).
Raspopovic, J., Marcon, L., Russo, L. & Sharpe, J. Modeling digits. Digit patterning is controlled by a Bmp-Sox9-Wnt Turing network modulated by morphogen gradients. Science 345, 566–570 (2014).
Kondo, S. & Asal, R. A reaction-diffusion wave on the skin of the marine angelfish Pomacanthus. Nature 376, 765–768 (1995).
Collier, J. R., Monk, N. A., Maini, P. K. & Lewis, J. H. Pattern formation by lateral inhibition with feedback: a mathematical model of delta-notch intercellular signalling. J. Theor. Biol. 183, 429–446 (1996).
Li, C., Hong, T. & Nie, Q. Quantifying the landscape and kinetic paths for epithelial-mesenchymal transition from a core circuit. Phys. Chem. Chem. Phys. 18, 17949–17956 (2016).
Tapscott, S. J. The circuitry of a master switch: Myod and the regulation of skeletal muscle gene transcription. Development 132, 2685–2695 (2005).
Davidson, E. H. et al. A provisional regulatory gene network for specification of endomesoderm in the sea urchin embryo. Dev. Biol. 246, 162–190 (2002).
Delile, J. et al. Single cell transcriptomics reveals spatial and temporal dynamics of gene expression in the developing mouse spinal cord. Development 146, dev173807 (2019).
Rayon, T., Maizels, R. J., Barrington, C. & Briscoe, J. Single-cell transcriptome profiling of the human developing spinal cord reveals a conserved genetic programme with human-specific features. Development 148, dev199711 (2021).
Nishi, Y. et al. A direct fate exclusion mechanism by Sonic hedgehog-regulated transcriptional repressors. Development 142, 3286–3293 (2015).
Peterson, K. A. et al. Neural-specific Sox2 input and differential Gli-binding affinity provide context and positional information in Shh-directed neural patterning. Genes Dev. 26, 2802–2816 (2012).
Badia-I-Mompel, P. et al. Gene regulatory network inference in the era of single-cell multi-omics. Nat. Rev. Genet. 24, 739–754 (2023).
Kamimoto, K. et al. Dissecting cell identity via network inference and in silico gene perturbation. Nature 614, 742–751 (2023).
Wang, L. et al. Dictys: dynamic gene regulatory network dissects developmental continuum with single-cell multiomics. Nat. Methods 20, 1368–1378 (2023).
González-Blas, C. B. et al. SCENIC+: single-cell multiomic inference of enhancers and gene regulatory networks. Nat. Methods 20, 1355–1367 (2023). This paper is a key example of modern GRN modelling from multiomics single-cell data.
Scholkopf, B. et al. Toward causal representation learning. Proc. IEEE 109, 612–634 (2021). This paper introduces the methods and applications of the field of causal representation learning.
Davidson, E. H. & Levine, M. S. Properties of developmental gene regulatory networks. Proc. Natl Acad. Sci. USA 105, 20063–20066 (2008).
Davidson, E. H. et al. A genomic regulatory network for development. Science 295, 1669–1678 (2002).
Balaskas, N. et al. Gene regulatory logic for reading the Sonic Hedgehog signaling gradient in the vertebrate neural tube. Cell 148, 273–84 (2012).
Huynh-Thu, V. A., Irrthum, A., Wehenkel, L. & Geurts, P. Inferring regulatory networks from expression data using tree-based methods. PLoS ONE 5, e12776 (2010).
Meyer, P. E., Kontos, K., Lafitte, F. & Bontempi, G. Information-theoretic inference of large transcriptional regulatory networks. Eur. J. Bioinform. Syst. Biol. 2007, 79879 (2007).
Maizels, R. J., Snell, D. M. & Briscoe, J. Reconstructing developmental trajectories using latent dynamical systems and time-resolved transcriptomics. Cell Syst. 15, 411–424.e9 (2024). This paper combines time-resolved single-cell transcriptomics and deep learning models to reconstruct cell differentiation trajectories and cell fate decisions.
Cao, J. et al. The single-cell transcriptional landscape of mammalian organogenesis. Nature 566, 496–502 (2019).
Mittnenzweig, M. et al. A single-embryo, single-cell time-resolved model for mouse gastrulation. Cell 184, 2825–2842.e22 (2021).
Pijuan-Sala, B. et al. A single-cell molecular map of mouse gastrulation and early organogenesis. Nature 566, 490–495 (2019).
