Abstract
Intrinsic optical signal imaging (IOSI) enables non-invasive monitoring of neural activity in the mouse cortex, yet quantitative analysis remains hindered by low signal intensity, complex spatiotemporal patterns, and the lack of standardized benchmarks. To tackle this challenge, we present MouseCortex-IOS, a novel open accessible video segmentation dataset specifically designed for standardizing IOS analysis in awake rodent models, comprising 5732 images from 14 experimental subjects. Meanwhile, we implement an efficient processing pipeline leveraging foundation models to ensure annotation consistency while minimizing manual intervention. The dataset supports quantitative characterization of neural activation parameters including signal propagation velocity and trajectory, serving as a critical benchmark for developing automated analysis tools. Furthermore, this resource facilitates technique development in neuroimaging studies and accelerates the integration of computational approaches in IOS-based neuroscience research.
Similar content being viewed by others
Background & Summary
Intrinsic optical signal imaging (IOSI) is a widely utilized functional brain imaging technique introduced in the 1980s1 and valued for its non-invasive nature, high resolution, and wide-field imaging capabilities. IOSI detects subtle changes in cortical optical reflectance generated by hemodynamic responses, which reflect neural activity within the cortex. However, the signal is highly susceptible to noise with typically intensity of only 0.1% to 6%2 of the background, presenting significant challenges for extracting reliable data on neural activity. Effective IOS image analysis requires quantitative methods to access signal strength, track propagation trajectories, determine durations, count cycles, and calculate coverage areas1.
Traditional IOSI analysis methods relied heavily on manual and statistical approaches, including event-related imaging with trial averaging and techniques like Principal Component Analysis (PCA) and Independent Component Analysis (ICA)1. While these approaches were instrumental in early studies, they often struggled with the noise introduced by factors such as subject movement, requiring complex setups and still yielding limited results. Despite advances in statistical and image-processing techniques, IOSI data remain challenging to analyze consistently, as no standardized datasets exist for developing and benchmarking automated analysis tools. This gap highlights the need for a dedicated dataset and improved methodologies that can more reliably segment signal regions, especially given IOSI’s increasing use in advanced neuroscience research3,4,5. Recent advances in AI (artificial intelligence), particularly in VOS (video object segmentation)6,7,8,9, have transformed image analysis across various fields, including medical imaging10,11,12. VOS methods capture spatial and temporal relationships in video data, making them particularly applicable to IOSI data, which consists of temporally sequenced frames showing subtle but correlated changes over time. One notable development in VOS is the SAM(Segment Anything Model)13, a foundation model trained on diverse datasets, providing broad applicability across segmentation tasks and featuring a prompt-based interface for interactive and adaptive segmentation. SAM2 (Segment Anything Model 2)14, an extension of SAM, integrates temporal consistency and motion awareness, making it well-suited for continuous object tracking tasks such as IOSI. Together, SAM and SAM2 address several limitations of traditional IOSI analysis by enabling the consistent segmentation of complex temporospatial data with minimal manual intervention. Thus, we employed SAM2 in building the MouseCortex-IOS, a novel video object segmentation dataset specifically designed for tracking and analyzing intrinsic optical signals in the rat cortex. It consisted of 5732 annotated frames from 14 experimental subjects and totally 194 signal segments.
In summary, we introduce the a new IOS dataset of mouse cortex, with detailed information provided in Table 1. For the “Name” column, the leading part of number corresponds to the experimental date of the respective mouse. The “Clips” column denotes the number of video clips acquired for each mouse, while the “Images” column represents the total number of frames collected per mouse. The column “Average and Standard deviation of clip duration” reports the mean and standard deviation of the video clip durations, respectively. The “Dataset size (GB)” column specifies the storage size of each individual dataset subset after decompression; the cumulative size of the entire MouseCortex-IOS dataset is approximately 4.363 GB.
