Abstract
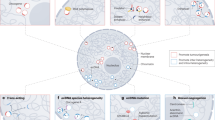
Extrachromosomal DNA (ecDNA) has emerged as a major driver of genomic instability and rapid tumour evolution in urothelial carcinoma. In urothelial carcinoma, ecDNA amplifies oncogenes, reshapes 3D chromatin interactions, reprogrammes transcription and modulates the tumour–immune interface. Together, these features fuel intratumour heterogeneity, accelerate APOBEC3-associated mutational evolution and contribute to aggressive disease. Advances in sequencing and imaging technologies have greatly expanded our understanding of ecDNA biology. Importantly, ecDNA can be detected through non-invasive liquid biopsies, including urine and plasma, and inferred from standard histopathology slides via digital pathology. These observations suggest that ecDNA could be a valuable adjunct biomarker, enhancing current strategies for early detection, patient stratification and dynamic monitoring of treatment response.
Key points
-
Extrachromosomal DNA (ecDNA) occurs in more than one-third of urothelial carcinomas and some precancerous lesions, driving strong oncogene amplification and defining a TP53 and chromothripsis-associated evolutionary trajectory distinct from the indolent FGFR3-mutant pathway.
-
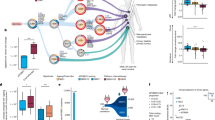
ecDNA shapes clonal evolution in urothelial carcinoma by enabling rapid copy-number remodelling and interacting with mutational processes such as APOBEC3-driven mutagenesis, thereby promoting tumour adaptation and multifocality.
-
ecDNA confers selective advantages under therapeutic stress and promotes immune evasion, in part by downregulating major histocompatibility complex class I expression in urothelial carcinoma cells.
-
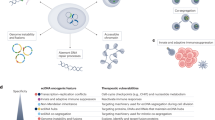
ecDNA is detectable through urothelial carcinoma-specific liquid biopsy approaches — including urine-based and plasma-based assays — and can be inferred from routine haematoxylin-and-eosin-stained slides using emerging digital pathology tools.
-
Targeting ecDNA biology offers a potential approach to modulating oncogene amplification and therapeutic resistance, and could be a promising avenue for future therapeutic development.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Glaser, A. P., Fantini, D., Shilatifard, A., Schaeffer, E. M. & Meeks, J. J. The evolving genomic landscape of urothelial carcinoma. Nat. Rev. Urol. 14, 215–229 (2017).
Barsoum, I. H. Y. & Roux, J. W. T. Upper tract urothelial cancer. Surgery 43, 673–678 (2025).
Zeng, Y. et al. Mapping the chromothripsis landscape in urothelial carcinoma unravels great intratumoral and intertumoral heterogeneity. iScience 28, 111510 (2025).
Dyrskjot, L. et al. Bladder cancer. Nat. Rev. Dis. Primers 9, 58 (2023).
Prip, F. et al. Comprehensive genomic characterization of early-stage bladder cancer. Nat. Genet. 57, 115–125 (2025).
Robertson, A. G. et al. Comprehensive molecular characterization of muscle-invasive bladder cancer. Cell 171, 540–556 (2017).
Fujii, Y. et al. Molecular classification and diagnostics of upper urinary tract urothelial carcinoma. Cancer Cell 39, 793–809 (2021).
Nakauma-Gonzalez, J. A. et al. Whole-genome mapping of APOBEC mutagenesis in metastatic urothelial carcinoma identifies driver hotspot mutations and a novel mutational signature. Cell Genom. 4, 100528 (2024). In this investigation APOBEC-driven hotspot mutations in metastatic urothelial carcinoma are mapped and recurrent driver events are identified, defining key mutational processes that shape tumour evolution.
Baard, J. et al. Diagnostic dilemmas in patients with upper tract urothelial carcinoma. Nat. Rev. Urol. 14, 181–191 (2017).
Knowles, M. A. & Hurst, C. D. Molecular biology of bladder cancer: new insights into pathogenesis and clinical diversity. Nat. Rev. Cancer 15, 25–41 (2015).
Tran, L., Xiao, J. F., Agarwal, N., Duex, J. E. & Theodorescu, D. Advances in bladder cancer biology and therapy. Nat. Rev. Cancer 21, 104–121 (2021).
