Abstract
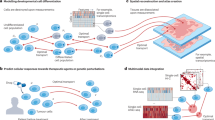
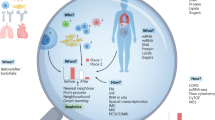
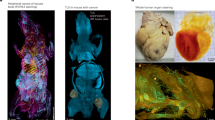
Cells have distinct molecular repertoires on their surfaces and unique intracellular biomolecular profiles that play pivotal roles in orchestrating a myriad of biological responses in the context of growth, development and disease. A persistent challenge in the deep exploration of these cues has been in our inability to effectively and precisely capture the temporal and spatial characteristics of living cells. In this Perspective, we delve into techniques for temporal and two- and three-dimensional spatial omics analyses and underscore how their harmonious fusion promises to unlock insights into the dynamics and diversity of individual cells within biological systems such as tissues and organoids. We then explore four-dimensional profiling, a nascent but promising frontier that adds a temporal (fourth-dimension) component to three-dimensional omics; highlight the advancements, challenges and gaps in the field; and discuss potential strategies for further technological development.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Vandereyken, K., Sifrim, A., Thienpont, B. & Voet, T. Methods and applications for single-cell and spatial multi-omics. Nat. Rev. Genet. 24, 494–515 (2023).
Deng, Y., Bai, Z. & Fan, R. Microtechnologies for single-cell and spatial multi-omics. Nat. Rev. Bioeng. 1, 769–784 (2023).
Ståhl, P. L. et al. Visualization and analysis of gene expression in tissue sections by spatial transcriptomics. Science 353, 78–82 (2016).
Maynard, K. R. et al. Transcriptome-scale spatial gene expression in the human dorsolateral prefrontal cortex. Nat. Neurosci. 24, 425–436 (2021).
Bressan, D., Battistoni, G. & Hannon, G. J. The dawn of spatial omics. Science 381, eabq4964 (2023).
Wahle, P. et al. Multimodal spatiotemporal phenotyping of human retinal organoid development. Nat. Biotechnol. 41, 1765–1775 (2023). An informative study that created comprehensive protein maps of retinal organoid development and primary human retinal tissue.
Li, X. et al. Profiling spatiotemporal gene expression of the developing human spinal cord and implications for ependymoma origin. Nat. Neurosci. 26, 891–901 (2023).
Chen, A. et al. Spatiotemporal transcriptomic atlas of mouse organogenesis using DNA nanoball-patterned arrays. Cell 185, 1777–1792 (2022). A comprehensive study that uses spatial enhanced resolution omics-sequencing for creating mouse organogenesis spatiotemporal transcriptomic atlas.
Chen, W. et al. Live-seq enables temporal transcriptomic recording of single cells. Nature 608, 733–740 (2022). An informative study that demonstrates one of the first concepts of live single-cell RNA-seq.
Guillaume-Gentil, O. et al. Tunable single-cell extraction for molecular analyses. Cell 166, 506–516 (2016).
Ansaryan, S. et al. High-throughput spatiotemporal monitoring of single-cell secretions via plasmonic microwell arrays. Nat. Biomed. Eng. 7, 943–958 (2023).
Lee, S., de Rutte, J., Dimatteo, R., Koo, D. & Di Carlo, D. Scalable fabrication and use of 3D structured microparticles spatially functionalized with biomolecules. ACS Nano 16, 38–49 (2022). A comprehensive study that uses a combination of hydrogels and flow cytometry for capturing and measuring single-cell secretions in high throughput.
Lin, D. et al. Time-tagged ticker tapes for intracellular recordings. Nat. Biotechnol. 41, 631–639 (2023). An informative study that bioengineers protein fibers designed to incorporate a variety of fluorescent markers during its growth, creating a ticker tape-like record of events.
Tang, F. et al. mRNA-seq whole-transcriptome analysis of a single cell. Nat. Methods 6, 377–382 (2009).
Ramsköld, D. et al. Full-length mRNA-seq from single-cell levels of RNA and individual circulating tumor cells. Nat. Biotechnol. 30, 777–782 (2012).
Macosko, E. Z. et al. Highly parallel genome-wide expression profiling of individual cells using nanoliter droplets. Cell 161, 1202–1214 (2015).
Klein, A. M. et al. Droplet barcoding for single-cell transcriptomics applied to embryonic stem cells. Cell 161, 1187–1201 (2015).
Fan, H. C., Fu, G. K. & Fodor, S. P. Expression profiling. Combinatorial labeling of single cells for gene expression cytometry. Science 347, 1258367 (2015).