Sanchez-Castillo, M., Blanco, D., Tienda-Luna, I. M., Carrion, M. C. & Huang, Y. A Bayesian framework for the inference of gene regulatory networks from time and pseudo-time series data. Bioinformatics 34, 964–970 (2018).
Matsumoto, H. et al. SCODE: an efficient regulatory network inference algorithm from single-cell RNA-Seq during differentiation. Bioinformatics 33, 2314–2321 (2017).
Specht, A. T. & Li, J. J. LEAP: constructing gene co-expression networks for single-cell RNA-sequencing data using pseudotime ordering. Bioinformatics 33, 764–766 (2017).
Qiu, X. et al. Inferring causal gene regulatory networks from coupled single-cell expression dynamics using scribe. Cell Syst. 10, 265–274.e11 (2020).
Pratapa, A., Jalihal, A. P., Law, J. N., Bharadwaj, A. & Murali, T. M. Benchmarking algorithms for gene regulatory network inference from single-cell transcriptomic data. Nat. Methods 17, 147–154 (2020).
Chen, S. & Mar, J. C. Evaluating methods of inferring gene regulatory networks highlights their lack of performance for single cell gene expression data. BMC Bioinformatics 19, 232 (2018).
Kernfeld, E., Keener, R., Cahan, P. & Battle, A. Transcriptome data are insufficient to control false discoveries in regulatory network inference. Cell Syst. 15, 709–724.e13 (2024).
Wanniarachchi, D. V., Viswakula, S. & Wickramasuriya, A. M. The evaluation of transcription factor binding site prediction tools in human and Arabidopsis genomes. BMC Bioinformatics 25, 371 (2024).
Keilwagen, J., Posch, S. & Grau, J. Accurate prediction of cell type-specific transcription factor binding. Genome Biol. 20, 9 (2019).
Zandvakili, A., Campbell, I., Gutzwiller, L. M., Weirauch, M. T. & Gebelein, B. Degenerate Pax2 and senseless binding motifs improve detection of low-affinity sites required for enhancer specificity. PLoS Genet 14, e1007289 (2018).
Lettice, L. A. et al. A long-range Shh enhancer regulates expression in the developing limb and fin and is associated with preaxial polydactyly. Hum. Mol. Genet. 12, 1725–1735 (2003).
Badia-i Mompel, P. et al. Comparison and evaluation of methods to infer gene regulatory networks from multimodal single-cell data. Preprint at bioRxiv https://doi.org/10.1101/2024.12.20.629764 (2025). This recent review identifies shortcomings and challenges of GRN modelling from multiome data.
Maizels, R. J. A dynamical perspective: moving towards mechanism in single-cell transcriptomics. Philos. Trans. R. Soc. Lond. B Biol. Sci. 379, 20230049 (2024).
Cotterell, J. & Sharpe, J. An atlas of gene regulatory networks reveals multiple three-gene mechanisms for interpreting morphogen gradients. Mol. Syst. Biol. 6, 425 (2010). This work demonstrates dynamical network structures that produce identical developmental patterns.
Isalan, M. Gene networks and liar paradoxes. BioEssays 31, 1110–1115 (2009).
Wieland, F.-G., Hauber, A. L., Rosenblatt, M., Tönsing, C. & Timmer, J. On structural and practical identifiability. Curr. Opin. Syst. Biol. 25, 60–69 (2021).
Raue, A. et al. Structural and practical identifiability analysis of partially observed dynamical models by exploiting the profile likelihood. Bioinformatics 25, 1923–1929 (2009).
Chis, O.-T., Villaverde, A. F., Banga, J. R. & Balsa-Canto, E. On the relationship between sloppiness and identifiability. Math. Biosci. 282, 147–161 (2016).
Gutenkunst, R. N. et al. Universally sloppy parameter sensitivities in systems biology models. PLoS Comput. Biol. 3, e189 (2007).
Transtrum, M. K. et al. Sloppiness and emergent theories in physics, biology, and beyond. J. Chem. Phys. 143, 010901 (2015).
Gábor, A., Hangos, K. M., Banga, J. R. et al. Reaction network realizations of rational biochemical systems and their structural properties. J. Math. Chem. 53, 1657–1686 (2015).
Babtie, A. C., Kirk, P. & Stumpf, M. P. H. Topological sensitivity analysis for systems biology. Proc. Natl Acad. Sci. USA 111, 18507–18512 (2014).