Zeng, Y. C. et al. Recent development of urinary biomarkers for bladder cancer diagnosis and monitoring. Clin. Transl. Discov. 3, e183 (2023).
Nguyen, D. D. et al. The interplay of mutagenesis and ecDNA shapes urothelial cancer evolution. Nature 635, 219–228 (2024). The results of this study show that CCND1 ecDNA drives treatment resistance in urothelial cancer, defining how mutational processes and ecDNA-forming structural variants shape tumour evolution.
Lv, W. et al. Spatial-temporal diversity of extrachromosomal DNA shapes urothelial carcinoma evolution and the tumor immune microenvironment. Cancer Discov. 15, 1225–1246 (2025). The results of this study provide the most comprehensive ecDNA landscape in urothelial carcinoma to date, showing that ecDNA drives malignant transformation, multifocality and immune evasion.
deCarvalho, A. C. et al. Discordant inheritance of chromosomal and extrachromosomal DNA elements contributes to dynamic disease evolution in glioblastoma. Nat. Genet. 50, 708–717 (2018).
Yi, E. et al. Live-cell imaging shows uneven segregation of extrachromosomal DNA elements and transcriptionally active extrachromosomal DNA hubs in cancer. Cancer Discov. 12, 468–483 (2022).
Pal Choudhuri, S. et al. Acquired cross-resistance in small cell lung cancer due to extrachromosomal DNA amplification of MYC paralogs. Cancer Discov. 14, 804–827 (2024).
Pang, J. et al. Extrachromosomal DNA in HPV-mediated oropharyngeal cancer drives diverse oncogene transcription. Clin. Cancer Res. 27, 6772–6786 (2021).
Fiorini, E. et al. MYC ecDNA promotes intratumour heterogeneity and plasticity in PDAC. Nature 640, 811–820 (2025).
Song, K. et al. Plasticity of extrachromosomal and intrachromosomal BRAF amplifications in overcoming targeted therapy dosage challenges. Cancer Discov. 12, 1046–1069 (2022).
Montuori, G. et al. Extrachromosomal DNA-driven oncogene dosage heterogeneity promotes rapid adaptation to therapy in MYCN-amplified cancers. Cancer Discov. 15, 2054–2077 (2025).
Zhang, J. et al. Amplification of extrachromosomal MYC paralogs shapes immunosuppressive tumor microenvironment in small cell lung cancer. Clin. Cancer Res. 31, 4529–4542 (2025).
Webb, M. G. et al. Resolving spatial subclonal genomic heterogeneity of loss of heterozygosity and extrachromosomal DNA in gliomas. Nat. Commun. 16, 5290 (2025).
Zhu, Y. et al. Oncogenic extrachromosomal DNA functions as mobile enhancers to globally amplify chromosomal transcription. Cancer Cell 39, 694–707 (2021).
Tang, W. et al. Integrated genomic and transcriptomic profiling of glioblastoma reveals ecDNA-driven heterogeneity and microenvironmental reprogramming. Cell Rep. 44, 116426 (2025).
Noorani, I. et al. Extrachromosomal DNA-driven oncogene spatial heterogeneity and evolution in glioblastoma. Cancer Discov. 15, 2078–2095 (2025).
Sun, R. et al. Landscape of extrachromosomal DNA characteristics in high-grade serous ovarian cancer via long-read sequencing. Cell Rep. 44, 116343 (2025).
Cox, D., Yuncken, C. & Spriggs, A. I. Minute chromatin bodies in malignant tumours of childhood. Lancet 7402, 55–58 (1965).
Kim, H. et al. Extrachromosomal DNA is associated with oncogene amplification and poor outcome across multiple cancers. Nat. Genet. 52, 891–897 (2020).
Kim, H. et al. Mapping extrachromosomal DNA amplifications during cancer progression. Nat. Genet. 56, 2447–2454 (2024).
Bailey, C. et al. Origins and impact of extrachromosomal DNA. Nature 635, 193–200 (2024). In this study, the pan-cancer landscape of ecDNA is mapped, revealing tissue-specific selection, immunomodulatory functions and major clinical implications including associations with therapy exposure, metastasis and poor survival.
Wu, S. et al. Circular ecDNA promotes accessible chromatin and high oncogene expression. Nature 575, 699–703 (2019).
Hung, K. L. et al. ecDNA hubs drive cooperative intermolecular oncogene expression. Nature 600, 731–736 (2021).
