Abstract
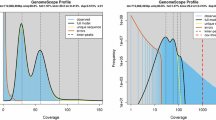
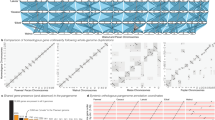
The pecan weevil, Curculio caryae (Horn), is an obligate feeder of pecan and native hickory trees (genus Carya) throughout North America. Subsequently it is a significant agricultural pest in pecan orchards. In this study, we present a reference quality genome using deep-coverage, ~40x PacBio HiFi genome sequence reads, and chromatin confirmation, Hi-C, scaffolding. The final genome assembly is approximately 2.2 Gb, which was confirmed by flow cytometry. The primary genome scaffolds have an N50 of 132 Mb and a BUSCO completeness of 95.4% [S:94.3%, D:1.1%]. Furthermore, we employed PacBio long-read RNA, Iso-seq, for de novo gene annotation, in conjunction with InterProscan to identify approximately 19,000 protein coding genes. Repeat content is extensive, contributing at least >80% of the total genome. This data set provides a valuable resource for comparative genomics and evolutionary studies of an economically impactful group of insect pests that currently lack extensive genomic resources.
Similar content being viewed by others
Data availability
The raw sequencing data, genome assembly, transcripts, and mitochondrial genome of Curculio caryae have been deposited at the National Center Biotechnology under project number PRJNA813156 and at the National Ag Library https://hdl.handle.net/10779/USDA.ADC.29329910. The custom annotations can be found at the National Ag Library https://doi.org/10.15482/USDA.ADC/30234490.
Code availability
Data processing was executed using published programs and default parameters unless otherwise specified in the Methods section. No custom code was used for these analyses.
References
Global Biodiversity Information Facility, https://www.gbif.org/species/4239 (2025).
Keeling, C. I. et al. Draft genome of the mountain pine beetle, Dendroctonus ponderosae Hopkins, a major forest pest. Genome Biol. 14, 1–20, https://doi.org/10.1186/gb-2013-14-3-r27 (2013).
Vega, F. E. et al. Draft genome of the most devastating insect pest of coffee worldwide: the coffee berry borer, Hypothenemus hampei. Sci. Rep. 5(1), 12525, https://doi.org/10.1038/srep12525 (2015).
Harrop, T. W. et al. Genetic diversity in invasive populations of argentine stem weevil associated with adaptation to biocontrol. Insects 11(7), 441, https://doi.org/10.3390/insects11070441 (2020).
Dias, G. B. et al. Haplotype-resolved genome assembly enables gene discovery in the red palm weevil Rhynchophorus ferrugineus. Sci. Rep. 11(1), 9987, https://doi.org/10.1038/s41598-021-89091-w (2021).
Parisot, N. et al. The transposable element-rich genome of the cereal pest Sitophilus oryzae. BMC Biol. 19, 241, https://doi.org/10.1186/s12915-021-01158-2 (2021).
Powell, D. et al. A highly-contiguous genome assembly of the Eurasian spruce bark beetle, Ips typographus, provides insight into a major forest pest. Commun. Biol. 4(1), 1059, https://doi.org/10.1038/s42003-021-02602-3 (2021).
Van Dam, M. H. et al. The Easter Egg Weevil (Pachyrhynchus) genome reveals syntenic patterns in Coleoptera across 200 million years of evolution. PLoS Genet. 17(8), e1009745, https://doi.org/10.1371/journal.pgen.1009745 (2021).
Cohen, Z. P. et al. Insight into weevil biology from a reference quality genome of the boll weevil, Anthonomus grandis grandis Boheman (Coleoptera: Curculionidae). G3 13(2), jkac309, https://doi.org/10.1093/g3journal/jkac309 (2023).
Gagalova, K. K. et al. The genome of the forest insect pest Pissodes strobi reveals genome expansion and evidence of a Wolbachia endosymbiont. G3 12(4), jkac038, https://doi.org/10.1093/g3journal/jkac038 (2022).
Liu, Z. et al. Chromosome-level genome assembly and population genomic analyses provide insights into adaptive evolution of the red turpentine beetle, Dendroctonus valens. BMC Biol. 20(1), 190, https://doi.org/10.1186/s12915-022-01388-y (2022).
McKenna, D. D., Sequeira, A. S., Marvaldi, A. E. & Farrell, B. D. Temporal lags and overlap in the diversification of weevils and flowering plants. PNAS 106(17), 7083–7088, https://doi.org/10.1073/pnas.0810618106 (2009).
Barry, R. M. & South, P. Costs of insect damage. Pecan South 1, 33 (1947).
Harris, M. K. Pecan arthropod management. ARS US Department of Agriculture, Agricultural Research Service (1991).
Harris, M. et al. Economic impact of pecan integrated pest management implementation in Texas. J. Econ. Entomol. 91(5), 1011–1020, https://doi.org/10.1093/jee/91.5.1011 (1998).
Mulder, P. G., Harris, M. K. & Grantham, R. A. Biology and Management of the Pecan Weevil (Coleoptera: Curculionidae). Integr. Pest Manag. 3(1), A1–A9, https://doi.org/10.1603/IPM10027 (2012).
Sim, S. B., Corpuz, R. L., Simmonds, T. J. & Geib, S. M. HiFiAdapterFilt, a memory efficient read processing pipeline, prevents occurrence of adapter sequence in PacBio HiFi reads and their negative impacts on genome assembly. BMC Gen. 23(1), 157, https://doi.org/10.1186/s12864-022-08375-1 (2022).
