Abstract
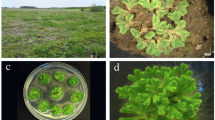
Bischofia polycarpa (2n = 68), belonging to Phyllanthaceae family, is a native deciduous tree with naturally distribution ranging from southern Qinling Mountains and Huaihe River basin to the northern regions of Fujian and Guangdong, China. It holds significant horticultural, ornamental, and medicinal value and serves as a crucial winter food resource for wild birds. Herein, we report a de novo genome assembly for B. polycarpa, utilizing a combination of PacBio HiFi Reads and Hi-C data. In total, the genome size reaches 585.68 Mb with a contig N50 of 12.62 Mb, and 99.06% (580.18 Mb) of the assembly successfully anchored on 34 chromosomes. The genome comprises approximately 62.77% repetitive sequences and 32,554 protein-coding genes, of which 96.15% could be functionally annotated. The BUSCO analysis reveals a genome completeness of 95.42% (n = 1,540), including 1,499 (92.87%) single-copy BUSCOs and 41 (2.54%) duplicated BUSCOs. This high-quality genome of the Phyllanthaceae enriches our understanding of the genetic underpinnings of plant reproductive ecology.
Similar content being viewed by others
Data availability
The finalized chromosome assembly were deposited in NCBI GenBank under BioProject (PRJNA1267844) with accession number GCA_053574235.1. RNA-seq data from various tissues are accessible under the BioProject (PRJNA1365770) with accession numbers SRR36186603. The genome annotation files (GFF3, GTF, FASTA) were available in the Figshare database. All datasets are publicly available without restriction.
Code availability
All sofware and pipelines were executed in strict accordance with the manuals and protocols provided by the published bioinformatics tools. No custom programming or coding was used.
References
Webster, G. L. Synopsis of the genera and suprageneric taxa of Euphorbiaceae. Annals of the Missouri Botanical Garden 81(1), 33–144 (1994).
GROUP TAP: An update of the Angiosperm Phylogeny Group classification for the orders and families of flowering plants: APG II. Botanical Journal of the Linnean Society 141, 399–436 (2003).
Kawakita, A. & Kato M. Diversity of Phyllanthaceae plants. Obligate pollination mutualism 81–115 (2017).
Mazumdar, A. B. & Chattopadhyay, S. Sequencing, de novo assembly, functional annotation and analysis of Phyllanthus amarus leaf transcriptome using the Illumina Platform. Frontiers in Plant Science 6, 1199 (2016).
Ahmad, B. et al. Phyllanthus emblica: a comprehensive review of its therapeutic benefits. South African Journal of Botany 138(1), 278–310 (2021).
Rani, N. Z. A. et al. Mechanistic studies of the antiallergic activity of Phyllanthus amarus Schum. & Thonn. and its compounds. Molecules 26(3), 695 (2021).
Zhang, W. T. et al. The first high-quality chromosome-level genome assembly of Phyllanthaceae (Phyllanthus cochinchinensis) provides insights into flavonoid biosynthesis. Planta 256(6), 109 (2022).
Xia, F. G. et al. Polyploid genome assembly provides insights into morphological development and ascorbic acid accumulation of Sauropus androgynus. International journal of molecular sciences 25(1), 300 (2024).
Li, F. et al. Haplotype-resolved genomes of octoploid species in Phyllanthaceae family reveal a critical role for polyploidization and hybridization in speciation. The Plant Journal 119(1), 348–363 (2024).
Huang, J. et al. Genome assembly provides insights into the genome evolution of Baccaurea ramiflora Lour. Scientific Reports 14(1), 4867 (2024).
Chen, B.-Z. et al. Chromosome-level genome assembly and annotation of Flueggea virosa (Phyllanthaceae). Scientific Data 11(1), 875 (2024).
Wannamethee, S. G. et al. Serum conjugated linoleic acid and risk of incident heart failure in older men: the British Regional heart study. Journal of the American Heart Association 7, e006653 (2018).
Allen, G. C., Flores-Vergara, M., Krasnyanski, K. & Thompson, W. A modified protocol for rapid DNA isolation from plant tissues using cetyltrimethylammonium bromide. Nature protocols 1(5), 2320–2325 (2006).
Chen, S., Zhou, Y., Chen, Y. & Gu, J. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34(17), i884–i890 (2018).
Marçais, G. & Kingsford, C. A fast, lock-free approach for efficient parallel counting of occurrences of k-mers. Bioinformatics 27(6), 764–770 (2011).
Ranallo-Benavidez, T. R., Jaron, K. S. & Schatz, M. C. GenomeScope 2.0 and Smudgeplot for reference-free profiling of polyploid genomes. Nature communications 11(1), 1432 (2020).
He, Z., Zhang, W., Luo X. & Huan, J. Five Fabaceae Karyotype and Phylogenetic Relationship Analysis Based on Oligo-FISH for 5S rDNA and (AG3T3)3. Genes. (Basel) 13(5), 768 (2022).