Hung, K. L., Mischel, P. S. & Chang, H. Y. Gene regulation on extrachromosomal DNA. Nat. Struct. Mol. Biol. 29, 736–744 (2022).
Bergstrom, E. N. et al. Mapping clustered mutations in cancer reveals APOBEC3 mutagenesis of ecDNA. Nature 602, 510–517 (2022).
Lange, J. T. et al. The evolutionary dynamics of extrachromosomal DNA in human cancers. Nat. Genet. 54, 1527–1533 (2022).
Li, Y. et al. A 41-year-old male with multifocal “grape-like” ureteral carcinoma involving nearly the entire ureter: a case report and management discussion. BMC Urol. https://doi.org/10.1186/s12894-026-02073-w (2026).
Luebeck, J. et al. Extrachromosomal DNA in the cancerous transformation of Barrett’s oesophagus. Nature 616, 798–805 (2023).
Lv, W. et al. Extrachromosomal circular DNA orchestrates genome heterogeneity in urothelial bladder carcinoma. Theranostics 14, 5102–5122 (2024). This article provides the first evidence that ecDNA can be detected in the urine of patients with urothelial carcinoma, supporting its potential as a non-invasive biomarker.
Vandekerkhove, G. et al. ERBB2/HER2 alterations in ctDNA and metachronous tissues of patients with metastatic urothelial cancer. Clin. Cancer Res. 31, 3725–3741 (2025). In this study, plasma ctDNA is shown to be able to sensitively capture ecDNA-derived ERBB2 amplification, highlighting ctDNA as a potential non-invasive tool for monitoring ecDNA-driven heterogeneity.
Choudhury, M. et al. ecPath detects ecDNA in tumors from histopathology images. Preprint at bioRxiv https://doi.org/10.1101/2024.11.13.623494 (2024). This article introduces ecPath, a deep-learning framework that enables scalable ecDNA detection directly from routine histology images.
Tang, J. et al. Enhancing transcription-replication conflict targets ecDNA-positive cancers. Nature 635, 210–218 (2024). Transcription–replication conflict is identified as a selective vulnerability of ecDNA-containing tumours and shows that CHK1 inhibition can preferentially eliminate ecDNA-driven cancers in this study.
Nakagawa, T. et al. Inhibition of human-HPV hybrid ecDNA enhancers reduces oncogene expression and tumor growth in oropharyngeal cancer. Nat. Commun. 16, 2964 (2025).
Qin, L. N. et al. Extrachromosomal DNA biogenesis is dependent on DNA looping and religation by YY1-Lig3-PARylation complex. Mol. Cell 85, 3090–3107 (2025).
Yi, E., Chamorro Gonzalez, R., Henssen, A. G. & Verhaak, R. G. W. Extrachromosomal DNA amplifications in cancer. Nat. Rev. Genet. 23, 760–771 (2022).
Yang, Q. L., Xie, Y., Qiao, K., Lim, J. Y. S. & Wu, S. Modern biology of extrachromosomal DNA: a decade-long voyage of discovery. Cell Res. 35, 11–22 (2025).
Yan, X., Mischel, P. & Chang, H. Extrachromosomal DNA in cancer. Nat. Rev. Cancer 24, 261–273 (2024).
Haber, D. A. & Schimke, R. T. Unstable amplification of an altered dihydrofolate reductase gene associated with double-minute chromosomes. Cell 26, 355–362 (1981).
Kaufman, R. J., Brown, P. C. & Schimke, R. T. Amplified dihydrofolate reductase genes in unstably methotrexate-resistant cells are associated with double minute chromosomes. Proc. Natl Acad. Sci. USA 76, 5669–5673 (1979).
Schwab, M., Alitalo, K., Varmus, H. E., Bishop, J. M. & George, D. A cellular oncogene (c-Ki-ras) is amplified, overexpressed, and located within karyotypic abnormalities in mouse adrenocortical tumour cells. Nature 303, 497–501 (1983).
Weiser, N. E., Watkins, T. B. K., Chang, H. Y. & Mischel, P. S. A guide to extrachromosomal DNA: cancer’s dynamic circular genome. Cancer Discov. 15, 1105–1114 (2025).
Noer, J. B., Horsdal, O. K., Xiang, X., Luo, Y. & Regenberg, B. Extrachromosomal circular DNA in cancer: history, current knowledge, and methods. Trends Genet. 38, 766–781 (2022).
