Abstract
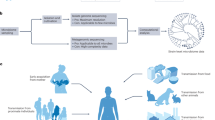
Indoor environments serve as habitat for humans and are replete with various reservoirs and niches for microorganisms. Microorganisms enter indoor spaces with their human and non-human hosts, as well as via exchange with outdoor sources, such as ventilation and plumbing. Once inside, many microorganisms do not survive, especially on dry, barren surfaces. Even reduced, this microbial biomass has critical implications for the health of human occupants. As urbanization escalates, exploring the intersection of the indoor environment with the human microbiome and health is increasingly vital. The indoor microbiome, a complex ecosystem of microorganisms influenced by human activities and environmental factors, plays a pivotal role in modulating infectious diseases and fostering healthy immune development. Recent advancements in microbiome research shed light on this unique ecological system, highlighting the need for innovative approaches in creating health-promoting living spaces. In this Review, we explore the microbial ecology of built environments — places where humans spend most of their lives — and its implications for immune, endocrine and neurological health. We further propose strategies to harness the indoor microbiome for better health outcomes.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Blaustein, R. A. et al. Toothbrush microbiomes feature a meeting ground for human oral and environmental microbiota. Microbiome 9, 32 (2021).
Ben Maamar, S., Hu, J. & Hartmann, E. M. Implications of indoor microbial ecology and evolution on antibiotic resistance. J. Expo. Sci. Environ. Epidemiol. 30, 1–15 (2020).
Parajuli, A. et al. Urbanization reduces transfer of diverse environmental microbiota indoors. Front. Microbiol. 9, 84 (2018).
Strachan, D. P. Hay fever, hygiene, and household size. BMJ 299, 1259–1260 (1989).
Qin, N. et al. Longitudinal survey of microbiome associated with particulate matter in a megacity. Genome Biol. 21, 55 (2020).
Gibbons, S. M. The built environment is a microbial wasteland. mSystems 1, e00033–e00116 (2016).
Ishaq, S. L. et al. Introducing the microbes and social equity working group: considering the microbial components of social, environmental, and health justice. mSystems 6, e0047121 (2021).
Gilbert, J. A. & Stephens, B. Microbiology of the built environment. Nat. Rev. Microbiol. 16, 661–670 (2018).
Prussin, A. J. II & Marr, L. C. Sources of airborne microorganisms in the built environment. Microbiome 3, 78 (2015).
Pausan, M.-R., Blohs, M., Mahnert, A. & Moissl-Eichinger, C. The sanitary indoor environment — a potential source for intact human-associated anaerobes. npj Biofilms Microbiomes 8, 44 (2022). This study demonstrates that most of bacterial and archaeal taxa in the built environment are of human origin and that aerobic and stress-resistant taxa have a survival advantage.
Scaturro, M. et al. Premise plumbing bacterial communities in four European cities and their association with Legionella. Front. Microbiomes https://doi.org/10.3389/frmbi.2023.1170824 (2023).
Stein, M. M. et al. Innate immunity and asthma risk in Amish and Hutterite farm children. N. Engl. J. Med. 375, 411–421 (2016).
Lax, S. et al. Bacterial colonization and succession in a newly opened hospital. Sci. Transl. Med. 9, eaah6500 (2017).
Qian, J., Hospodsky, D., Yamamoto, N., Nazaroff, W. W. & Peccia, J. Size-resolved emission rates of airborne bacteria and fungi in an occupied classroom. Indoor Air 22, 339–351 (2012).
Lax, S. et al. Longitudinal analysis of microbial interaction between humans and the indoor environment. Science 345, 1048–1052 (2014).
Young, G. R., Sherry, A. & Smith, D. L. Built environment microbiomes transition from outdoor to human-associated communities after construction and commissioning. Sci. Rep. 13, 15854 (2023).
Kembel, S. W. et al. Architectural design influences the diversity and structure of the built environment microbiome. ISME J. 6, 1469–1479 (2012).
Hoisington, A. J. et al. Ten questions concerning the built environment and mental health. Build. Environ. 155, 58–69 (2019).
Meadow, J. F. et al. Indoor airborne bacterial communities are influenced by ventilation, occupancy, and outdoor air source. Indoor Air 24, 41–48 (2014).
Carstens, C. K., Salazar, J. K., Sharma, S. V., Chan, W. & Darkoh, C. Evaluation of the kitchen microbiome and food safety behaviors of predominantly low-income families. Front. Microbiol. 13, 987925 (2022).