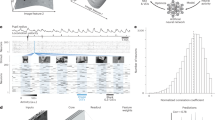
Furthermore, we have developed a pipeline for generating high-quality IOS annotations for each frame, and Fig. 1 illustrates the overview of the experimental procedures. Three stages of preprocessing convert massive indistinguishable raw grayscale images into interpretable color-coded signal maps while reducing data volume by two orders of magnitude. The preprocessed image sequences are imported into an open-sourced annotation tool, where a preloaded SAM2 model enables manual or automated annotation of prompts for the initial frames. A single-click inference operation then generates annotations for all frames. The annotated data is used for downstream tasks such as signal analysis and tracking.
Data processing framework. Overall, raw data undergoes preprocessing and subsequent processing via the SAM2 model to generate our dataset.
Method
Data collection
This dataset was collected from experimental data acquired in collaboration with the Department of Neurosurgery, Guangzhou First People’s Hospital, utilizing 14 adult C57BL6 mice (23–30 g, obtained from Guangdong Zhiyuan Biotechnology Co., Ltd.). These 14 subjects were randomly selected from a larger experimental group, covering diverse intervention groups including control conditions, potassium chloride (KCl) application, vagus nerve stimulation (VNS), and sciatic nerve stimulation (SNS). All procedures were approved by the Animal Ethics Committee of Guangzhou First People’s Hospital (approval number: K-2021-065). At the conclusion of the study, mice designated for histological analysis were deeply anesthetized and perfused, while others were euthanized via CO₂ inhalation, as specified in protocol K-2021-065. Death was confirmed by the absence of heartbeat, respiration, and pedal reflexes.
Mice were anesthetized via intraperitoneal injection of 1% sodium pentobarbital (60 mg/kg), and their rectal temperature was maintained at 37 °C throughout the experiments. Following anesthesia, cranial windows were created in the skull (under saline cooling) to facilitate subsequent stimulations. For neural modulation, either the vagus or sciatic nerve was isolated, and copper wire electrodes were applied; stimulation parameters included 0.5 mA intensity, 20 Hz frequency, 30-second stimulation trains, and 5-minute intervals, totaling 60 minutes of stimulation. Sham controls received electrode placement without electrical current. Cortical spreading depression (CSD) was induced by topical application of 1 μL of 0.125 M or 1 M KCl at the cranial window sites (administered every 20 minutes for 2 hours), starting ~10 minutes post-stimulation to allow physiological stabilization.
Cortical signals were recorded using a custom-developed intrinsic optical signal (IOS) imaging system: the field of view was 10 × 10 mm (corresponding to 512 × 512 pixels resolution), with illumination from 545/558/578 nm LEDs positioned 10 cm above the skull. Reflected light was captured by a cooled frame-transfer EM-CCD camera (Andor iXon 897) at 60 Hz, with continuous image acquisition lasting 2.5 hours after KCl administration. Raw image data was stored on external storage devices, and the 14 selected mice contributed 5732 frames to the MouseCortex-IOS dataset.
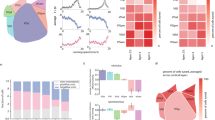
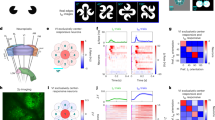
All image data was stored on multiple 1TB West Digital external hard drives. The experimental setup and the schematic structure of the imaging system are illustrated in Fig. 2a and b, respectively. Figure 2a depicts the actual image acquisition scene of the experimental platform, with professionals performing stimulation on the experimental mouse. Figure 2b shows a schematic diagram of the experimental platform’s structure, including a centrally positioned camera, an illumination light source, and an adjustable fixed platform for the experimental subject at the bottom. Figure 2c presents a representative collected image, with a resolution of 512 × 512 pixels and a 16-bit grayscale depth per pixel.
Data preprocessing
Raw IOS signal data has significant limitations, including weak signal intensity and motion artifacts, as exemplified by the raw image shown in Fig. 2(c). This makes it unsuitable for direct signal analysis, so efficient preprocessing steps are essential before further analysis (details are presented in the left part of Fig. 1). Temporal averaging of consecutive frames helped minimize environmental noise and motion artifacts through per-second integration. This computation reduced the volume of raw video data by two orders of magnitude while preserving IOS dynamics, as these signals propagate inherently slowly. Subsequent interval differencing amplified subtle signal variations; we used an optimized 5-frame interval for this step, which balances distortion prevention and data compression, reducing the data volume by a further five times. Pseudo-color transformation improved the visualization of differential signals by converting grayscale intensity gradients into visually distinct color scales. This sequential preprocessing workflow produced temporally resolved IOS propagation maps that are suitable for quantitative analysis.