Bose, S. et al. Scalable microfluidics for single-cell RNA printing and sequencing. Genome Biol. 16, 120 (2015).
Han, Q. et al. Polyfunctional responses by human T cells result from sequential release of cytokines. Proc. Natl Acad. Sci. USA 109, 1607–1612 (2012).
Junkin, M. et al. High-content quantification of single-cell immune dynamics. Cell Rep. 15, 411–422 (2016).
Wu, T., Womersley, H. J., Wang, J. R., Scolnick, J. & Cheow, L. F. Time-resolved assessment of single-cell protein secretion by sequencing. Nat. Methods 20, 723–734 (2023).
Kirschenbaum, D. et al. Time-resolved single-cell transcriptomics defines immune trajectories in glioblastoma. Cell 187, 149–165 (2024).
Wang, W. et al. Live-cell imaging and analysis reveal cell phenotypic transition dynamics inherently missing in snapshot data. Sci. Adv. 6, eaba9319 (2020).
Genshaft, A. S. et al. Live cell tagging tracking and isolation for spatial transcriptomics using photoactivatable cell dyes. Nat. Commun. 12, 4995 (2021).
Shi, L. et al. Optical imaging of metabolic dynamics in animals. Nat. Commun. 9, 2995 (2018).
Erhard, F. et al. Time-resolved single-cell RNA-seq using metabolic RNA labelling. Nat. Rev. Methods Primers 2, 77 (2022).
Perrin, J. et al. Cell tracking in cancer immunotherapy. Front. Med. 7, 34 (2020).
Trapnell, C. et al. The dynamics and regulators of cell fate decisions are revealed by pseudotemporal ordering of single cells. Nat. Biotechnol. 32, 381–386 (2014).
van Dijk, D. et al. Recovering gene interactions from single-cell data using data diffusion. Cell 174, 716–729 (2018).
La Manno, G. et al. RNA velocity of single cells. Nature 560, 494–498 (2018).
Lange, M. et al. CellRank for directed single-cell fate mapping. Nat. Methods 19, 159–170 (2022).
Sashittal, P. et al. Inferring cell differentiation maps from lineage tracing data. Preprint at bioRxiv https://doi.org/10.1101/2024.09.09.611835 (2024).
Ding, J., Sharon, N. & Bar-Joseph, Z. Temporal modelling using single-cell transcriptomics. Nat. Rev. Genet. 23, 355–368 (2022).
Shroff, H., Testa, I., Jug, F. & Manley, S. Live-cell imaging powered by computation. Nat. Rev. Mol. Cell Biol. 25, 443–463 (2024).
Femino, A. M., Fay, F. S., Fogarty, K. & Singer, R. H. Visualization of single RNA transcripts in situ. Science 280, 585–590 (1998).
Nilsson, M. et al. Padlock probes: circularizing oligonucleotides for localized DNA detection. Science 265, 2085–2088 (1994).
Ke, R. et al. In situ sequencing for RNA analysis in preserved tissue and cells. Nat. Methods 10, 857–860 (2013).
Rodriques, S. G. et al. Slide-seq: a scalable technology for measuring genome-wide expression at high spatial resolution. Science 363, 1463–1467 (2019).
Liu, Y. et al. High-spatial-resolution multi-omics sequencing via deterministic barcoding in tissue. Cell 183, 1665–1681 (2020).
Gerdes, M. J. et al. Highly multiplexed single-cell analysis of formalin-fixed, paraffin-embedded cancer tissue. Proc. Natl Acad. Sci. USA 110, 11982–11987 (2013).
Wang, X. et al. Three-dimensional intact-tissue sequencing of single-cell transcriptional states. Science 361, eaat5691 (2018).
Fu, X. et al. Polony gels enable amplifiable DNA stamping and spatial transcriptomics of chronic pain. Cell 185, 4621–4633 (2022).
Lin, J. R. et al. Multiplexed 3D atlas of state transitions and immune interaction in colorectal cancer. Cell 186, 363–381 (2023).
Legnini, I. et al. Spatiotemporal, optogenetic control of gene expression in organoids. Nat. Methods 20, 1544–1552 (2023).
Lundin, E. et al. Spatiotemporal mapping of RNA editing in the developing mouse brain using in situ sequencing reveals regional and cell-type-specific regulation. BMC Biol. 18, 6 (2020).
Chow, K.-H. K. et al. Imaging cell lineage with a synthetic digital recording system. Science 372, eabb3099 (2021). An informative study that develops a synthetic digital recording system to simultaneously evaluate gene expression, single-cell lineage, and spatial organization within the same tissue.