Flores, G. E. et al. Diversity, distribution and sources of bacteria in residential kitchens. Environ. Microbiol. 15, 588–596 (2013).
Adams, R. I., Bateman, A. C., Bik, H. M. & Meadow, J. F. Microbiota of the indoor environment: a meta-analysis. Microbiome 3, 49 (2015).
Gibbons, S. M. et al. Ecological succession and viability of human-associated microbiota on restroom surfaces. Appl. Environ. Microbiol. 81, 765–773 (2015).
Cox, J. et al. Associations of observed home dampness and mold with the fungal and bacterial dust microbiomes. Environ. Sci. Process. Impacts 23, 491–500 (2021).
Jayaprakash, B. et al. Indoor microbiota in severely moisture damaged homes and the impact of interventions. Microbiome 5, 138 (2017).
Lai, P. S. et al. The classroom microbiome and asthma morbidity in children attending 3 inner-city schools. J. Allergy Clin. Immunol. 141, 2311–2313 (2018).
Green, J. L. Can bioinformed design promote healthy indoor ecosystems? Indoor Air 24, 113–115 (2014).
Jin, L. et al. Integrating environmental dimensions of ‘One Health’ to combat antimicrobial resistance: essential research needs. Environ. Sci. Technol. 56, 14871–14874 (2022).
Fahimipour, A. K. et al. Antimicrobial chemicals associate with microbial function and antibiotic resistance indoors. mSystems 3, e00200-18 (2018).
Hartmann, E. M. et al. Antimicrobial chemicals are associated with elevated antibiotic resistance genes in the indoor dust microbiome. Environ. Sci. Technol. 50, 9807–9815 (2016).
Mahnert, A. et al. Man-made microbial resistances in built environments. Nat. Commun. 10, 968 (2019). This study demonstrates that increasing isolation within an indoor space is associated with decreased microbial diversity and a shift from Gram-positive bacteria such as Actinobacteria and Firmicutes, to Gram-negative Proteobacteria, many of which are antibiotic-resistant.
Guevarra, R. B. et al. Metagenomic characterization of bacterial community and antibiotic resistance genes found in the mass transit system in Seoul, South Korea. Ecotoxicol. Environ. Saf. 246, 114176 (2022).
Leung, M. H. Y. et al. Characterization of the public transit air microbiome and resistome reveals geographical specificity. Microbiome 9, 112 (2021).
Kang, K. et al. The environmental exposures and inner- and intercity traffic flows of the metro system may contribute to the skin microbiome and resistome. Cell Rep. 24, 1190–1202.e5 (2018).
Danko, D. et al. A global metagenomic map of urban microbiomes and antimicrobial resistance. Cell 184, 3376–3393.e17 (2021).
Singh, N. K., Wood, J. M., Karouia, F. & Venkateswaran, K. Succession and persistence of microbial communities and antimicrobial resistance genes associated with International Space Station environmental surfaces. Microbiome 6, 204 (2018).
Klassert, T. E. et al. Bacterial colonization dynamics and antibiotic resistance gene dissemination in the hospital environment after first patient occupancy: a longitudinal metagenetic study. Microbiome 9, 169 (2021).
Gupta, M., Lee, S., Bisesi, M. & Lee, J. Indoor microbiome and antibiotic resistance on floor surfaces: an exploratory study in three different building types. Int. J. Environ. Res. Public Health 16, 4160 (2019).
Chng, K. R. et al. Cartography of opportunistic pathogens and antibiotic resistance genes in a tertiary hospital environment. Nat. Med. 26, 941–951 (2020).
Brooks, B. et al. The developing premature infant gut microbiome is a major factor shaping the microbiome of neonatal intensive care unit rooms. Microbiome 6, 112 (2018).
Brooks, B. et al. Strain-resolved analysis of hospital rooms and infants reveals overlap between the human and room microbiome. Nat. Commun. 8, 1814 (2017). This study reports the application of strain tracking to demonstrate the transfer of microorganisms from the environment to human inhabitants.
Hu, J., Shuai, W., Sumner, J. T., Moghadam, A. A. & Hartmann, E. M. Clinically relevant pathogens on surfaces display differences in survival and transcriptomic response in relation to probiotic and traditional cleaning strategies. npj Biofilms Microbiomes 8, 72 (2022).