Experimental setup for data collection and representative results.
Data labeling
An additional challenge still need to be addressed prior to data labeling. Preprocessed frames contain physiological blank intervals between cyclically triggered IOS events, which confound automated analysis by intermixing with noise-induced gaps. To resolve this, we performed manual validation by visually identifying complete signal episodes. Operators recorded the start and end frame timestamps in structured sequences. Python scripts were then used to automatically extract frame ranges between these boundaries, systematically excluding the blank intervals between triggers. For cases with consecutive signal overlaps, an adaptive merging strategy was applied to merge overlapping events into unified video segments. The integrity of individual waveforms was preserved using suffix-based tagging (e.g., SignalA-1, SignalA-2).
The SAM2-based segmentation and labeling process is a core component of our workflow and the primary contributor to reducing manual effort. SAM2 excels in its ability to track multiple objects simultaneously, even in the presence of temporary occlusion, and exhibits strong adaptability across various application fields. We adopted an excellent open-source tool that offers an interactive labeling interface supporting the integration of segmentation models. This tool, named “ISAT with Segment Anything” (abbreviated as “ISAT”), is a semi-automatic annotation software on GitHub with 1.9k stars. One of the interfaces utilized during labeling is shown in Fig. 3, which depicts the basic interface for annotating images. At the top of the interface are various setting buttons, corresponding with most software applications: the “File” menu is used to select the target folder, and the “SAM” menu to choose the segmentation model. Below these buttons are shortcut keys for functional modes, such as single-frame propagation mode and video propagation mode. The largest area in center displays the image currently being labeled. To the left of this central area is the signal color setting panel, where operators can select colors for labels. The right-hand area contains information panels, which show details such as the image size, the color of the current label, and the active folder.
Interface of the open-source ISAT labeling tool.
We streamlined the labeling process by loading the SAM2 model into this software, and selected the “tiny” version to balance performance and speed. The tool provides access to four SAM2 models. For an average video in our dataset, the other three models required approximately 1.5, 2, and 3 times the inference time of the tiny version, but offered no significant improvement in segmentation precision. The workflow starts with loading a folder of image frames, where each folder corresponds to a single signal trigger. Labeling begin with the first frame of each sequence, as SAM2 needs an initial reference to define the target object for segmentation and tracking. Once the initial frame is labeled, segmentation for all subsequent frames can be completed with a single click, greatly enhancing efficiency. The first-frame labeling can also be simplified by SAM2’s one-click segmentation feature, eliminating the need for tedious manual contour selection. However, manual labeling is required if the first frame is ambiguous. Each folder corresponds to one signal trigger, and finishing its labeling completes the processing of that trigger. This process is repeated for all data folders. Compared to traditional point-by-point and frame-by-frame labeling methods, SAM2’s interactive workflow speeds up the labeling of IOS datasets by an order of magnitude.
In the case of special scenarios mentioned before, where a single video segment contains two or more overlapping signals, additional steps are necessary. If the signals have different start and end frames, label the earlier signal as “label_name-1” at its first frame. Use SAM2 to propagate this label until the second signals’ start frame. Then, manually label the second signal’s start frame and propagate it using SAM2 until the end frame. During this process, SAM2 retains the segmentation and tracking of the first signal. For cases where two signals appear simultaneously in the same starting frame, we label both objects in the initial frame with distinct labels (e.g., “label_name-1” and “label_name-2”) and proceed with SAM2’s propagation for each signal. This approach ensures precise labeling even in complex scenarios, leveraging SAM2’s robust segmentation capabilities while maintaining efficiency and accuracy. Moreover, in cases where noise introduces blank frames within a signal segment, SAM2’s capability to track occluded objects allows it to skip signal segmentation for these frames without losing its ability to track signals in later frames. However, since we aim to label the entire signal variation process, these noise-induced blank frames need to be labeled manually, like traditional methods. Although this process is time-consuming, the proportion of such noise-induced blank frames is small, so they add little to the total labeling time.