Schaerli, Y. et al. Synthetic circuits reveal how mechanisms of gene regulatory networks constrain evolution. Mol. Syst. Biol. 14, e8102 (2018).
Otero-Muras, I., Perez-Carrasco, R., Banga, J. R. & Barnes, C. P. Automated design of gene circuits with optimal mushroom-bifurcation behavior. iScience 26, 106836 (2023).
Jiménez, A., Cotterell, J., Munteanu, A. & Sharpe, J. A spectrum of modularity in multi-functional gene circuits. Mol. Syst. Biol. 13, 925 (2017).
Bhalla, U. S. & Iyengar, R. Emergent properties of networks of biological signaling pathways. Science 283, 381–387 (1999). This paper provides an example of emergent functionality in biological networks.
Yurchenko, S. B. Is information the other face of causation in biological systems? BioSystems 229, 104925 (2023).
Artime, O. & De Domenico, M. From the origin of life to pandemics: emergent phenomena in complex systems. Philos. Trans. R. Soc. A 380, 20200410 (2022).
Ellis, G. F. Top-down causation and emergence: some comments on mechanisms. Interface Focus 2, 126–140 (2012).
Klein, B. & Hoel, E. The emergence of informative higher scales in complex networks. Complexity https://doi.org/10.1155/2020/8932526 (2020).
Hoel, E. When the map is better than the territory. Entropy 19, 188 (2017).
Zhang, P. et al. Negative cross-talk between hematopoietic regulators: GATA proteins repress PU.1. Proc. Natl Acad. Sci. USA 96, 8705–8710 (1999).
Levy, C., Khaled, M. & Fisher, D. E. MITF: master regulator of melanocyte development and melanoma oncogene. Trends Mol. Med. 12, 406–414 (2006).
Frith, T. J., Briscoe, J. & Boezio, G. L. in Current Topics in Developmental Biology Vol. 159 (ed. Mallo, M.) Ch. 5 (Elsevier, 2024).
Kassouf, M. T. et al. The α-globin super-enhancer acts in an orientation-dependent manner. Nat. Commun. 16, 1033 (2025).
Small, S., Blair, A. & Levine, M. Regulation of even-skipped stripe 2 in the Drosophila embryo. EMBO J. 11, 4047–4057 (1992).
Panne, D., Maniatis, T. & Harrison, S. C. An atomic model of the interferon-β enhanceosome. Cell 129, 1111–1123 (2007).
Kmiecik, S. et al. Coarse-grained protein models and their applications. Chem. Rev. 116, 7898–7936 (2016).
Ingólfsson, H. I. et al. The power of coarse graining in biomolecular simulations. Wiley Interdiscip. Rev. Comput. Mol. Sci. 4, 225–248 (2014).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Marr, D. Vision: A Computational Approach (MIT Press, 1982).
Marr, D. C. & Poggio, T. From Understanding Computation to Understanding Neural Circuitry (MIT Artificial Intelligence Laboratory, 1976).
Lazebnik, Y. Can a biologist fix a radio? — or, what I learned while studying apoptosis. Cancer Cell 2, 179–182 (2002).
Pezzotta, A. & Briscoe, J. Optimal control of gene regulatory networks for morphogen-driven tissue patterning. Cell Syst. 14, 940–952.e11 (2023).
Tkačik, G. & Wolde, P. R. T. Information processing in biochemical networks. Annu. Rev. Biophys. 54, 249–274 (2025).
Peter, I. S. & Davidson, E. H. Evolution of gene regulatory networks controlling body plan development. Cell 144, 970–985 (2011).
Hatleberg, W. L. & Hinman, V. F. in Current Topics in Developmental Biology Vol. 141 (ed. Gilbert, S. F.) Ch. 2 (Elsevier, 2021). This paper explores modularity and hierarchy in GRNs, which can help understand their developmental function.
Alon, U. Network motifs: theory and experimental approaches. Nat. Rev. Genet. 8, 450–461 (2007).
Lorenz, D. M., Jeng, A. & Deem, M. W. The emergence of modularity in biological systems. Phys. Life Rev. 8, 129–160 (2011).
Erwin, D. H. & Davidson, E. H. The evolution of hierarchical gene regulatory networks. Nat. Rev. Genet. 10, 141–148 (2009).
