Abstract
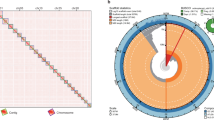
The Turpan minnow (Phoxinus grumi) is a small endemic fish species inhabiting the extreme environment of the Turpan Basin in Xinjiang, China, holding significant value for evolutionary and conservation biology research. However, the absence of a high-quality reference genome has severely constrained studies on its adaptive evolution and conservation genetics, in stark contrast to the available chromosome-level genomes of its congeners, such as Phoxinus phoxinus. A total of 240.38 Gb of sequencing data was generated in this study, comprising 44.12Gb (53.35×) of PacBio HiFi reads, 50.36 Gb (60.90×) of Illumina reads, 120.59 Gb (133.95×) of Hi-C data and 25.31 Gb of RNA sequencing data, which enabled the successful assembly of a chromosome-level genome for P. grumi. The assembled genome has a total size of 900.41 Mb, with 97.58% of the sequences anchored onto 25 chromosomes. The contig N50 and scaffold N50 reached 17.52 Mb and 34.99 Mb, respectively. BUSCO assessment indicated a genome completeness of 98.1%. We predicted a total of 24,224 protein-coding genes, of which 90.8% were functionally annotated. This high-quality reference genome will serve as a key genetic resource for in-depth exploration of the environmental adaptation mechanisms and species conservation of P. grumi.
Similar content being viewed by others
Data availability
The raw sequencing data are available in the NCBI databases under Bioproject accession number PRJNA1399684. Additionally, the assembled genome has been deposited in GenBank. Furthermore, all datasets are available under the BioProject accession number PRJCA050662 in the Genome Warehouse (GWH) at the National Genomics Data Center (NGDC). The data are publicly accessible via the following link at https://ngdc.cncb.ac.cn/gwh. Raw reads have been deposited in NGDC (Hi-C: SAMC6098139; Illumina: SAMC6098138; PacBio HiFi: SAMC6098137). The final genome assembled and annotation files have been deposited in Figshare platform via https://doi.org/10.6084/m9.figshare.30572321.v1.
Code availability
No custom code or scripts were utilized in this study, all commands and pipelines involved in data processing were executed in accordance with the manuals and protocols provided by the bioinformatic software employed. The specific versions of software packages and corresponding parameters implemented for each analytical step are explicitly detailed in the Methods section to ensure reproducibility.
References
Zardoya, R. & Doadrio, I. Molecular evidence on the evolutionary and biogeographical patterns of European cyprinids. J Mol Evol. 49, 227–237 (1999).
Imoto, J. M. et al. Phylogeny and biogeography of highly diverged freshwater fish species (Leuciscinae, Cyprinidae, Teleostei) inferred from mitochondrial genome analysis. Gene. 514, 112–124 (2013).
Schönhuth, S. et al. Phylogenetic relationships and classification of the Holarctic family Leuciscidae (Cypriniformes: Cyprinoidei). Mol Phylogenet Evol. 127, 781–799 (2018).
Palandačić, A., Witman, K. & Spikmans, F. Molecular analysis reveals multiple native and alien Phoxinus species (Leusciscidae) in the Netherlands and Belgium. Biol Invasions. 24, 2273–2283 (2022).
Page, L. M. et al. Common and Scientific Names of Fishes from the United States, Canada, and Mexico (8th ed.). Fisheries. 48, 497–498 (2023).
Zhou, Y. et al. Telomere-to-telomere genome assembly of Phoxinus lagowskii. Sci Data. 12, 1025 (2025).
Zheng, H. et al. Chromosome-level genome assembly of the Phoxinus lagowskii. Sci Data. 12, 1400 (2025).
Zhang, C. & Zhao, Y. Species Diversity and Distribution of Inland Fishes in China. (Science Press, 2016).
Bridle, J. R., Pedro, P. M. & Butlin, R. K. Habitat fragmentation and biodiversity: testing for the evolutionary effects of refugia. Evolution. 58, 1394–1400 (2004).