Data Records
The MouseCortex-IOS dataset is publicly available on figshare under a CC-BY 4.0 license, comprising 0.46 GB of compressed ZIP files that can be accessed without restrictions at https://doi.org/10.6084/m9.figshare.2860181315. The dataset includes preprocessed pseudo-color images (TIFF format) and corresponding annotation files (JSON format) generated by our pipeline, with strict one-to-one correspondence between images and annotations. Each experimental subject is organized into an independent folder named based on the recording date and subject ID (e.g., 20230922-shu-101). Subfolders within these directories correspond to individual signal-triggered events, and are named by video segment numbers as well as timestamped of start and end frames (e.g., segment_0_20230922_221431_20230922_221556). Additionally, there is a configuration files named ‘isat.yaml’ which is automatically generated from the open-source labeling tool, requiring no user intervention. The decompressed directory structure is illustrated in Fig. 4. The figure only displays the example of the structure, more detailed items can be found in Table 1.
File structure overview of MouseCortex-IOS dataset.
Technical Validation
To assess the reliability of the MouseCortex-IOS dataset, we conducted a stratified sampling evaluation of the labels generated by our pipeline using Dice coefficient and Intersection over Union (IoU). A subset of 30 signal segments was selected from the total of 194 segments across all experimental subjects. Prior to sampling, a neurophysiology expert categorized all segments into three levels based on noise severity and frequency of ambiguous frames. This ensured representative sampling across segments with heterogeneous signal qualities, and 10 segments were randomly selected from each level.
The criteria for these levels were established based on the expertise of our team members, including clinical doctor and experts in medical image processing. Level 1 includes segments with consistently clear IOS signals across all frames. Level 2 comprises segments that are predominantly clear but contain sporadic noise-affected frames, such as transient blanks or motion blur. Although foundation models like SAM2 may perform poorly on corrupted frames, they typically recover accurate tracking in subsequent frames. Human annotators can efficiently identify and correct these localized anomalies. Level 3 consists of segments with severe noise degradation, where even human annotators struggle to discern signals. These segments represent the most challenging cases for automated segmentation. Representative images of the three levels are illustrated in Fig. 5.
Representative images of the three levels. Panels (a–c) correspond to the labeled masked images for Level 1, Level 2, and Level 3, respectively, while panels (d–f) are the corresponding original preprocessed images. It can be observed that the IOS signals in (d–f) become increasingly indistinct.
For the selected 30 segments, an experienced operator was firstly employed to label them manually treated as ground-truth annotation. Then, we evaluated segmentation performance across three approaches: (1) U-Net, a classical deep learning model; (2) SAM2 (Segment Anything Model V2); and (3) our pipeline. The U-Net model adopted the classic encoder-decoder architecture combined with skip connections, designed for the binary segmentation task of intrinsic optical signal (IOS) regions. Implemented based on the PyTorch framework, this model was trained on a NVIDIA GeForce GTX 1080Ti GPU. A total of 30 selected datasets were divided into training and test sets at an 8:2 ratio within each quality level, resulting in a training set consisting of 24 video clips (680 images in total) and a test set with 6 video clips (172 images in total). We observed that the final image ratio of the training to test set was approximately 7.98:2.02. This slight deviation arose from the fact that our initial division was performed on a per-video basis, and the number of frames varied across different videos. Nevertheless, the final image ratio remained highly close to the 8:2 split, which fully demonstrates that this level-based division method not only ensures a relatively consistent proportion of images with different quality levels in both the training and test sets but also maintains a high consistency between the image ratio and the video clip ratio of the two sets, thus guaranteeing the scientific reliability of the U-Net experiment. To mitigate overfitting, a data augmentation strategy of random shuffling was applied during the training phase. The model was set to train for 100 epochs with a batch size of 4, using the Adam optimizer and Binary Cross-Entropy (BCE) as the loss function. After 10 epochs of training, the loss value of the model dropped below 0.1, with the entire training process taking about 4.5 hours; the trained weights were then used for inference on the test set. For SAM2, inference was directly performed on the selected video clips using the ISAT annotation tool, with the tiny SAM2 model loaded.