Logan-Jackson, A. R. et al. A critical review on the factors that influence opportunistic premise plumbing pathogens: from building entry to fixtures in residences. Environ. Sci. Technol. 57, 6360–6372 (2023).
Mathews, C. L. et al. Crosslinked polyethylene (PEX) drinking water pipe: carbon leaching, impacts on microbial growth, and developmental toxicity to zebrafish. Environ. Adv. 13, 100386 (2023).
Webster, T. M. et al. Structure and functional attributes of bacterial communities in premise plumbing across the United States. Environ. Sci. Technol. 55, 14105–14114 (2021).
Diorio-Toth, L. et al. Intensive care unit sinks are persistently colonized with multidrug resistant bacteria and mobilizable, resistance-conferring plasmids. mSystems 8, e0020623 (2023).
Franco, L. C. et al. A microbiological survey of handwashing sinks in the hospital built environment reveals differences in patient room and healthcare personnel sinks. Sci. Rep. 10, 8234 (2020).
Bosch, T. C. G. et al. The potential importance of the built-environment microbiome and its impact on human health. Proc. Natl Acad. Sci. USA 121, e2313971121 (2024).
Fahimipour, A. K. et al. Daylight exposure modulates bacterial communities associated with household dust. Microbiome 6, 175 (2018).
Lax, S. et al. Microbial and metabolic succession on common building materials under high humidity conditions. Nat. Commun. 10, 1767 (2019).
Mhuireach, G. Á. et al. Differing effects of four building materials on viable bacterial communities and VOCs. Dev. Built Environ. 7, 100055 (2021).
Hu, J. et al. Impacts of indoor surface finishes on bacterial viability. Indoor Air 29, 551–562 (2019).
Haines, S. R. et al. Microbial growth and volatile organic compound (VOC) emissions from carpet and drywall under elevated relative humidity conditions. Microbiome 9, 209 (2021). This study proposes a quantitative framework that links microbial metabolic activity to the availability of water, addressing the critical question of when indoor microbiota are viable or active.
Hegarty, B., Haverinen-Shaughnessy, U., Shaughnessy, R. J. & Peccia, J. Spatial gradients of fungal abundance and ecology throughout a damp building. Environ. Sci. Technol. Lett. 6, 329–333 (2019).
Ling, F., Whitaker, R., LeChevallier, M. W. & Liu, W.-T. Drinking water microbiome assembly induced by water stagnation. ISME J. 12, 1520–1531 (2018).
Angert, D. M. et al. Water quality during extended stagnation and flushing in a college residential hall. Environ. Sci. Water Res. Technol. https://doi.org/10.1039/D3EW00038A (2023).
Ley, C. J. et al. Drinking water microbiology in a water-efficient building: stagnation, seasonality, and physicochemical effects on opportunistic pathogen and total bacteria proliferation. Environ. Sci. Water Res. Technol. 6, 2902–2913 (2020).
Montagnino, E. et al. Over the weekend: water stagnation and contaminant exceedances in a green office building. PLOS Water 1, e0000006 (2022).
Prussin, A. J. II et al. Viruses in the built environment (VIBE) meeting report. Microbiome 8, 1 (2020).
Prussin, A. J. II et al. Seasonal dynamics of DNA and RNA viral bioaerosol communities in a daycare center. Microbiome 7, 53 (2019).
Du, S. et al. Highly host-linked viromes in the built environment possess habitat-dependent diversity and functions for potential virus–host coevolution. Nat. Commun. 14, 2676 (2023).
Mokili, J. L., Rohwer, F. & Dutilh, B. E. Metagenomics and future perspectives in virus discovery. Curr. Opin. Virol. 2, 63–77 (2012).
Bahram, M., Anslan, S., Hildebrand, F., Bork, P. & Tedersoo, L. Newly designed 16S rRNA metabarcoding primers amplify diverse and novel archaeal taxa from the environment. Environ. Microbiol. Rep. 11, 487–494 (2019).
Hsu, T. et al. Urban transit system microbial communities differ by surface type and interaction with humans and the environment. mSystems 1, e00018-16 (2016).
Pakpour, S. et al. Presence of Archaea in the indoor environment and their relationships with housing characteristics. Microb. Ecol. 72, 305–312 (2016). This study focuses on the presence of Archaea on indoor surfaces, complementing sequencing data with culture-based experiments.
