Abstract
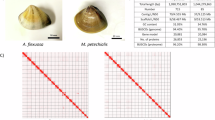
The genus Meretrix contains marine bivalve molluscs commonly known as the Venus clams or hard clams, which can be found in the estuarine and marine habitats in Asia. Given their edibility, they have been exploited in clam digging activities and also been farmed in some places. Here, we provide two new high-quality genomes of species M. meretrix and M. lamarckii. Utilising a combination of PacBio HiFi and Omni-C sequencing technologies, genome assemblies of M. meretrix and M. lamarcki are obtained with sizes 835.1 Mb (scaffold N50 = 46 Mb) and 890.5 Mb (scaffold N50 = 46 Mb), respectively. More than 99% of sequences were anchored to 19 pseudochromosomes, and high completeness was also obtained estimated by BUSCO scores (~99.5%, mollusca_odb12). The two new genomic resources provided in this study will be useful for further understanding biology, ecology, and evolution of edible clams.
Similar content being viewed by others
Data availability
The genome assemblies of M. meretrix and M. lamarcki are available at NCBI/ENA GenBank under accessions JBTXPG000000000 and JBTORC000000000, DDBJ accessions PRJDB40269 and PRJDB40270, NGDC accessions GWHHOJO00000000 and GWHHOOY00000000, and CUHK Research Data Repository (https://doi.org/10.48668/DOVLYS), respectively. The raw sequencing data of M. meretrix and M. lamarcki are deposited in the NCBI/ENA database under the SRA accession numbers SRP617717 and SRP618173, DDBJ database under the DRA accession numbers DRR911404-DRR911405 and DRR911157-DRR911158, NGDC database under the GSA accession numbers CRA039134 and CRA039136, and CUHK Research Data Repository (https://doi.org/10.48668/DOVLYS), respectively. The genome annotation files of M. meretrix and M. lamarckii can be found at CUHK Research Data Repository (https://doi.org/10.48668/DOVLYS).
Code availability
No specific script was used in this work.
References
Srinun, N., Vidthayanon, C. & Klangnurak, W. Morphological and genetic variations of Meretrix spp. (Bivalvia: Veneridae) recently distributed in the coastal waters of the Gulf of Thailand. Estuar. Coast. Shelf Sci. 307, 108889 (2024).
Soon, T. K., Sing, O. F., Denil, D. J. & Ransangan, J. Distribution and fishing pressure of hard clam, Meretrix meretrix in Marudu Bay, Sabah. Int. J. Oceans Oceanogr. 11 (2017).
Ye, Y. et al. Genetic Population Structure of the Hard Clam Meretrix meretrix Along the Chinese Coastlines Revealed by Microsatellite DNA Markers. Front. Mar. Sci. 7 (2020).
Chen, Y. et al. The biological resources survey of mudflat clams and other major creatures in Panjin Geligang and Xiao He. Hebei Fish 1, 46–49 (2012).
So, K. J. Y., Cheang, C. C., Hui, T. Y. & Chan, J. K. Y. Understanding the behavioural gap between perceived and actual environmental behaviour: Investigating the clam-harvesting pattern in Hong Kong SAR, China. J. Clean. Prod. 316, 128259 (2021).
WoRMS. World Register of Marine Species. https://www.marinespecies.org, https://doi.org/10.14284/170 (2025).
Chen, C.-C., Hsu, T.-H., Lu, H.-Y., Tang, S.-L. & Ho, Y.-N. High-quality chromosome-level genome of three Meretrix species using Nanopore and Hi-C technologies. Sci. Data 12, 1141 (2025).
Morton, B., Leung, S. F. & Leung, K. F. The biology and functional morphology of Meretrix cf. meretrix (Bivalvia: Veneridae: Meretricinae) at Tong Fuk Miu Wan, Shui Hau, Lantau Island, Hong Kong. Reg. Stud. Mar. Sci. 45, 101842 (2021).
Hsiao, S.-T. & Chuang, S.-C. Meretrix taiwanica (Bivalvia: Veneridae), a previously misidentified new species in Taiwan. Molluscan Res. 43, 12–21 (2023).
Law, S. T. S. et al. Genomes of two indigenous clams Anomalocardia flexuosa (Linnaeus, 1767) and Meretrix petechialis (Lamarck, 1818). Sci. Data 12, 409 (2025).
Folmer, O., Black, M., Hoeh, W., Lutz, R. & Vrijenhoek, R. DNA primers for amplification of mitochondrial cytochrome c oxidase subunit I from diverse metazoan invertebrates. Mol. Mar. Biol. Biotechnol. 3, 294–299 (1994).