Lv, W. et al. Circle-Seq reveals genomic and disease-specific hallmarks in urinary cell-free extrachromosomal circular DNAs. Clin. Transl. Med. 12, e817 (2022).
Moller, H. D. et al. Circular DNA elements of chromosomal origin are common in healthy human somatic tissue. Nat. Commun. 9, 1069 (2018).
Kumar, P. et al. Normal and cancerous tissues release extrachromosomal circular DNA (eccDNA) into the circulation. Mol. Cancer Res. 15, 1197–1205 (2017).
Xu, Z. et al. Plasma extrachromosomal circular DNA is a pathophysiological hallmark of short-term intensive insulin therapy for type 2 diabetes. Clin. Transl. Med. 13, e1437 (2023).
Feng, W. et al. Targeted removal of mitochondrial DNA from mouse and human extrachromosomal circular DNA with CRISPR-Cas9. Comput. Struct. Biotechnol. J. 20, 3059–3067 (2022).
Kumar, P. et al. ATAC-seq identifies thousands of extrachromosomal circular DNA in cancer and cell lines. Sci. Adv. 6, eaba2489 (2020).
Chamorro Gonzalez, R. et al. Parallel sequencing of extrachromosomal circular DNAs and transcriptomes in single cancer cells. Nat. Genet. 55, 880–890 (2023).
Jiang, X. et al. Genome-wide characterization of extrachromosomal circular DNA in gastric cancer and its potential role in carcinogenesis and cancer progression. Cell. Mol. Life Sci. 80, 191 (2023).
Paulsen, T., Shibata, Y., Kumar, P., Dillon, L. & Dutta, A. Small extrachromosomal circular DNAs, microDNA, produce short regulatory RNAs that suppress gene expression independent of canonical promoters. Nucleic Acids Res. 47, 4586–4596 (2019).
Wang, Y. et al. eccDNAs are apoptotic products with high innate immunostimulatory activity. Nature 599, 308–314 (2021).
Chen, Y. A. et al. Extrachromosomal telomere repeat DNA is linked to ALT development via cGAS-STING DNA sensing pathway. Nat. Struct. Mol. Biol. 24, 1124–1131 (2017).
Cortes-Ciriano, I. et al. Comprehensive analysis of chromothripsis in 2,658 human cancers using whole-genome sequencing. Nat. Genet. 52, 331–341 (2020).
Helmsauer, K. et al. Enhancer hijacking determines extrachromosomal circular MYCN amplicon architecture in neuroblastoma. Nat. Commun. 11, 5823 (2020).
Koche, R. P. et al. Extrachromosomal circular DNA drives oncogenic genome remodeling in neuroblastoma. Nat. Genet. 52, 29–34 (2020).
Nathanson, D. A. et al. Targeted therapy resistance mediated by dynamic regulation of extrachromosomal mutant EGFR DNA. Science 343, 72–76 (2014).
Kang, X. et al. Extrachromosomal DNA replication and maintenance couple with DNA damage pathway in tumors. Cell 188, 3405–3421 (2025). The results of this study demonstrate that ecDNA replication activates ATM-dependent DNA damage responses and relies on alt-NHEJ repair, revealing essential pathways that maintain ecDNA and potential therapeutic vulnerabilities.
Jaworski, J. J. et al. ecDNA replication is disorganized and vulnerable to replication stress. Nucleic Acids Res. 53, gkaf711 (2025).
Nichols, A. et al. Chromosomal tethering and mitotic transcription promote ecDNA nuclear inheritance. Mol. Cell 85, 2839–2853 (2025).
Rose, J. C. et al. Disparate pathways for extrachromosomal DNA biogenesis and genomic DNA repair. Cancer Discov. 15, 69–82 (2025).
Moller, H. D. et al. CRISPR-C: circularization of genes and chromosome by CRISPR in human cells. Nucleic Acids Res. 46, e131 (2018).
Stephens, P. J. et al. Massive genomic rearrangement acquired in a single catastrophic event during cancer development. Cell 144, 27–40 (2011).
Engel, J. L. et al. The Fanconi anemia pathway induces chromothripsis and ecDNA-driven cancer drug resistance. Cell 187, 6055–6070 (2024).