Krumlauf, R. Hox genes in vertebrate development. Cell 78, 191–201 (1994).
Shubin, N., Tabin, C. & Carroll, S. Deep homology and the origins of evolutionary novelty. Nature 457, 818–823 (2009).
Carroll, S. B. Evo-devo and an expanding evolutionary synthesis: a genetic theory of morphological evolution. Cell 134, 25–36 (2008).
Ohta, T. Slightly deleterious mutant substitutions in evolution. Nature 246, 96–98 (1973).
Lynch, M. & Conery, J. S. The origins of genome complexity. Science 302, 1401–1404 (2003).
Wray, G. A. The evolutionary significance of cis-regulatory mutations. Nat. Rev. Genet. 8, 206–216 (2007).
Schmidt, D. et al. Five-vertebrate ChIP-seq reveals the evolutionary dynamics of transcription factor binding. Science 328, 1036–1040 (2010).
Wagner, G. P. The developmental genetics of homology. Nat. Rev. Genet. 8, 473–479 (2007).
Wagner, G. P. Homology, Genes, and Evolutionary Innovation (Princeton Univ. Press, 2014).
DiFrisco, J., Love, A. C. & Wagner, G. P. Character identity mechanisms: a conceptual model for comparative-mechanistic biology. Biol. Philos. 35, 44 (2020).
Marks, D. S. et al. Protein 3D structure computed from evolutionary sequence variation. PLoS ONE 6, 1–20 (2011).
Marks, D. S., Hopf, T. A. & Sander, C. Protein structure prediction from sequence variation. Nat. Biotechnol. 30, 1072–1080 (2012).
Hopf, T. A. et al. Mutation effects predicted from sequence co-variation. Nat. Biotechnol. 35, 128–135 (2017).
Hecker, N. et al. Enhancer-driven cell type comparison reveals similarities between the mammalian and bird pallium. Science 387, eadp3957 (2025).
McQueen, E. & Rebeiz, M. in Gene Regulatory Networks (ed. Peter, I. S.) 375–405 (Academic, 2020).
Halfon, M. S. Perspectives on gene regulatory network evolution. Trends Genet. 33, 436–447 (2017).
Booth, H. & Hadjivasiliou, Z. Gene network organization, mutation, and selection collectively drive developmental pattern evolvability and predictability. PRX Life 3, 033023 (2025).
Pavlicev, M., Cheverud, J. M. & Wagner, G. P. A model of developmental evolution: selection, pleiotropy and compensation. Trends Ecol. Evol. 27, 316–322 (2012).
McColgan, Á & DiFrisco, J. Understanding developmental system drift. Development 151, dev203054 (2024).
McGinnis, C. S. et al. MULTI-seq: sample multiplexing for single-cell RNA sequencing using lipid-tagged indices. Nat. Methods 16, 619–626 (2019).
van der Maaten, L. J. P. & Hinton, G. E. Visualizing data using t-SNE. J. Mach. Learn. Res. 9, 2579–2605 (2008).
Ternes, L. et al. A multi-encoder variational autoencoder controls multiple transformational features in single-cell image analysis. Commun. Biol. 5, 255 (2022).
Wani, S. A., Khan, S. A. & Quadri, S. scJVAE: a novel method for integrative analysis of multimodal single-cell data. Comput. Biol. Med. 158, 106865 (2023).
Kalafut, N. C., Huang, X. & Wang, D. Joint variational autoencoders for multimodal imputation and embedding. Nat. Mach. Intell. 5, 631–642 (2023).
Gong, B., Zhou, Y. & Purdom, E. Cobolt: integrative analysis of multimodal single-cell sequencing data. Genome Biol. 22, 351 (2021).
Danino, R., Nachman, I. & Sharan, R. Batch correction of single-cell sequencing data via an autoencoder architecture. Bioinform. Adv. 4, vbad186 (2024).
Tran, H. T. N. et al. A benchmark of batch-effect correction methods for single-cell RNA sequencing data. Genome Biol. 21, 12 (2020).
Zhang, Z. et al. scDisInFact: disentangled learning for integration and prediction of multi-batch multi-condition single-cell RNA-sequencing data. Nat. Commun. 15, 912 (2024).
Kana, O. et al. Generative modeling of single-cell gene expression for dose-dependent chemical perturbations. Patterns 4, 100817 (2023).
Yang, Y. et al. The Manatee variational autoencoder model for predicting gene expression alterations caused by transcription factor perturbations. Sci. Rep. 14, 11794 (2024).