Amin, H. et al. Indoor airborne microbiome and endotoxin: meteorological events and occupant characteristics are important determinants. Environ. Sci. Technol. 57, 11750–11766 (2023).
Mahnert, A., Haratani, M., Schmuck, M. & Berg, G. Enriching beneficial microbial diversity of indoor plants and their surrounding built environment with biostimulants. Front. Microbiol. 9, 2985 (2018).
Weiss, H. et al. The airplane cabin microbiome. Microb. Ecol. 77, 87–95 (2019).
Sun, Y. et al. Shotgun metagenomics of dust microbiome from flight deck and cabin in civil aviation aircraft. Indoor Air 30, 1199–1212 (2020).
Mahnert, A. et al. Microbiome dynamics during the HI-SEAS IV mission, and implications for future crewed missions beyond Earth. Microbiome 9, 27 (2021).
Avila-Herrera, A. et al. Crewmember microbiome may influence microbial composition of ISS habitable surfaces. PLoS ONE 15, e0231838 (2020).
Checinska Sielaff, A. et al. Characterization of the total and viable bacterial and fungal communities associated with the International Space Station surfaces. Microbiome 7, 50 (2019).
Klimenko, N. S. et al. Co-occurrence patterns of bacteria within microbiome of Moscow subway. Comput. Struct. Biotechnol. J. 18, 314–322 (2020).
Gohli, J. et al. The subway microbiome: seasonal dynamics and direct comparison of air and surface bacterial communities. Microbiome 7, 160 (2019).
Hernández, A. M., Vargas-Robles, D., Alcaraz, L. D. & Peimbert, M. Station and train surface microbiomes of Mexico City’s metro (subway/underground). Sci. Rep. 10, 8798 (2020).
Fu, X. et al. Continental-scale microbiome study reveals different environmental characteristics determining microbial richness, composition, and quantity in hotel rooms. mSystems 5, e00119–e00120 (2020).
Nygaard, A. B. & Charnock, C. Longitudinal development of the dust microbiome in a newly opened Norwegian kindergarten. Microbiome 6, 159 (2018).
Richardson, M., Gottel, N., Gilbert, J. A. & Lax, S. Microbial similarity between students in a common dormitory environment reveals the forensic potential of individual microbial signatures. mBio 10, e01054–e01119 (2019).
Sharma, A. et al. Longitudinal homogenization of the microbiome between both occupants and the built environment in a cohort of United States air force cadets. Microbiome 7, 70 (2019). This study is a rare example of an indoor microbiome study involving a highly behaviourally homogenous participant cohort and a reproducible building design.
Lee, M. D. et al. Reference-guided metagenomics reveals genome-level evidence of potential microbial transmission from the ISS environment to an astronaut’s microbiome. iScience 24, 102114 (2021).
Sharma, A. & Gilbert, J. A. Microbial exposure and human health. Curr. Opin. Microbiol. 44, 79–87 (2018).
Hoare, E., Jacka, F. & Berk, M. The impact of urbanization on mood disorders: an update of recent evidence. Curr. Opin. Psychiatry 32, 198–203 (2019).
Kaplan, R. C. et al. Gut microbiome composition in the Hispanic Community Health Study/Study of Latinos is shaped by geographic relocation, environmental factors, and obesity. Genome Biol. 20, 219 (2019).
Fakunle, A. G., Jafta, N., Okekunle, A. P. & Naidoo, R. N. Indoor microbiome and risk of lower respiratory tract infections among children under-five years: a meta-analysis. Indoor Air 30, 795–804 (2020). This study presents a meta-analysis to demonstrate that exposure to high indoor microbial concentration increases the risk of lower respiratory tract infections.
Richardson, M. et al. Concurrent measurement of microbiome and allergens in the air of bedrooms of allergy disease patients in the Chicago area. Microbiome 7, 82 (2019).
Pearson, A. L. et al. Associations detected between measures of neighborhood environmental conditions and human microbiome diversity. Sci. Total Environ. 745, 141029 (2020).
Arnold, W. A. et al. Quaternary ammonium compounds: a chemical class of emerging concern. Environ. Sci. Technol. 57, 7645–7665 (2023).
Fu, X. et al. Associations between respiratory infections and bacterial microbiome in student dormitories in Northern China. Indoor Air 30, 816–826 (2020).