Rosswog, C. et al. Chromothripsis followed by circular recombination drives oncogene amplification in human cancer. Nat. Genet. 53, 1673–1685 (2021).
Shoshani, O. et al. Chromothripsis drives the evolution of gene amplification in cancer. Nature 591, 137–141 (2021).
Coquelle, A., Rozier, L., Dutrillaux, B. & Debatisse, M. Induction of multiple double-strand breaks within an hsr by meganucleaseI-SceI expression or fragile site activation leads to formation of double minutes and other chromosomal rearrangements. Oncogene 21, 7671–7679 (2002).
Hackett, J. A., Feldser, D. M. & Greider, C. W. Telomere dysfunction increases mutation rate and genomic instability. Cell 106, 275–286 (2001).
Cleal, K. & Baird, D. M. Catastrophic endgames: emerging mechanisms of telomere-driven genomic instability. Trends Genet. 36, 347–359 (2020).
Kraft, K. et al. Enhancer activation from transposable elements in extrachromosomal DNA. Nat. Cell Biol. 27, 1914–1924 (2025). The results of this study show that transposable-element-derived sequences can be reactivated and co-opted as functional enhancers on ecDNA, revealing a mechanism by which ecDNA reshapes oncogene regulation and tumour evolution.
Whongsiri, P. et al. Many different LINE-1 retroelements are activated in bladder cancer. Int. J. Mol. Sci. 21, 9433 (2020).
Xu, K. et al. Structure and evolution of double minutes in diagnosis and relapse brain tumors. Acta Neuropathol. 137, 123–137 (2019).
Yi, E. et al. Selective depletion of cancer cells with extrachromosomal DNA via lentiviral infection. Cancer Res. Commun. 5, 1458–1465 (2025).
Kaufman, R. J., Brown, P. C. & Schimke, R. T. Loss and stabilization of amplified dihydrofolate reductase genes in mouse sarcoma S-180 cell lines. Mol. Cell Biol. 1, 1084–1093 (1981).
Alitalo, K., Schwab, M., Lin, C. C., Varmus, H. E. & Bishop, J. M. Homogeneously staining chromosomal regions contain amplified copies of an abundantly expressed cellular oncogene (c-myc) in malignant neuroendocrine cells from a human colon carcinoma. Proc. Natl Acad. Sci. USA 80, 1707–1711 (1983).
Beverley, S. M., Coderre, J. A., Santi, D. V. & Schimke, R. T. Unstable DNA amplifications in methotrexate-resistant Leishmania consist of extrachromosomal circles which relocalize during stabilization. Cell 38, 431–439 (1984).
Turner, K. M. et al. Extrachromosomal oncogene amplification drives tumour evolution and genetic heterogeneity. Nature 543, 122–125 (2017).
Taghbalout, A. et al. Extrachromosomal DNA associates with nuclear condensates and reorganizes chromatin structures to enhance oncogenic transcription. Cancer Cell 43, 2191–2205 (2025).
Purshouse, K. et al. Oncogene expression from extrachromosomal DNA is driven by copy number amplification and does not require spatial clustering in glioblastoma stem cells. eLife 11, e80207 (2022).
Hung, K. L. et al. Coordinated inheritance of extrachromosomal DNAs in cancer cells. Nature 635, 201–209 (2024).
Deshpande, V. et al. Exploring the landscape of focal amplifications in cancer using AmpliconArchitect. Nat. Commun. 10, 392 (2019).
Luebeck, J. et al. AmpliconReconstructor integrates NGS and optical mapping to resolve the complex structures of focal amplifications. Nat. Commun. 11, 4374 (2020).
Chang, L. et al. Single-cell third-generation sequencing-based multi-omics uncovers gene expression changes governed by ecDNA and structural variants in cancer cells. Clin. Transl. Med. 13, e1351 (2023).
Moller, H. D. et al. Genome-wide purification of extrachromosomal circular DNA from eukaryotic cells. J. Vis. Exp. 4, e54239 (2016).
Hung, K. L. et al. Targeted profiling of human extrachromosomal DNA by CRISPR-CATCH. Nat. Genet. 54, 1746–1754 (2022).
Pradella, D. et al. Engineered extrachromosomal oncogene amplifications promote tumorigenesis. Nature 637, 955–964 (2025).