Comparative results in Table 2 demonstrate that our semi-interactive pipeline achieves a high overlap with manual annotations (average Dice coefficient: 0.82; average IoU: 0.71). For reference, we also include the performance of two baselines: the U-Net model and SAM2. Note that our comparison with SAM2 is not for capability rivalry but to highlight the value of our IOS-customized pipeline in balancing annotation efficiency and accuracy, because we know that it’s unfair to compare our specialized pipeline with a foundation model. Bold values in the table denote the optimal metrics for each evaluation dimension. The result confirms that our dataset reliably captures biologically relevant IOS dynamics, providing a solid foundation for model training. Additionally, our semi-automated framework maintains high annotation quality even under heterogeneous noise conditions, which general baselines lack.
For the manual annotation parts involved in the above process, we have two experienced members, including a doctor, who jointly annotate the parts that need manual annotation and have reached a consensus on the best annotation area. To ensure the reliability of the annotated data, we conducted an additional annotation consistency test on the two annotators. We selected two video clips from each of Level 1 (60 frames), Level 2 (51 frames), and Level 3 (61 frames), totaling 172 frames of images, and required the two annotators to annotate them independently. Finally, we calculated the Cohen’s16 coefficient for each level and the entire samples, and the results are recorded in Table 3. The coefficients in the results are all between 0.7 and 0.8, which to some extent indicates that the two annotators have good consistency in annotating this data, and the annotation results are relatively reliable.
In addition, the original purpose of our dataset is to facilitate spatiotemporal tracking and analysis of cortical signals. Accordingly, we computed key parameters including the signal’s starting point, ending point, movement trajectory, duration, velocity, and coverage area using the outputs generated by our method. These metrics were computed through pixel-level analysis of annotation masks, with spatial calibration to their corresponding physical dimensions (11μm per 512pixel) achieved using sensor-specific parameters. The computational algorithms have been fully documented in our open-source code repository; for details, please refer to the “Code Availability” section.
Figure 6 illustrates representative results, displaying five evenly spaced frames extracted from a 25-frame propagation sequence. The ‘F’ label in blue box denotes the frame number within 25-frame sequence. This example corresponds to one analytical case randomly choose from the dataset entry “20231012-shu-113”, and it depicts spreading depression (CSD) recorded following induction with 0.125 M KCL following electrical stimulation of the vagus nerve. This demonstrates one of the applications of IOS in investigating vagal nerve responses to stimulation. The visualized image and video integrate signal trajectory and average velocity vectors. The complete results are provided in the supplementary materials. These results have been confirmed by neuroscience experts to be consistent with the characteristics of real cortical signals, supporting the research on vagus nerve stimulation.
Display of some extracted frames from a vagus nerve stimulation signal video segment.
Data availability
The dataset is publicly available on figshare: https://doi.org/10.6084/m9.figshare.28601813.
Code availability
The preprocessing code and other scripts that have been used in the pipeline are accessible at: https://github.com/ZhangWang-hub/MouseCortex-IOS. The open-source labeling tool can be download at: https://github.com/yatengLG/ISAT_with_segment_anything.
References
Zepeda, A., Arias, C. & Sengpiel, F. Optical imaging of intrinsic signals: Recent developments in the methodology and its applications. J. Neurosci. Methods 136, 1–21 (2004).
Li, P., Luo, W., Luo, Q. & Cheng, S. An evaluation of data analysis methods for optical intrinsic signal imaging. in (eds. Luo, Q., Tuchin, V. V., Gu, M. & Wang, L. V.) 542, https://doi.org/10.1117/12.546565 (Wuhan, China, 2003).
Crouzet, C., Phan, T., Wilson, R. H., Shin, T. J. & Choi, B. Intrinsic, widefield optical imaging of hemodynamics in rodent models of alzheimer’s disease and neurological injury. Neurophotonics 10 (2023).
Trizna, E. Y. et al. Brightfield vs fluorescent staining dataset–a test bed image set for machine learning based virtual staining. Sci. Data 10, 160 (2023).