Du, L. et al. Hydroclimatic Change in Turpan Basin under Climate Change. Water. 15, 3422 (2025).
Di Giulio, M., Holderegger, R. & Tobias, S. Effects of habitat and landscape fragmentation on humans and biodiversity in densely populated landscapes. J Environ Manage. 90, 2959–2968 (2009).
Nunn, A. D. et al. The genome sequence of the Eurasian minnow, Phoxinus phoxinus (Linnaeus, 1758). Wellcome Open Res. 9, 504 (2024).
Oriowo, T. O. et al. A chromosome-level, haplotype-resolved genome assembly and annotation for the Eurasian minnow (Leuciscidae: Phoxinus phoxinus) provide evidence of haplotype diversity. Gigascience. 14, giae116 (2025).
Wenger, A. M. et al. Accurate circular consensus long-read sequencing improves variant detection and assembly of a human genome. Nat Biotechnol. 37, 1155–1162 (2019).
van Berkum, N. L. et al. Hi-C: a method to study the three-dimensional architecture of genomes. J Vis Exp, 1869 (2010).
Chen, S. et al. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics. 34, i884–i890 (2018).
Marçais, G. & Kingsford, C. A fast, lock-free approach for efcient parallel counting of occurrences of k-mers. Bioinformatics. 27, 764–770 (2011).
Ranallo-Benavidez, T. R., Jaron, K. S. & Schatz, M. C. GenomeScope 2.0 and Smudgeplot for reference-free profling of polyploid genomes. Nat Commun. 11, 1432 (2020).
Cheng, H. et al. Haplotype-resolved de novo assembly using phased assembly graphs with hifasm. Nat Methods. 18, 170–175 (2021).
Durand, N. C. et al. Juicer Provides a One-Click System for Analyzing Loop-Resolution Hi-C Experiments. Cell Syst. 3, 95–98 (2016).
Dudchenko, O. et al. De novo assembly of the Aedes aegypti genome using Hi-C yields chromosome-length scafolds. Science. 356, 92–95 (2017).
Durand, N. C. et al. Juicebox provides a visualization system for Hi-C contact maps with unlimited zoom. Cell Syst. 3, 99–101 (2016).
Krzywinski, M. et al. Circos: an information aesthetic for comparative genomics. Genome Res. 19, 1639–1645 (2009).
Simão, F. A. et al. BUSCO: assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics. 31, 3210–3212 (2015).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics. 25, 1754–1760 (2009).
Rhie, A. et al. Merqury: reference-free quality, completeness, and phasing assessment for genome assemblies. Genome Biol. 21, 245 (2020).
Tarailo‐Graovac, M. & Chen, N. Using RepeatMasker to identify repetitive elements in genomic sequences. Curr Protoc Bioinformatics. 25, 4–10 (2009).
Flynn, J. M. et al. RepeatModeler2 for automated genomic discovery of transposable element families. Proc. Natl. Acad. Sci. USA. 117, 9451–9457 (2020).
Stanke, M. et al. AUGUSTUS: a web server for gene finding in eukaryotes. Nucleic Acids Res. 32, W309–W312 (2004).
Stanke, M. & Morgenstern, B. AUGUSTUS: a web server for gene prediction in eukaryotes that allows user-defined constraints. Nucleic Acids Res. 33, W465–W467.
Stanke, M. et al. AUGUSTUS: ab initio prediction of alternative transcripts. Nucleic Acids Res. 34, W435–W439.
Korf, I. Gene fnding in novel genomes. BMC Bioinformatics. 5, 59 (2004).
Gertz, E. M. et al. Composition-based statistics and translated nucleotide searches:improving the TBLASTN module of BLAST. BMC Biol. 4, 41 (2006).
Birney, E., Clamp, M. & Durbin, R. GeneWise and Genomewise. Genome Res. 14, 988–3995 (2004).
Kim, D., Langmead, B. & Salzberg, S. L. HISAT: A fast spliced aligner with low memory requirements. Nat. Methods. 12, 357–360 (2015).