Marotz, C. et al. SARS-CoV-2 detection status associates with bacterial community composition in patients and the hospital environment. Microbiome 9, 132 (2021).
Minich, J. J. et al. Feasibility of using alternative swabs and storage solutions for paired SARS-CoV-2 detection and microbiome analysis in the hospital environment. Microbiome 9, 25 (2021).
Weber, D. J., Anderson, D. & Rutala, W. A. The role of the surface environment in healthcare-associated infections. Curr. Opin. Infect. Dis. 26, 338–344 (2013).
Alverdy, J. C., Hyman, N. & Gilbert, J. Re-examining causes of surgical site infections following elective surgery in the era of asepsis. Lancet Infect. Dis. 20, e38–e43 (2020). This perspective highlights extensive new data suggesting that the occurrence of infectious disease in human subjects indoors is more likely owing to opportunistic pathobionts already present in a person’s microbiome rather than acquisition from fomite.
Gilbert, J. A. & Alverdy, J. Where do the pathogens that cause surgical site infections come from? Sci. Transl. Med. 16, eado1449 (2024).
Johnson, C. R. et al. Nano-enabled, antimicrobial toothbrushes — how physical and chemical properties relate to antibacterial capabilities. J. Hazard. Mater. 396, 122445 (2020).
Kelley, S. T. et al. Altered microbiomes in thirdhand smoke-exposed children and their home environments. Pediatr. Res. 90, 1153–1160 (2021).
Fu, X. et al. Association between indoor microbiome exposure and sick building syndrome (SBS) in junior high schools of Johor Bahru, Malaysia. Sci. Total Environ. 753, 141904 (2021).
Indoor Air Quality Research. Report on a WHO meeting. Stockholm, 27–31 August 1984. EURO Rep. Stud. 1–64 (1986).
Ha, C. W. Y. et al. Translocation of viable gut microbiota to mesenteric adipose drives formation of creeping fat in humans. Cell 183, 666–683.e17 (2020).
Turnbaugh, P. J. et al. An obesity-associated gut microbiome with increased capacity for energy harvest. Nature 444, 1027–1031 (2006).
Scepanovic, P. et al. A comprehensive assessment of demographic, environmental, and host genetic associations with gut microbiome diversity in healthy individuals. Microbiome 7, 130 (2019).
Fyfe-Johnson, A. L. et al. Nature contact and children’s health: a systematic review. In Council on Community Pediatrics Program. https://doi.org/10.1542/peds.147.3_meetingabstract.50 (American Academy of Pediatrics, 2021).
Haahtela, T. Clinical application of the biodiversity hypothesis in the management of allergic disorders. in: Evolution, Biodiversity and a Reassessment of the Hygiene Hypothesis 393–414 (Springer International Publishing, 2022).
Lowry, C. A. et al. The microbiota, immunoregulation, and mental health: implications for public health. Curr. Environ. Health Rep. 3, 270–286 (2016).
Diaz Heijtz, R. et al. Normal gut microbiota modulates brain development and behavior. Proc. Natl Acad. Sci. USA 108, 3047–3052 (2011).
Cryan, J. F. et al. The microbiota–gut–brain axis. Physiol. Rev. 99, 1877–2013 (2019).
Foster, J. A., Rinaman, L. & Cryan, J. F. Stress & the gut–brain axis: regulation by the microbiome. Neurobiol. Stress 7, 124–136 (2017).
Deckers, J., Marsland, B. J. & von Mutius, E. Protection against allergies: microbes, immunity, and the farming effect. Eur. J. Immunol. 51, 2387–2398 (2021).
Kirjavainen, P. V. et al. Farm-like indoor microbiota in non-farm homes protects children from asthma development. Nat. Med. 25, 1089–1095 (2019).
von Mutius, E. Allergies, infections and the hygiene hypothesis — the epidemiological evidence. Immunobiology 212, 433–439 (2007).
Sun, Y. et al. Indoor metabolites and chemicals outperform microbiome in classifying childhood asthma and allergic rhinitis. Eco-Environ. Health 2, 208–218 (2023).
Fu, X. et al. Associations between species-level indoor microbiome, environmental characteristics, and asthma in junior high schools of Terengganu, Malaysia. Air Qual. Atmos. Health 15, 1043–1055 (2022).
Fu, X. et al. Indoor microbiome, environmental characteristics and asthma among junior high school students in Johor Bahru, Malaysia. Environ. Int. 138, 105664 (2020).