Tang, Z., Zhou, M., Zhang, K. & Song, Q. scPerb: predict single-cell perturbation via style transfer-based variational autoencoder. J. Adv. Res. 75, 189–198 (2024).
Rampášek, L., Hidru, D., Smirnov, P., Haibe-Kains, B. & Goldenberg, A. Dr.VAE: improving drug response prediction via modeling of drug perturbation effects. Bioinformatics 35, 3743–3751 (2019).
Chari, T. & Pachter, L. The specious art of single-cell genomics. PLoS Comput. Biol. 19, e1011288 (2023).
Gorban, A. N. & Tyukin, I. Y. Blessing of dimensionality: mathematical foundations of the statistical physics of data. Philos. Trans. R. Soc. A 15, 20170237 (2018).
Kunes, R. Z. et al. Supervised discovery of interpretable gene programs from single-cell data. Nat. Biotechnol. 42, 1084–1095 (2024).
Li, Y. et al. MetaQ: fast, scalable and accurate metacell inference via single-cell quantization. Nat. Commun. 16, 1205 (2025).
Saelens, W., Cannoodt, R., Todorov, H. & Saeys, Y. A comparison of single-cell trajectory inference methods. Nat. Biotechnol. 37, 547–554 (2019).
Lopez, R. et al. Learning causal representations of single cells via sparse mechanism shift modeling. In Proc. 2nd Conference on Causal Learning and Reasoning (eds van der Schaar, M. et al.) 662–691 (PMLR, 2023). This paper is an early example of the application of causal representation learning in biology.
Zhang, J. et al. Identifiability guarantees for causal disentanglement from soft interventions. In Proc. Advances in Neural Information Processing Systems 36 (eds Oh, A. et al.) (NeurIPS, 2023).
Gao, Y., Dong, K., Shan, C., Li, D. & Liu, Q. Causal disentanglement for single-cell representations and controllable counterfactual generation. Nat. Commun. 16, 6775 (2025).
An, S. et al. scCausalVI disentangles single-cell perturbation responses with causality-aware generative model. Cell Syst. 16, 101443 (2025).
Bereket, M. & Karaletsos, T. Modelling cellular perturbations with the sparse additive mechanism shift variational autoencoder. In Proc. Advances in Neural Information Processing Systems 36 (eds Oh, A. et al.) (NeurIPS, 2023).
Baek, S. et al. CRADLE-VAE: enhancing single-cell gene perturbation modeling with counterfactual reasoning-based artifact disentanglement. In Proc. Thirty-Ninth AAAI Conference on Artificial Intelligence 15445–15452 (AAAI Press, 2025).
Mao, H. et al. Learning identifiable factorized causal representations of cellular responses. In Proc. Advances in Neural Information Processing Systems 37 (eds Globerson, A. et al.) (NeurIPS, 2024).
Tejada-Lapuerta, A. et al. Causal machine learning for single-cell genomics. Nat. Genet. 57, 797–808 (2025).
Aliee, H., Kapl, F., Hediyeh-Zadeh, S. & Theis, F. J. Conditionally invariant representation learning for disentangling cellular heterogeneity. Preprint at https://doi.org/10.48550/arXiv.2307.00558 (2023).
Lopez, R., Hütter, J.-C., Pritchard, J. K. & Regev, A. Large-scale differentiable causal discovery of factor graphs. In Proc. Advances in Neural Information Processing Systems 35 (eds Koyejo, S. et al.) (NeurIPS, 2022).
Kvon, E. Z., Waymack, R., Gad, M. & Wunderlich, Z. Enhancer redundancy in development and disease. Nat. Rev. Genet. 22, 324–336 (2021).
Hong, J. W., Hendrix, D. A. & Levine, M. S. Shadow enhancers as a source of evolutionary novelty. Science 321, 1314 (2008).
Thompson, J. J. et al. Extensive co-binding and rapid redistribution of NANOG and GATA6 during emergence of divergent lineages. Nat. Commun. 13, 4257 (2022).
de Boer, C. G. & Taipale, J. Hold out the genome: a roadmap to solving the cis-regulatory code. Nature 625, 41–50 (2024).
Gibney, E. R. & Nolan, C. M. Epigenetics and gene expression. Heredity 105, 4–13 (2010).