Ren, J. et al. Spatiotemporally resolved transcriptomics reveals the subcellular RNA kinetic landscape. Nat. Methods 20, 695–705 (2023). A comprehensive study that uses metabolic labeling and multiplexed 3D ISS to profile the age and spatial location of individual RNA molecules.
Ratz, M. et al. Clonal relations in the mouse brain revealed by single-cell and spatial transcriptomics. Nat. Neurosci. 25, 285–294 (2022). An informative study integrating spatial transcriptomics and clonal barcoding to reconstruct cell phenotypes and lineage relationships at both the single-cell and tissue scales.
Sampath Kumar, A. et al. Spatiotemporal transcriptomic maps of whole mouse embryos at the onset of organogenesis. Nat. Genet. 55, 1176–1185 (2023).
Wang, M. et al. High-resolution 3D spatiotemporal transcriptomic maps of developing Drosophila embryos and larvae. Dev. Cell 57, 1271–1283 (2022).
Mahdessian, D. et al. Spatiotemporal dissection of the cell cycle with single-cell proteogenomics. Nature 590, 649–654 (2021).
Kolb, H. C., Finn, M. G. & Sharpless, K. B. Click chemistry: diverse chemical function from a few good reactions. Angew. Chem. Int. Ed. Engl. 40, 2004–2021 (2001).
Scinto, S. L. et al. Bioorthogonal chemistry. Nat. Rev. Methods Primers 1, 30 (2021).
Ko, J. et al. In vivo click chemistry enables multiplexed intravital microscopy. Adv. Sci. 9, e2200064 (2022).
Ko, J., Oh, J., Ahmed, M. S., Carlson, J. C. T. & Weissleder, R. Ultra-fast cycling for multiplexed cellular fluorescence imaging. Angew. Chem. Int. Ed. Engl. 59, 6839–6846 (2020).
Carlson, J. C. T., Mikula, H. & Weissleder, R. Unraveling tetrazine-triggered bioorthogonal elimination enables chemical tools for ultrafast release and universal cleavage. J. Am. Chem. Soc. 140, 3603–3612 (2018).
Reynolds, D. E. et al. Live organoid cyclic imaging. Adv. Sci. 11, e2309289 (2024).
Ko, J. et al. Spatiotemporal multiplexed immunofluorescence imaging of living cells and tissues with bioorthogonal cycling of fluorescent probes. Nat. Biotechnol. 40, 1654–1662 (2022). A comprehensive study that applies bioorthogonal click chemistry for capturing spatiotemporal patterns in living cells and tissues.
Wang, F. et al. A light-initiated chemical reporter strategy for spatiotemporal labeling of biomolecules. RSC Chem. Biol. 3, 539–545 (2022).
Liu, Y. et al. Spatiotemporally resolved subcellular phosphoproteomics. Proc. Natl Acad. Sci. USA 118, e2025299118 (2021).
Das, K. K., Basu, B., Maiti, P. & Dubey, A. K. Piezoelectric nanogenerators for self-powered wearable and implantable bioelectronic devices. Acta Biomater. 171, 85–113 (2023).
Zhao, S. et al. Tracking neural activity from the same cells during the entire adult life of mice. Nat. Neurosci. 26, 696–710 (2023). An informative study utilizing bioelectronic mesh networks to record single-unit action potentials from the same neurons throughout the entire lifespan of adult mice.
Li, Q. et al. Multimodal charting of molecular and functional cell states via in situ electro-sequencing. Cell 186, 2002–2017 (2023). A comprehensive study that integrates flexible bioelectronics with spatial transcriptomics for mapping electrical activity and profiling single-cell expression.
Yang, X. et al. Kirigami electronics for long-term electrophysiological recording of human neural organoids and assembloids. Nat. Biotechnol. 42, 1836–1843 (2024).
Le Floch, P. et al. Stretchable mesh nanoelectronics for 3D single-cell chronic electrophysiology from developing brain organoids. Adv. Mater. 34, e2106829 (2022).
Trujillo, C. A. et al. Complex oscillatory waves emerging from cortical organoids model early human brain network development. Cell Stem Cell 25, 558–569 (2019).
Sempionatto, J. R., Lasalde-Ramírez, J. A., Mahato, K., Wang, J. & Gao, W. Wearable chemical sensors for biomarker discovery in the omics era. Nat. Rev. Chem. 6, 899–915 (2022).
Lin, M., Hu, H., Zhou, S. & Xu, S. Soft wearable devices for deep-tissue sensing. Nat. Rev. Mater. 7, 850–869 (2022).