Servant, N. et al. HiC-Pro: an optimized and flexible pipeline for Hi-C data processing. Genome biology 16, 1–11 (2015).
Cheng, H., Concepcion, G. T., Feng, X., Zhang, H. & Li, H. Haplotype-resolved de novo assembly using phased assembly graphs with hifiasm. Nature Methods 18(2), 170–175 (2021).
Simão, F. A., Waterhouse, R. M., Panagiotis, I., Kriventseva, E. V. & Zdobnov, E. M. BUSCO: assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics 31(19), 3210–3212 (2015).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25(14), 1754–1760 (2009).
Burton, J. N. et al. Chromosome-scale scaffolding of de novo genome assemblies based on chromatin interactions. Nature biotechnology 31(12), 1119–1125 (2013).
Robinson, J. T. et al. Juicebox.js provides a Cloud-Based Visualization System for Hi-C Data. Cell Systems 6(2), 256–258.e251 (2018).
Haas, B. J. et al. Improving the Arabidopsis genome annotation using maximal transcript alignment assemblies. Nucleic acids research 31(19), 5654–5666 (2003).
Haas, B. J. et al. De novo transcript sequence reconstruction from RNA-seq using the Trinity platform for reference generation and analysis. Nature Protocols 8(8), 1494–1512 (2013).
Hart, A. J. et al. EnTAP: Bringing faster and smarter functional annotation to non‐model eukaryotic transcriptomes. Molecular ecology resources 20(2), 591–604 (2020).
Flynn, J. M., Hubley, R., Rosen, J., Clark, A. G. & Smit, A. F. RepeatModeler2 for automated genomic discovery of transposable element families. PNAS 117(17), 9451–9457 (2020).
Bao, Z. & Eddy, S. R. Automated de novo identification of repeat sequence families in sequenced genomes. Genome Research 12(8), 1269–1276 (2002).
Price, A. L., Jones, N. C. & Pevzner, P. A. De novo identification of repeat families in large genomes. Bioinformatics 21(suppl_1), i351–i358 (2005).
Flynn, J. M. et al. AFJPotNAoS: RepeatModeler2 for automated genomic discovery of transposable element families, 117(17):9451-9457 (2020).
Hubley, R. et al. The Dfam database of repetitive DNA families. Nucleic Acids Research 44(D1), D81–D89 (2016).
Ellinghaus, D., Kurtz, S. & Willhoeft, U. LTRharvest, an efficient and flexible software for de novo detection of LTR retrotransposons. BMC Bioinformatics 9, 18 (2008).
Xu, Z. & Wang, H. LTR_FINDER: an efficient tool for the prediction of full-length LTR retrotransposons. Nucleic Acids Research 35, W265–W268 (2007).
Ou, S. & Jiang, N. LTR_retriever: A Highly Accurate and Sensitive Program for Identification of Long Terminal Repeat Retrotransposons. Plant Physiology 176(2), 1410–1422 (2018).
Tarailo‐Graovac, M. & Chen, N. S. Using RepeatMasker to identify repetitive elements in genomic sequences. Current protocols in bioinformatics 25(1), 4–10 (2009).
Benson, G. Tandem repeats finder: a program to analyze DNA sequences. Nucleic Acids Research 27(2), 573–580 (1999).
Beier, S., Thiel, T., Scholz, T. M. & Mascher, U. M MISA-web: a web server for microsatellite prediction. Bioinformatics 33(16), 2583–2585 (2017).
Stanke, M., Diekhans, M., Baertsch, R. & Haussler, D. Using native and syntenically mapped cDNA alignments to improve de novo gene finding. Bioinformatics 24(5), 637–644 (2008).
Korf, I. Gene finding in novel genomes. BMC Bioinformatics 14(5), 59 (2004).
Jens, K. et al. Using intron position conservation for homology-based gene prediction. Nucleic acids research 44(9), e89–e89 (2016).
Kim, D., Langmead, B. & Salzberg, S. L. HISAT: A fast spliced aligner with low memory requirements. Nature methods 12(4), 357–360 (2015).
Pertea, M. et al. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nature biotechnology 33(3), 290–295 (2015).
Tang, S., Lomsadze, A. & Borodovsky, M. Identification of protein coding regions in RNA transcripts. Nucleic Acids Research 43(12), e78 (2015).
Grabherr, M. G., Haas, B. J., Yassour, M. & Levin, J. Z. Trinity: reconstructing a full-length transcriptome without a genome from RNA-Seq data. Nature Biotechnology 29, 644 (2013).
Wu, T. D., Reeder, J., Lawrence, M., Becker, G. & Brauer M. J. GMAP and GSNAP for genomic sequence alignment: enhancements to speed, accuracy, and functionality. Statistical genomics: methods and protocols: 283–334 (2016).
Cantarel, B. L. et al. Yandell MJGr: MAKER: an easy-to-use annotation pipeline designed for emerging model organism genomes, 18(1), 188–196 (2008).