Bolt, C. C. & Duboule, D. The regulatory landscapes of developmental genes. Development 147, dev171736 (2020).
Blassberg, R. et al. Sox2 levels regulate the chromatin occupancy of WNT mediators in epiblast progenitors responsible for vertebrate body formation. Nat. Cell Biol. 24, 633–644 (2022).
Kim, S. et al. Deciphering the multi-scale, quantitative cis-regulatory code. Mol. Cell 83, 373–392 (2023).
Mayran, A. et al. Pioneer and nonpioneer factor cooperation drives lineage specific chromatin opening. Nat. Commun. 10, 3807 (2019).
Liu, B. B. et al. An automated ATAC-seq method reveals sequence determinants of transcription factor dose response in the open chromatin. Preprint at bioRxiv https://doi.org/10.1101/2025.07.24.666684 (2025).
Li, D. et al. Chromatin accessibility dynamics during iPSC reprogramming. Cell Stem Cell 21, 819–833.e6 (2017).
Chronis, C. et al. Cooperative binding of transcription factors orchestrates reprogramming. Cell 168, 442–459.e20 (2017).
Narita, T. et al. Acetylation of histone H2B marks active enhancers and predicts CBP/p300 target genes. Nat. Genet. 55, 679–692 (2023).
Lynch, M. The evolution of genetic networks by non-adaptive processes. Nat. Rev. Genet. 8, 803–813 (2007).
Kosicki, M. et al. In vivo mapping of mutagenesis sensitivity of human enhancers. Nature 643, 839–846 (2025).
Koeppel, J. et al. Randomizing the human genome by engineering recombination between repeat elements. Science 387, eado3979 (2025).
Koeppel, J. et al. Resolution of a human super-enhancer by targeted genome randomisation. Preprint at bioRxiv https://doi.org/10.1101/2025.01.14.632548 (2025).
Gasperini, M. et al. A genome-wide framework for mapping gene regulation via cellular genetic screens. Cell 176, 377–390.e19 (2019).
Liu, W. et al. Dissecting the impact of transcription factor dose on cell reprogramming heterogeneity using scTF-seq. Nat. Genet. 57, 2522–2535 (2025).
Frömel, R. et al. Design principles of cell-state-specific enhancers in hematopoiesis. Cell 188, 3202–3218 (2025).
Lalanne, J.-B. et al. Multiplex profiling of developmental cis-regulatory elements with quantitative single-cell expression reporters. Nat. Methods 21, 983–993 (2024).
Cornwall-Scoones, J. et al. Predictable engineering of signal-dependent cis-regulatory elements. Preprint at bioRxiv https://doi.org/10.1101/2025.03.07.642002 (2025).
Buckley, M. et al. Saturation genome editing maps the functional spectrum of pathogenic VHL alleles. Nat. Genet. 56, 1446–1455 (2024).
Zhang, P. et al. Deep flanking sequence engineering for efficient promoter design using DeepSEED. Nat. Commun. 14, 6309 (2023).
Taskiran, I. I. et al. Cell-type-directed design of synthetic enhancers. Nature 626, 212–220 (2024). This study exemplifies the emerging capability to engineer cis-regulatory sequences, supporting the vision of synthetic GRN construction.
Gosai, S. J. et al. Machine-guided design of cell-type-targeting cis-regulatory elements. Nature 634, 1211–1220 (2024).
Gao, X. J., Chong, L. S., Kim, M. S. & Elowitz, M. B. Programmable protein circuits in living cells. Science 361, 1252–1258 (2018).
Chen, Z. et al. A synthetic protein-level neural network in mammalian cells. Science 386, 1243–1250 (2024).
Green, A. A. et al. Complex cellular logic computation using ribocomputing devices. Nature 548, 117–121 (2017).
Chen, Y. et al. Genetic circuit design automation for yeast. Nat. Microbiol. 5, 1349–1360 (2020).
Bunne, C. et al. How to build the virtual cell with artificial intelligence: priorities and opportunities. Cell 187, 7045–7063 (2024).
Johnson, G. T. et al. Building the next generation of virtual cells to understand cellular biology. Biophys. J. 122, 3560–3569 (2023).
Heimberg, G. et al. A cell atlas foundation model for scalable search of similar human cells. Nature 638, 1085–1094 (2024).
Abramson, J. et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature 630, 493–500 (2024).