Sun, Y. et al. Indoor microbiome, microbial and plant metabolites, chemical compounds, and asthma symptoms in junior high school students: a multicentre association study in Malaysia. Eur. Respir. J. 60, 2200260 (2022).
Karvonen, A. M. et al. Indoor bacterial microbiota and development of asthma by 10.5 years of age. J. Allergy Clin. Immunol. 144, 1402–1410 (2019). This paper shows an inverse correlation between bacterial diversity and the development of asthma by age 10, revealing that the proportion of Lactococcus spp. is a significant risk factor for asthma onset, whereas the proportion of Actinomycetales is associated with lower asthma risk.
Lynch, S. V. et al. Effects of early-life exposure to allergens and bacteria on recurrent wheeze and atopy in urban children. J. Allergy Clin. Immunol. 134, 593–601.e12 (2014).
von Mutius, E. & Vercelli, D. Farm living: effects on childhood asthma and allergy. Nat. Rev. Immunol. 10, 861–868 (2010).
Huang, Y. J. & Boushey, H. A. The microbiome in asthma. J. Allergy Clin. Immunol. 135, 25–30 (2015).
Meggs, W. J. Epidemics of mold poisoning past and present. Toxicol. Ind. Health 25, 571–576 (2009).
Fisk, W. J., Eliseeva, E. A. & Mendell, M. J. Association of residential dampness and mold with respiratory tract infections and bronchitis: a meta-analysis. Environ. Health 9, 72 (2010).
Chowdhary, A., Agarwal, K. & Meis, J. F. Filamentous fungi in respiratory infections. What lies beyond aspergillosis and mucormycosis? PLoS Pathog. 12, e1005491 (2016).
Hedayati, M. T., Pasqualotto, A. C., Warn, P. A., Bowyer, P. & Denning, D. W. Aspergillus flavus: human pathogen, allergen and mycotoxin producer. Microbiology 153, 1677–1692 (2007).
Barbeau, D. N., Grimsley, L. F., White, L. E., El-Dahr, J. M. & Lichtveld, M. Mold exposure and health effects following Hurricanes Katrina and Rita. Annu. Rev. Public Health 31, 165–178 (2010).
Zaramela, L. S., Tjuanta, M., Moyne, O., Neal, M. & Zengler, K. SynDNA — a synthetic DNA spike-in method for absolute quantification of shotgun metagenomic sequencing. mSystems 7, e0044722 (2022).
Shen, J. et al. An improved workflow for accurate and robust healthcare environmental surveillance using metagenomics. Microbiome 10, 206 (2022). This paper discusses the technical issues associated with sequencing-based surveillance of microbial communities on indoor surfaces and provides tools and protocols to address these challenges.
Minich, J. J. et al. KatharoSeq enables high-throughput microbiome analysis from low-biomass samples. mSystems 3, e00218–e00317 (2018).
Jackson, R. J., Dannenberg, A. L. & Frumkin, H. Health and the built environment: 10 years after. Am. J. Public Health 103, 1542–1544 (2013).
Frumkin, H. et al. Nature contact and human health: a research agenda. Environ. Health Perspect. 125, 075001 (2017).
Wells, N. M. & Evans, G. W. Nearby nature: a buffer of life stress among rural children. Environ. Behav. 35, 311–330 (2003).
Balseviciene, B. et al. Impact of residential greenness on preschool children’s emotional and behavioral problems. Int. J. Environ. Res. Public Health 11, 6757–6770 (2014).
Fjørtoft, I. The natural environment as a playground for children: the impact of outdoor play activities in pre-primary school children. Early Child. Educ. J. 29, 111–117 (2001).
Copeland, K. A., Khoury, J. C. & Kalkwarf, H. J. Child care center characteristics associated with preschoolers’ physical activity. Am. J. Prev. Med. 50, 470–479 (2016).
Kim, J.-H., Lee, C. & Sohn, W. Urban natural environments, obesity, and health-related quality of life among Hispanic children living in inner-city neighborhoods. Int. J. Environ. Res. Public Health 13, 121 (2016).
Bell, J. F., Wilson, J. S. & Liu, G. C. Neighborhood greenness and 2-year changes in body mass index of children and youth. Am. J. Prev. Med. 35, 547–553 (2008).
Dadvand, P. et al. Risks and benefits of green spaces for children: a cross-sectional study of associations with sedentary behavior, obesity, asthma, and allergy. Environ. Health Perspect. 122, 1329–1335 (2014).