Jahed, Z. et al. Nanocrown electrodes for parallel and robust intracellular recording of cardiomyocytes. Nat. Commun. 13, 2253 (2022).
Ren, H., Walker, B. L., Cang, Z. & Nie, Q. Identifying multicellular spatiotemporal organization of cells with SpaceFlow. Nat. Commun. 13, 4076 (2022).
Pham, D. et al. Robust mapping of spatiotemporal trajectories and cell-cell interactions in healthy and diseased tissues. Nat. Commun. 14, 7739 (2023).
Lefebvre, M. et al. Learning a conserved mechanism for early neuroectoderm morphogenesis. Preprint at https://arxiv.org/html/2405.18382v1 (2024).
Velten, B. & Stegle, O. Principles and challenges of modeling temporal and spatial omics data. Nat. Methods 20, 1462–1474 (2023).
Depaoli, M. R. et al. Live cell imaging of signaling and metabolic activities. Pharmacol. Ther. 202, 98–119 (2019).
Bridges, K. & Miller-Jensen, K. Mapping and validation of scRNA-seq-derived cell-cell communication networks in the tumor microenvironment. Front. Immunol. 13, 885267 (2022).
Rugg-Gunn, P. J., Moris, N. & Tam, P. P. L. Technical challenges of studying early human development. Development 150, dev201797 (2023).
Apostolou, E. et al. Progress and challenges in stem cell biology. Nat. Cell Biol. 25, 203–206 (2023).
Harrison, M. M., Jenkins, B. V., O’Connor-Giles, K. M. & Wildonger, J. A CRISPR view of development. Genes Dev. 28, 1859–1872 (2014).
Krueger, D. et al. Principles and applications of optogenetics in developmental biology. Development 146, dev175067 (2019).
Binnewies, M. et al. Understanding the tumor immune microenvironment (TIME) for effective therapy. Nat. Med. 24, 541–550 (2018).
Wareham, L. K. et al. Solving neurodegeneration: common mechanisms and strategies for new treatments. Mol. Neurodegener. 17, 23 (2022).
Runa, F. et al. Tumor microenvironment heterogeneity: challenges and opportunities. Curr. Mol. Biol. Rep. 3, 218–229 (2017).
van Driel, M. A. & Brunner, H. G. Bioinformatics methods for identifying candidate disease genes. Hum. Genomics 2, 429–432 (2006).
Chen, C. et al. Applications of multi-omics analysis in human diseases. MedComm 4, e315 (2023).
Jucker, M. The benefits and limitations of animal models for translational research in neurodegenerative diseases. Nat. Med. 16, 1210–1214 (2010).
Velasco, V., Shariati, S. A. & Esfandyarpour, R. Microtechnology-based methods for organoid models. Microsyst. Nanoeng. 6, 76 (2020).
Wang, X. & Fan, J. Spatiotemporal molecular medicine: a new era of clinical and translational medicine. Clin. Transl. Med. 11, e294 (2021).
Zhang, J. et al. Spatiotemporal omics-refining the landscape of precision medicine. Life Med. 1, 84–102 (2022).
Roisman, L. C. et al. Radiological artificial intelligence - predicting personalized immunotherapy outcomes in lung cancer. NPJ Precis. Oncol. 7, 125 (2023).
Ho, D. et al. Enabling technologies for personalized and precision medicine. Trends Biotechnol. 38, 497–518 (2020).
Whitesides, G. M. The origins and the future of microfluidics. Nature 442, 368–373 (2006).
Duncombe, T. A., Tentori, A. M. & Herr, A. E. Microfluidics: reframing biological enquiry. Nat. Rev. Mol. Cell Biol. 16, 554–567 (2015).
Sharei, A. et al. A vector-free microfluidic platform for intracellular delivery. Proc. Natl Acad. Sci. USA 110, 2082–2087 (2013).
Joo, B., Hur, J., Kim, G. B., Yun, S. G. & Chung, A. J. Highly efficient transfection of human primary T lymphocytes using droplet-enabled mechanoporation. ACS Nano 15, 12888–12898 (2021).
Park, S. E., Georgescu, A. & Huh, D. Organoids-on-a-chip. Science 364, 960–965 (2019).
Baig, N., Kammakakam, I. & Falath, W. Nanomaterials: a review of synthesis methods, properties, recent progress, and challenges. Mater. Adv. 2, 1821–1871 (2021).