Pertea, M. et al. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat. Biotechnolo. 33, 290–295 (2015).
Haas, B. J. et al. Automated eukaryotic gene structure annotation using EVidenceModeler and the Program to Assemble Spliced Alignments. Genome Biol. 9, 1–22 (2008).
Boeckmann, B. et al. The SWISS-PROT protein knowledgebase and its supplement TrEMBL in 2003. Nucleic Acids Res. 31, 365–370 (2003).
McGinnis, S. & Madden, T. L. BLAST: at the core of a powerful and diverse set of sequence analysis tools. Nucleic Acids Research. 32, W20–W25 (2004).
Zdobnov, E. M. & Apweiler, R. InterProScan–an integration platform for the signature-recognition methods in InterPro. Bioinformatics 17, 847–848 (2001).
Bru, C. et al. The ProDom database of protein domain families: more emphasis on 3D. Nucleic Acids Res. 33, D212–D215 (2005).
Corpet, F., Gouzy, J. & Kahn, D. The ProDom database of protein domain families. Nucleic Acids Res. 26, 323–326 (1998).
Attwood, T. K. The PRINTS database: a resource for identification of protein families. Brief Bioinform. 3, 252–263 (2002).
Mistry, J. et al. Pfam: Te protein families database in 2021. Nucleic Acids Res. 49, D412–D419 (2021).
Letunic, I. & Bork, P. 20 years of the SMART protein domain annotation resource. Nucleic Acids Res. 46, D493–D496 (2018).
Mi, H. et al. The PANTHER database of protein families, subfamilies, functions and pathways. Nucleic Acids Res. 33, D284–D288 (2005).
Hulo, N. et al. The PROSITE database. Nucleic Acids Res. 34, D227–D230 (2006).
Buchfnk, B., Xie, C. & Huson, D. H. Fast and sensitive protein alignment using DIAMOND. Nat Methods. 12, 59–60 (2015).
Chan, P. P. et al. tRNAscan-SE 2.0: improved detection and functional classifcation of transfer RNA genes. Nucleic Acids Res. 49, 9077–9096 (2021).
Nawrocki, E. P. & Eddy, S. R. Infernal 1.1: 100-fold faster RNA homology searches. Bioinformatics. 29, 2933–2935 (2013).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:SRR36766626 (2026).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:SRR36766627 (2026).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:SRR36766628 (2026).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:SRR36766624 (2026).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:SRR36766625 (2026).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:SRR36766622 (2026).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:SRR36766623 (2026).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:SRR36843988 (2026).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:SRR36843989 (2026).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:SRR36843990 (2026).
NCBI GenBank https://identifiers.org/ncbi/insdc.gca:GCA_055048795.1 (2026).
Members, C.-N., Partners Database resources of the National Genomics Data Center. China National Center for Bioinformation in 2024. Nucleic Acids Res. 52, D18–D32 (2024).
Chang, H. Genome Annotation Dataset of Phoxinus grumi. Figshare. https://doi.org/10.6084/m9.figshare.30572321.v1 (2025).
Acknowledgements
This research was supported by the Third Xinjiang Scientific Expedition Program (No. 2022xjkk1505), the Xinjiang Key Laboratory for Ecological Adaptation and Evolution of Extreme Environment Organisms (No. KFKT2402), the China Postdoctoral Science Foundation (No. 339494), and the Xinjiang Uygur Autonomous Region Tianchi Talent Introduction Program.
Author information
Authors and Affiliations
Contributions
J.W. and H.C. conducted the bioinformatic analyses including genome assembly and gene annotation, and drafted the manuscript. P.Y. processed and refined the images and contributed to data analysis. W.G., X.W., X.L., Y.H. and M.G. collected the samples and performed the animal experiments. J.W. and W.G. revised and edited the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Wang, J., Chang, H., Yang, P. et al. A chromosomal-level genome assembly of Phoxinus grumi (Cypriniformes: Leuciscidae). Sci Data (2026). https://doi.org/10.1038/s41597-026-07087-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41597-026-07087-5