Chen, J., Li, Q., Kong, L. & Zheng, X. Molecular phylogeny of venus clams (Mollusca, Bivalvia, Veneridae) with emphasis on the systematic position of taxa along the coast of mainland China. Zool. Scr. 40, 260–271 (2011).
Wang, X., Kong, L., Chen, J., Matsukuma, A. & Li, Q. Phylogeography of bivalve Meretrix petechialis in the Northwestern Pacific indicated by mitochondrial and nuclear DNA data. PLOS ONE 12, e0183221 (2017).
Zhang, S. P., Wang, H. X. & Xu, F. S. Taxonomic study on Meretrix (Bivalvia, Veneridae) from China seas. Acta Zootaxononomica Sin. 37 (2012).
Katoh, K. & Standley, D. M. MAFFT Multiple Sequence Alignment Software Version 7: Improvements in Performance and Usability. Mol. Biol. Evol. 30, 772–780 (2013).
Price, M. N., Dehal, P. S. & Arkin, A. P. FastTree 2 – Approximately Maximum-Likelihood Trees for Large Alignments. PLOS ONE 5, e9490 (2010).
Cheng, H., Concepcion, G. T., Feng, X., Zhang, H. & Li, H. Haplotype-resolved de novo assembly using phased assembly graphs with hifiasm. Nat. Methods 18, 170–175 (2021).
Guan, D. et al. Identifying and removing haplotypic duplication in primary genome assemblies. Bioinformatics 36, 2896–2898 (2020).
Laetsch, D. R. & Blaxter, M. L. BlobTools: Interrogation of genome assemblies. Preprint at https://doi.org/10.12688/f1000research.12232.1 (2017).
Zhou, C., McCarthy, S. A. & Durbin, R. YaHS: yet another Hi-C scaffolding tool. Bioinformatics 39, btac808 (2023).
Durand, N. C. et al. Juicebox Provides a Visualization System for Hi-C Contact Maps with Unlimited Zoom. Cell Syst. 3, 99–101 (2016).
Astashyn, A. et al. Rapid and sensitive detection of genome contamination at scale with FCS-GX. Genome Biol. 25, 60 (2024).
Hoff, K. J., Lomsadze, A., Borodovsky, M. & Stanke, M. Whole-Genome Annotation with BRAKER. in Gene Prediction: Methods and Protocols (ed. Kollmar, M.) 65–95. https://doi.org/10.1007/978-1-4939-9173-0_5 (Springer, New York, NY, 2019).
Stanke, M., Diekhans, M., Baertsch, R. & Haussler, D. Using native and syntenically mapped cDNA alignments to improve de novo gene finding. Bioinformatics 24, 637–644 (2008).
Baril, T., Galbraith, J. & Hayward, A. Earl Grey: A Fully Automated User-Friendly Transposable Element Annotation and Analysis Pipeline. Mol. Biol. Evol. 41, msae068 (2024).
Emms, D. M. & Kelly, S. OrthoFinder: phylogenetic orthology inference for comparative genomics. Genome Biol. 20, 238 (2019).
Wei, M. et al. Chromosome-Level Clam Genome Helps Elucidate the Molecular Basis of Adaptation to a Buried Lifestyle. iScience 23, 101148 (2020).
Song, H. et al. The hard clam genome reveals massive expansion and diversification of inhibitors of apoptosis in Bivalvia. BMC Biol. 19, 15 (2021).
McCartney, M. A. et al. The genome of the zebra mussel, Dreissena polymorpha: a resource for comparative genomics, invasion genetics, and biocontrol. G3 GenesGenomesGenetics 12, jkab423 (2022).
Li, J. et al. Chromosome-level genome assembly and annotation of rare and endangered tropical bivalve, Tridacna crocea. Sci. Data 11, 186 (2024).
Guo, Y. et al. Hologenome analysis reveals independent evolution to chemosymbiosis by deep-sea bivalves. BMC Biol. 21, 51 (2023).
Bai, Y. et al. Multi-omic insights into the formation and evolution of a novel shell microstructure in oysters. BMC Biol. 21, 204 (2023).
Regan, T., Hori, T. S. & Bean, T. P. A chromosome-scale Mytilus edulis genome assembly for aquaculture, marine ecology, and evolution. G3 GenesGenomesGenetics 14, jkae138 (2024).