Harish, V. et al. Review on nanoparticles and nanostructured materials: bioimaging, biosensing, drug delivery, tissue engineering, antimicrobial, and agro-food applications. Nanomaterials 12, 457 (2022).
Kumar, S., Bhushan, P. & Bhattacharya, S. Fabrication of nanostructures with bottom-up approach and their utility in diagnostics, therapeutics, and others. Environmental, Chemical and Medical Sensors 167–198 (2018).
Garnett, E., Mai, L. & Yang, P. Introduction: 1D nanomaterials/nanowires. Chem. Rev. 119, 8955–8957 (2019).
Chattrairat, K. et al. All-in-one nanowire assay system for capture and analysis of extracellular vesicles from an ex vivo brain tumor model. ACS Nano 17, 2235–2244 (2023).
Yee Mon, K. J. et al. Functionalized nanowires for miRNA-mediated therapeutic programming of naïve T cells. Nat. Nanotechnol. 19, 1190–1202 (2024).
Xie, X. et al. Nanostraw-electroporation system for highly efficient intracellular delivery and transfection. ACS Nano 7, 4351–4358 (2013).
Cao, Y. et al. Nondestructive nanostraw intracellular sampling for longitudinal cell monitoring. Proc. Natl Acad. Sci. USA 114, E1866–E1874 (2017).
Golshadi, M., Wright, L. K., Dickerson, I. M. & Schrlau, M. G. High-efficiency gene transfection of cells through carbon nanotube arrays. Small 12, 3014–3020 (2016).
Thiruppathi, R., Mishra, S., Ganapathy, M., Padmanabhan, P. & Gulyás, B. Nanoparticle functionalization and its potentials for molecular imaging. Adv. Sci. 4, 1600279 (2017).
Cameron, D. E., Bashor, C. J. & Collins, J. J. A brief history of synthetic biology. Nat. Rev. Microbiol. 12, 381–390 (2014).
Yan, X., Liu, X., Zhao, C. & Chen, G. Q. Applications of synthetic biology in medical and pharmaceutical fields. Signal Transduct. Target Ther. 8, 199 (2023).
Kebschull, J. M. & Zador, A. M. Cellular barcoding: lineage tracing, screening and beyond. Nat. Methods 15, 871–879 (2018).
Vallot, C. RNA barcoding: the catalyst for the single-cell revolution. Nat. Rev. Genet. 24, 491 (2023).
Lovatt, D. et al. Transcriptome in vivo analysis (TIVA) of spatially defined single cells in live tissue. Nat. Methods 11, 190–196 (2014).
Horns, F. et al. Engineering RNA export for measurement and manipulation of living cells. Cell 186, 3642–3658 (2023).
Metzcar, J., Wang, Y., Heiland, R. & Macklin, P. A review of cell-based computational modeling in cancer biology. JCO Clin. Cancer Inf. 3, 1–13 (2019).
Walpole, J., Papin, J. A. & Peirce, S. M. Multiscale computational models of complex biological systems. Annu. Rev. Biomed. Eng. 15, 137–154 (2013).
Tuszynski, J. A. et al. Mathematical and computational modeling in biology at multiple scales. Theor. Biol. Med Model 11, 52 (2014).
Alber, M. et al. Integrating machine learning and multiscale modeling-perspectives, challenges, and opportunities in the biological, biomedical, and behavioral sciences. NPJ Digit. Med. 2, 115 (2019).
Acknowledgements
This work was supported by the NSF Graduate Research Fellowship Program to D.E.R. and National Institutes of Health award R00CA256353 to J.K.
Author information
Authors and Affiliations
Contributions
D.E.R. led the writing and editing of the paper and created the figures. Y.H.R., D.O. and P.V. contributed to the writing and editing. R.F. provided feedback and assisted with revisions. J.K. supervised and reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
R.F. is a scientific founder and adviser for IsoPlexis, Singleron Biotechnologies and AtlasXomics. J.K. is a scientific cofounder and adviser for Aperture Bio. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Methods thanks Guangdun Peng and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Madhura Mukhopadhyay, in collaboration with the Nature Methods team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Reynolds, D.E., Roh, Y.H., Oh, D. et al. Temporal and spatial omics technologies for 4D profiling. Nat Methods 22, 1408–1419 (2025). https://doi.org/10.1038/s41592-025-02683-6
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41592-025-02683-6
This article is cited by
-
Interpretation, extrapolation and perturbation of single cells
Nature Reviews Genetics (2026)
-
Spatial omics in 3D culture model systems: decoding cellular positioning mechanisms and microenvironmental dynamics
Journal of Translational Medicine (2025)