Lovasi, G. S. et al. Neighborhood safety and green space as predictors of obesity among preschool children from low-income families in New York City. Prev. Med. 57, 189–193 (2013).
Gordon, J. et al. A simple novel device for air sampling by electrokinetic capture. Microbiome 3, 79 (2015).
Mills, J. G. et al. Revegetation of urban green space rewilds soil microbiotas with implications for human health and urban design. Restor. Ecol. https://doi.org/10.1111/rec.13175 (2020).
Beckett, R. Probiotic design. J. Archit. 26, 6–31 (2021).
González, L. M., Mukhitov, N. & Voigt, C. A. Resilient living materials built by printing bacterial spores. Nat. Chem. Biol. 16, 126–133 (2020).
Berg, N. W. et al. Safety assessment of the use of Bacillus-based cleaning products. Food Chem. Toxicol. 116, 42–52 (2018).
Spök, A., Arvanitakis, G. & McClung, G. Status of microbial based cleaning products in statutory regulations and ecolabelling in Europe, the USA, and Canada. Food Chem. Toxicol. 116, 10–19 (2018).
Neu, L. & Hammes, F. Feeding the building plumbing microbiome: the importance of synthetic polymeric materials for biofilm formation and management. Water 12, 1774 (2020).
Nodehi, M., Ozbakkaloglu, T. & Gholampour, A. A systematic review of bacteria-based self-healing concrete: biomineralization, mechanical, and durability properties. J. Build. Eng. 49, 104038 (2022).
Birch, E., Bridgens, B., Zhang, M. & Dade-Robertson, M. Bacterial spore-based hygromorphs: a novel active material with potential for architectural applications. Sustain. Sci. Pract. Policy 13, 4030 (2021).
Ramos, T., Dedesko, S., Siegel, J. A., Gilbert, J. A. & Stephens, B. Spatial and temporal variations in indoor environmental conditions, human occupancy, and operational characteristics in a new hospital building. PLoS ONE 10, e0118207 (2015). This study demonstrates the potential of integrating genetically modified microorganisms into 3D printed materials, paving the way for novel interventions in the indoor environment to reduce infectious disease risk and improve immune health.
Stephens, B. et al. Microbial exchange via fomites and implications for human health. Curr. Pollut. Rep. 5, 198–213 (2019).
Gomez-Silvan, C. et al. A comparison of methods used to unveil the genetic and metabolic pool in the built environment. Microbiome 6, 71 (2018).
Sui, H.-Y. et al. Impact of DNA extraction method on variation in human and built environment microbial community and functional profiles assessed by shotgun metagenomics sequencing. Front. Microbiol. 11, 953 (2020).
Maestre, J. P. et al. Filter forensics: microbiota recovery from residential HVAC filters. Microbiome 6, 22 (2018).
Hampton-Marcell, J. T., Ghosh, A., Gukeh, M. J. & Megaridis, C. M. A new approach of microbiome monitoring in the built environment: feasibility analysis of condensation capture. Microbiome 11, 129 (2023).
Kapono, C. A. et al. Creating a 3D microbial and chemical snapshot of a human habitat. Sci. Rep. 8, 3669 (2018).
Zolfo, M. et al. Profiling microbial strains in urban environments using metagenomic sequencing data. Biol. Direct 13, 9 (2018).
Acknowledgements
K. Gilbert (UCSD Health Sciences) was responsible for drafting the included figures.
Author information
Authors and Affiliations
Contributions
The authors contributed equally to all aspects of the article.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Reviews Microbiology thanks Mark Hernandez, Martin Täubel and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Gilbert, J.A., Hartmann, E.M. The indoors microbiome and human health. Nat Rev Microbiol 22, 742–755 (2024). https://doi.org/10.1038/s41579-024-01077-3
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41579-024-01077-3
This article is cited by
-
Occupants and surface types drive microbial dynamics in controlled indoor environments
Environmental Microbiome (2025)
-
Human microbiome acquisition and transmission
Nature Reviews Microbiology (2025)
-
An early microbial landscape: inspiring endeavor from the China Space Station Habitation Area Microbiome Program (CHAMP)
Science China Life Sciences (2025)
-
Fungal-Associated Endotypes as a Treatable Trait in Bronchiectasis
Pulmonary Therapy (2025)