McElroy, K. E., Masonbrink, R., Chudalayandi, S., Severin, A. J. & Serb, J. M. A chromosome-level genome assembly of the disco clam, Ctenoides ales. G3 GenesGenomesGenetics 14, jkae115 (2024).
Sanderson, M. J. r8s: inferring absolute rates of molecular evolution and divergence times in the absence of a molecular clock. Bioinformatics 19, 301–302 (2003).
Hedges, S. B., Dudley, J. & Kumar, S. TimeTree: a public knowledge-base of divergence times among organisms. Bioinformatics 22, 2971–2972 (2006).
Manni, M., Berkeley, M. R., Seppey, M. & Zdobnov, E. M. BUSCO: Assessing Genomic Data Quality and Beyond. Curr. Protoc. 1, e323 (2021).
Lovell, J. T. et al. GENESPACE tracks regions of interest and gene copy number variation across multiple genomes. eLife 11, e78526 (2022).
Wang, Y. et al. MCScanX: a toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic Acids Res. 40, e49 (2012).
Li, H. Minimap2: pairwise alignment for nucleotide sequences. Bioinformatics 34, 3094–3100 (2018).
Robinson, J. T., Thorvaldsdottir, H., Turner, D. & Mesirov, J. P. igv.js: an embeddable JavaScript implementation of the Integrative Genomics Viewer (IGV). Bioinformatics 39, btac830 (2023).
NCBI GenBank https://identifiers.org/ncbi/insdc:JBTXPG000000000 (2026).
European Nucleotide Archive https://www.ebi.ac.uk/ena/browser/view/GCA_055551385.1 (2026).
NCBI GenBank https://identifiers.org/ncbi/insdc:JBTORC000000000 (2026).
European Nucleotide Archive https://www.ebi.ac.uk/ena/browser/view/GCA_055550865.1 (2026).
DNA Data Bank of Japan https://ddbj.nig.ac.jp/search/entry/bioproject/PRJDB40269 (2026).
DNA Data Bank of Japan https://ddbj.nig.ac.jp/search/entry/bioproject/PRJDB40270 (2026).
National Genomics Data Center https://ngdc.cncb.ac.cn/gwh/Assembly/111533/show (2026).
National Genomics Data Center https://ngdc.cncb.ac.cn/gwh/Assembly/111604/show (2026).
Law, S. T. S. et al. Chromosome-level genomes of hard clams Meretrix lamarckii (Deshayes, 1853) and Meretrix meretrix (Linnaeus, 1758). CUHK Research Data Repository https://doi.org/10.48668/DOVLYS (2025).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:SRP617717 (2026).
European Nucleotide Archive https://www.ebi.ac.uk/ena/browser/view/PRJNA1312090 (2026).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:SRP618173 (2026).
European Nucleotide Archive https://www.ebi.ac.uk/ena/browser/view/PRJNA1315236 (2026).
DNA Data Bank of Japan https://ddbj.nig.ac.jp/search/entry/sra-run/DRR911404 (2026).
DNA Data Bank of Japan https://ddbj.nig.ac.jp/search/entry/sra-run/DRR911405 (2026).
DNA Data Bank of Japan https://ddbj.nig.ac.jp/search/entry/sra-run/DRR911157 (2026).
DNA Data Bank of Japan https://ddbj.nig.ac.jp/search/entry/sra-run/DRR911158 (2026).
National Genomics Data Center https://ngdc.cncb.ac.cn/gsa/browse/CRA039134 (2026).
National Genomics Data Center https://ngdc.cncb.ac.cn/gsa/browse/CRA039136 (2026).
Acknowledgements
This work was supported by Lantau Conservation Fund (RE-2020-39), Hong Kong Research Grant Council Collaborative Research Fund (C4015-20EF), Innovation Technology Fund of Innovation Technology Commission: Funding Support to State Key Laboratory of Agrobiotechnology, and Direct Grant of The Chinese University of Hong Kong.
Author information
Authors and Affiliations
Contributions
S.Y.L., S.G.C. and J.H.L.H. conceived and supervised the study; S.T.S.L., M.F.F.A., L.H.T.C. and C.W.Y.S. carried out sample collection; S.T.S.L. and W.N. performed data curation on the analysis; J.H.L.H. and S.T.S.L. wrote the initial manuscript; all authors revised and contributed to the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Law, S.T.S., Nong, W., Au, M.F.F. et al. Chromosome-level genomes of hard clams Meretrix lamarckii (Deshayes, 1853) and Meretrix meretrix (Linnaeus, 1758). Sci Data (2026). https://doi.org/10.1038/s41597-026-07119-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41597-026-07119-0