Booeshaghi, A. S., Galvez-Merchán, Á. & Pachter, L. Algorithms for a Commons Cell Atlas. Preprint at bioRxiv https://doi.org/10.1101/2024.03.23.586413 (2024).
Brixi, G. et al. Genome modeling and design across all domains of life with Evo 2. Preprint at bioRxiv https://doi.org/10.1101/2025.02.18.638918 (2025).
Minsky, M. & Papert, S. Perceptrons (MIT Press, 1969).
Sáez, M. et al. Statistically derived geometrical landscapes capture principles of decision-making dynamics during cell fate transitions. Cell Syst. 13, 12–28.e3 (2022).
Sáez, M., Briscoe, J. & Rand, D. A. Dynamical landscapes of cell fate decisions. Interface Focus 12, 20220002 (2022).
Rand, D. A., Raju, A., Sáez, M., Corson, F. & Siggia, E. D. Geometry of gene regulatory dynamics. Proc. Natl Acad. Sci. USA 118, e2109729118 (2021).
Acknowledgements
The authors are grateful to T. Brown, J. DiFrisco, D. Erwin, F. Fröhlich, L. Parts, P. Badia-i-Mompel and the members of the Briscoe Lab for their constructive comments. This work was supported by the Wellcome Trust (220379/D/20/Z) and the Francis Crick Institute, which receives its core funding from Cancer Research UK, the UK Medical Research Council and the Wellcome Trust (all under FC001051).
Author information
Authors and Affiliations
Contributions
R.J.M. and J.B. researched the literature, R.J.M. wrote the article, and R.J.M. and J.B. contributed substantially to the discussion of the content, and reviewed and edited the manuscript before submission.
Corresponding author
Ethics declarations
Competing interests
R.J.M. is a consultant for Omnipotent Biotechnologies.
Peer review
Peer review information
Nature Reviews Genetics thanks Mohammad Lotfollahi and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Glossary
- Bayesian modelling
-
A probabilistic modelling approach that combines prior beliefs (prior distributions) with observed data (likelihoods) to estimate an updated belief (posterior distributions), allowing inference and prediction with robust uncertainty quantification.
- Coarse-graining
-
A method commonly used in statistical mechanics and chemical simulation methods; details at a lower scale of resolution (for example, atomic level) are removed or averaged such that only features that are essential to preserve macroscopic behaviour at a higher scale (for example, bio-molecular) are retained.
- Dimension reduction
-
A class of methods used to reduce the number of variables in a dataset while retaining the major sources of variation in the data. Methods such as principal component analysis, uniform manifold approximation and projection, and t-distributed stochastic neighbor embedding are commonly used to reduce the dimension of sequencing data.
- Dynamical systems
-
Models of systems whose state evolves through time, often expressed in the form of differential equations; often used to describe biological systems such as population dynamics, biochemical reactions and gene regulatory networks.
- Ground truths
-
Real-world models or outcome of a system, serving as a benchmark against which trained models can be compared to assess performance.
- Marr’s levels of analysis
-
A framework introduced by David Marr to describe cognitive and computational systems that breaks these systems into three layers: the computational level (what the system does and why), the algorithmic or representational level (organization of the system required to fulfill the computational purpose), and the implementational level (the physical material and substrates used to construct the algorithmic organization).
- Multi-perspectival
-
The idea that the representation of a phenomenon may depend on the perspective of the viewer or the question of the researcher; the most useful representation of a gene regulatory system may differ, for example, for questions concerning global cell-type control versus questions concerning regulation of specific genes.
- Multi-scale
-
Models or analysis that represent phenomena across different scales of resolution in time, space or organization, for example, the atomic, molecular, bio-molecular, genetic, cellular, organismal and population scales of biology.
- Representation learning
-
Related to dimension reduction, a field of machine learning focused on learning meaningful, compact representations of data rather than using only the variables observed in the original data; an example of representation learning is the variational autoencoder, a type of deep generative model that encodes data to a latent representation.
- Sloppy models
-
Systems biology models that show extreme sensitivity to a small number of parameters, with the majority of parameters having no effect on model performance.
- Structural non-identifiability
-
When wide variation in the parameters of a model produce small changes in model output such that there is not a unique solution for fitting a model to a dataset.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Maizels, R.J., Briscoe, J. Gene regulatory networks: from correlative models to causal explanations. Nat Rev Genet (2026). https://doi.org/10.1038/s41576-026-00939-1
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41576-026-00939-1