Cheng, H., Concepcion, G. T., Feng, X., Zhang, H. & Li, H. Haplotype-resolved de novo assembly using phased assembly graphs with hifiasm. Nat. Methods. 18(2), 170–175, https://doi.org/10.1038/s41592-020-01056-5 (2021).
Guan, D. et al. Identifying and removing haplotypic duplication in primary genome assemblies. Bioinform. 36(9), 2896–2898, https://doi.org/10.1093/bioinformatics/btaa025 (2020).
Zhou, C., McCarthy, S. A. & Durbin, R. YaHS: yet another Hi-C scaffolding tool. Bioinform. 39(1), btac808, https://doi.org/10.1093/bioinformatics/btac808 (2023).
Durand, N. C. et al. Juicebox provides a visualization system for Hi-C contact maps with unlimited zoom. Cell Syst. 3(1), 99–101, https://doi.org/10.1016/j.cels.2015.07.012 (2016).
Dudchenko, O. et al. The Juicebox Assembly Tools module facilitates de novo assembly of mammalian genomes with chromosome-length scaffolds for under $1000. BioRxiv. 1, 254797 (2018).
Li, H. Minimap2: pairwise alignment for nucleotide sequences. Bioinform. 34(18), 3094–3100, https://doi.org/10.1093/bioinformatics/bty191 (2018).
Laetsch, D. R. & Blaxter, M. L. BlobTools: Interrogation of genome assemblies. F1000Research 6(1287), 1287, https://doi.org/10.12688/f1000research.12232.1 (2017).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinform. 26(6), 841–842, https://doi.org/10.1093/bioinformatics/btq033 (2010).
Lachowska, D., Holecova, M. & Rozek, M. Karyotypic data on weevils (Coleoptera, Curculionidae). FOLIA BIOLOGICA-KRAKOW- 46, 129–136 (1998).
Li, H. et al. 1000 Genome Project Data Processing Subgroup. The Sequence Alignment/Map format and SAMtools. Bioinform. 25(16), 2078–9, https://doi.org/10.1093/bioinformatics/btp352 (2009).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:SRP362700 (2024).
NCBI Genbank https://identifiers.org/ncbi/insdc:JAKZMK000000000
National Ag Library https://hdl.handle.net/10779/USDA.ADC.29329910 (2024).
National Ag Library https://doi.org/10.15482/USDA.ADC/30234490 (2014).
Ranallo-Benavidez, T. R., Jaron, K. S. & Schatz, M. C. GenomeScope 2.0 and Smudgeplot for reference-free profiling of polyploid genomes. Nat. Commun. 11(1), 1432, https://doi.org/10.1038/s41467-020-14998-3 (2020).
Kokot, M., Długosz, M. & Deorowicz, S. KMC 3. counting and manipulating k-mer statistics. Bioinform. 33(17), 2759–2761, https://doi.org/10.1093/bioinformatics/btx304 (2017).
Mapleson, D., Garcia Accinelli, G., Kettleborough, G., Wright, J. & Clavijo, B. J. KAT: a K-mer analysis toolkit to quality control NGS datasets and genome assemblies. Bioinform. 33(4), 574–576, https://doi.org/10.1093/bioinformatics/btw663 (2017).
Manni, M., Berkeley, M. R., Seppey, M., Simão, F. A. & Zdobnov, E. M. BUSCO update: novel and streamlined workflows along with broader and deeper phylogenetic coverage for scoring of eukaryotic, prokaryotic, and viral genomes. MBE 38(10), 4647–4654, https://doi.org/10.1093/molbev/msab199 (2021).
Acknowledgements
This work was partially supported through funds from the Texas Pecan Board (Agreement # 58-3091-0-019) and the U.S. Department of Agriculture, Agricultural Research Service (USDA-ARS, CRIS Projects 3091-22000-038-000D and 2040-22430-028-000-D). The genome assembly was generated as part of the USDA-ARS Ag100Pest Initiative. The authors thank members of the USDA-ARS Ag100Pest Team for sequencing and analysis support. This research used resources provided by the SCINet project of the USDA-ARS project number 0500-00093-001-00-D. Mention of trade names or commercial products in this publication is solely for the purpose of providing specific information and does not imply recommendation or endorsement by the U.S. Department of Agriculture. USDA is an equal opportunity provider and employer. Special thanks to Mike Barry (Comanche Co., TX, Extension Agent), and Patrick Dudley (Texas Department of Agriculture) for their assistance in setting up Circle traps. The US Department of Agriculture, Agricultural Research Service is an equal opportunity/affirmative action employer, and all agency services are available without discrimination.
Author information
Authors and Affiliations
Contributions
Designed research: Lindsey C. Perkin, Zachary P. Cohen, and Charles P.-C. Suh. Collection of samples: Lindsey C Perkin and Charles P.-C. Suh. Genome assembly and data analysis: Zachary P. Cohen, Sheina B. Sims, Scott M. Geib. Flow Cytometry: J. Spencer Johnston. Resources: Timothy P.L. Smith and Perot Saelao. Manuscript writing: Lindsey C. Perkin and Zachary P. Cohen. All authors provided suggestions for the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Perkin, L.C., Cohen, Z.P., Sim, S.B. et al. A chromosome level reference genome for the pecan weevil, Curculio caryae. Sci Data (2026). https://doi.org/10.1038/s41597-026-07030-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41597-026-07030-8