Levan, A. & Levan, G. Have double minutes functioning centromeres? Hereditas 88, 81–92 (1978).
Chapman, O. S. et al. Circular extrachromosomal DNA promotes tumor heterogeneity in high-risk medulloblastoma. Nat. Genet. 55, 2189–2199 (2023).
Kamoun, A. et al. A consensus molecular classification of muscle-invasive bladder cancer. Eur. Urol. 77, 420–433 (2020).
Hayashi, Y. et al. TERT C228T mutation in non-malignant bladder urothelium is associated with intravesical recurrence for patients with non-muscle invasive bladder cancer. Mol. Oncol. 14, 2375–2383 (2020).
Audenet, F. et al. Clonal relatedness and mutational differences between upper tract and bladder urothelial carcinoma. Clin. Cancer Res. 25, 967–976 (2019).
Wu, T. et al. Extrachromosomal DNA formation enables tumor immune escape potentially through regulating antigen presentation gene expression. Sci. Rep. 12, 3590 (2022).
Lin, M. S. et al. Transcriptional immune suppression and up-regulation of double-stranded DNA damage and repair repertoires in ecDNA-containing tumors. eLife 12, RP88895 (2024).
Rose, K. M. et al. Circulating and urinary tumour DNA in urothelial carcinoma-upper tract, lower tract and metastatic disease. Nat. Rev. Urol. 20, 406–419 (2023).
Pongor, L. S. et al. Extrachromosomal DNA amplification contributes to small cell lung cancer heterogeneity and is associated with worse outcomes. Cancer Discov. 13, 928–949 (2023).
Marra, A. et al. Artificial intelligence entering the pathology arena in oncology: current applications and future perspectives. Ann. Oncol. 36, 712–725 (2025).
Zack, T. I. et al. Pan-cancer patterns of somatic copy number alteration. Nat. Genet. 45, 1134–1140 (2013).
Xue, Y. et al. An approach to suppress the evolution of resistance in BRAF(V600E)-mutant cancer. Nat. Med. 23, 929–937 (2017).
Aoki, Y. et al. Clinical significance of extrachromosomal DNA in FGFR2-amplified gastric cancer: therapeutic implications from clinical experience. JCO Precis. Oncol. 9, e2500033 (2025).
Canute, G. W. et al. Hydroxyurea accelerates the loss of epidermal growth factor receptor genes amplified as double-minute chromosomes in human glioblastoma multiforme. Neurosurgery 39, 976–983 (1996).
Bei, Y. et al. Passenger gene coamplifications create collateral therapeutic vulnerabilities in cancer. Cancer Discov. 14, 492–507 (2024).
Barretina, J. et al. The cancer cell line encyclopedia enables predictive modelling of anticancer drug sensitivity. Nature 483, 603–607 (2012).
Fessler, J. et al. CytoCellDB: a comprehensive resource for exploring extrachromosomal DNA in cancer cell lines. NAR. Cancer 6, zcae035 (2024).
Zhao, H. et al. Patient-derived bladder cancer organoids as a valuable tool for understanding tumor biology and developing personalized treatment. Adv. Sci. 12, e2414558 (2025).
Giurgiu, M. et al. Reconstructing extrachromosomal DNA structural heterogeneity from long-read sequencing data using Decoil. Genome Res. 34, 1355–1364 (2024).
Li, F. et al. FLED: a full-length eccDNA detector for long-reads sequencing data. Brief. Bioinform. 24, bbad388 (2023).
Rajkumar, U. et al. EcSeg: semantic segmentation of metaphase images containing extrachromosomal DNA. iScience 21, 428–435 (2019).
Acknowledgements
The authors thank all members of the W.L. Lab for their constructive suggestions.
Author information
Authors and Affiliations
Contributions
C. Li, Z.H., W.Z., Y.J., C. Lin and W.L. researched data for the article. C. Li, Z.H., W.Z., F.M., Y.J. and W.L. wrote the article. All authors made substantial contributions to discussions of the content and reviewed and edited the manuscript before submission.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Reviews Urology thanks Gabriel Malouf, Wenxuan Cheng, Yujiro Hayashi and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, C., Hu, Z., Zhang, W. et al. Extrachromosomal DNA in urothelial carcinoma: mechanisms and clinical applications. Nat Rev Urol (2026). https://doi.org/10.1038/s41585-026-01134-x
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41585-026-01134-x