Haas, B. J., Salzberg, S. L., Zhu, W. & Mihaela, P. Automated eukaryotic gene structure annotation using EVidenceModeler and the program to assemble spliced alignments. Genome biology 9, 1–22 (2008).
Zhang, R.-G. et al. TEsorter: an accurate and fast method to classify LTR-retrotransposons in plant genomes. Horticulture Research 9, uhac017 (2022).
Huerta-Cepas, J. et al. eggNOG 5.0: a hierarchical, functionally and phylogenetically annotated orthology resource based on 5090 organisms and 2502 viruses. Nucleic acids research 47(D1), D309–D314 (2019).
Boeckmann, B. et al. The SWISS-PROT protein knowledgebase and its supplement TrEMBL in 2003. Nucleic Acids Research 31(1), 365–370 (2003).
Buchfink, B., Xie, C. & Huson, D. H. Fast and sensitive protein alignment using DIAMOND. Nature Methods 12(12), 59–60 (2015).
Kanehisa, M., Sato, Y., Kawashima, M. & Mao, T. KEGG as a reference resource for gene and protein annotation. Nucleic Acids Research 44(D1), D457–D462 (2015).
Jones, P. et al. InterProScan 5: genome-scale protein function classification. Bioinformatics 30(9), 1236–1240 (2014).
Finn, R. D. et al. Pfam: clans, web tools and services, 34(Database issue):D247-251 (2006).
Lowe, T. M. & Eddy, S. R. tRNAscan-SE: a program for improved detection of transfer RNA genes in genomic sequence. Nucleic acids research 25(5), 955–964 (1997).
Torkel, L. A novel method for predicting ribosomal RNA genes in prokaryotic genomes. Lund University (2017).
Nawrocki, E. P. & Eddy, S. R. Infernal 1.1: 100-fold faster RNA homology searches. Bioinformatics 29(22), 2933–2935 (2013).
Griffiths-Jones, S. et al. Rfam: annotating non-coding RNAs in complete genomes. Nucleic acids research 33(Database issue), D121–124 (2005).
She, R., Chu, J. S. C., Wang, K., Pei, J. & Chen, N. genBlastA: Enabling BLAST to identify homologous gene sequences. Genome research 19(1), 143–149 (2009).
Birney, E., Clamp, M. & Durbin, R. GeneWise and Genomewise. Frontiers in Plant Science 14(5), 988 (2004).
Partners C-NMA. Database resources of the national genomics data center, China national center for bioinformation in 2024. Nucleic acids research 52(D1), D18–D32 (2024).
Chen, M. et al. Genome Warehouse: a public repository housing genome-scale data. Genomics Proteomics Bioinformatics 19(4), 584–589 (2021).
NCBI GenBank https://identifiers.org/ncbi/insdc.gca:GCA_053574235.1 (2025).
NCBI GenBank https://identifiers.org/ncbi/insdc.sra:SRR36186603 (2025).
NCBI GenBank https://identifiers.org/ncbi/insdc.sra:SRR36589530 (2025).
Xin, G. et al. The chromosome-scale genome assembly, annotation of Bischofia polycarpa (Levl.) Airy Shaw, Phyllanthaceae. Figshare https://doi.org/10.6084/m9.figshare.27458694 (2025).
Parra, G., Bradnam, K. & Korf, I. CEGMA: a pipeline to accurately annotate core genes in eukaryotic genomes. Bioinformatics 23(9), 1061–1067 (2007).
Rhie, A., Walenz, B. P., Koren, S. & Phillippy, A. M. J. G. B. Merqury: reference-free quality, completeness, and phasing assessment for genome assemblies, 21, 1–27 (2020).
Li, H. Minimap2: pairwise alignment for nucleotide sequences. Bioinformatics 31(18), 3094–3100 (2018).
Acknowledgements
This work was supported by the Program for Young Talents of Science and Technology in Universities of Yancheng Teachers University (grant number: 206670157, and 204670012); Hunan Provincial Natural Science Foundation of China (grant number: 2024JJ5295), and General Project of Philosophy and Social Sciences in Hunan Province (grant number: 22YBA306); Key Scientific Research Projects of Hunan Provincial Education Department (grant number: 24A0751). Thanks to Professor Chenglang Pan from Minjiang University for providing the photographs of Bischofia.
Author information
Authors and Affiliations
Contributions
L. Wang and B.B. conceived and designed the study, C.Y. revised the manuscript. G.L. prepared the materials. G.L. and C.Y. analyzed the data and wrote the manuscript. G. Wang, D.Z., B.P., and B.B. edited and improved the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Xin, G., Wang, G., Liu, B. et al. The chromosome-scale genome assembly, annotation of Bischofia polycarpa (H. Lév.) Airy Shaw, Phyllanthaceae. Sci Data (2026). https://doi.org/10.1038/s41597-026-06554-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41597-026-06554-3