Lu, H. D., Chen, G., Cai, J. & Roe, A. W. Intrinsic signal optical imaging of visual brain activity: Tracking of fast cortical dynamics. NeuroImage 148, 160–168 (2017).
Li, J., Zheng, A., Chen, X. & Zhou, B. Primary video object segmentation via complementary CNNs and neighborhood reversible flow. in 2017 IEEE International Conference on Computer Vision (ICCV) 1426–1434, https://doi.org/10.1109/ICCV.2017.158 (IEEE, Venice, 2017).
Wang, W., Lu, X., Shen, J., Crandall, D. & Shao, L. Zero-shot video object segmentation via attentive graph neural networks. in 2019 IEEE/CVF International Conference on Computer Vision (ICCV) 9235–9244, https://doi.org/10.1109/ICCV.2019.00933 (IEEE, Seoul, Korea (South), 2019).
Fragkiadaki, K., Arbelaez, P., Felsen, P. & Malik, J. Learning to segment moving objects in videos. in 2015 IEEE Conference on Computer Vision and Pattern Recognition (CVPR) 4083–4090, https://doi.org/10.1109/CVPR.2015.7299035 (IEEE, Boston, MA, USA, 2015).
Li, M., Li, S., Zhang, X. & Zhang, L. UniVS: Unified and universal video segmentation with prompts as queries. in 2024 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR) 3227–3238, https://doi.org/10.1109/CVPR52733.2024.00311 (IEEE, Seattle, WA, USA, 2024).
Mattjie, C. et al. “Zero-Shot Performance of the Segment Anything Model (SAM) in 2D Medical Imaging: A Comprehensive Evaluation and Practical Guidelines,” 2023 IEEE 23rd International Conference on Bioinformatics and Bioengineering (BIBE), Dayton, OH, USA, pp. 108-112, https://doi.org/10.1109/BIBE60311.2023.00025 (2023).
Yan, Z. et al. Biomedical SAM2: Segment Anything in Biomedical Images and Videos. Preprint at http://arxiv.org/abs/2408.03286 (2024).
Zhu, J., Hamdi, A., Qi, Y., Jin, Y. & Wu, J. Medical SAM2: Segment medical images as video via segment anything model 2. Preprint at, https://doi.org/10.48550/arXiv.2408.00874 (2024).
Kirillov A. et al. Segment Anything. 2023 IEEE/CVF International Conference on Computer Vision (ICCV), Paris, France, pp. 3992–4003, https://doi.org/10.1109/ICCV51070.2023.00371 (2023).
Ravi, N. et al. SAM 2: Segment anything in images and videos. Preprint at, https://doi.org/10.48550/arXiv.2408.00714 (2024).
Zhang, W. et al. MouseCortex-IOS dataset. Figshare https://doi.org/10.6084/m9.figshare.28601813 (2024).
Cohen, J. A coefficient of agreement for nominal scales. Educational and psychological measurement 20(1), 37–46 (1960).
Acknowledgements
The research was funded by Guangdong Basic and Applied Basic Research Foundation (2021A1515220151), Guangzhou Science and Technology Planning Program (No. 202201020518), Guangzhou Science and Technology Projects (2023A03J0958), Guangzhou Municipal Key R and D Program, China (2024B03J0947). Additionally, the authors would like to express sincere gratitude to the developers of the open-sourced AI toolkit ISAT_with_segment_anything (ISAT) for making this valuable resource publicly available.
Author information
Authors and Affiliations
Contributions
W.Z. created the dataset, performed the analysis and post-processing, and wrote the manuscript. G.Z. designed and constructed the optical acquisition platform, conducted experimental data acquisition, and performed partial data preprocessing. Z.Z., H.L., W.Q. and J.L. analyzed the results and provided suggestions. All authors actively participated in revising the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhang, W., Zeng, G., Zheng, Z. et al. A Mouse Cortex Video Segmentation Dataset for Intrinsic Optical Signal Tracking and Neural Activity Analysis. Sci Data 13, 255 (2026). https://doi.org/10.1038/s41597-026-06580-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41597-026-06580-1