Abstract
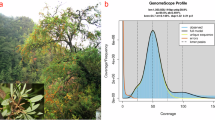
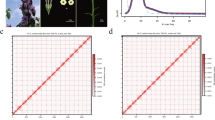
Piptanthus nepalensis (Hook.) Sweet is a leguminous shrub native to the Himalayan region and is valued for its medicinal uses and ornamental yellow flowers. Here, we report a chromosome-scale genome assembly of P. nepalensis generated using PacBio HiFi long reads, Illumina short reads, and Hi-C chromatin interaction data. The assembled genome spans approximately 1.04 Gb, with a contig N50 of 39.5 Mb and a scaffold N50 of 111.5 Mb. Benchmarking universal single-copy orthologs (BUSCO) analysis indicated high completeness for both the genome (99.3%) and predicted protein set (99.2%). Approximately 99.0% of the assembled sequences were anchored onto nine pseudo-chromosomes. We annotated 26,035 protein-coding genes, and repetitive elements were estimated to comprise ~75.2% of the genome. This high-quality genome assembly provides a foundational genomic resource for P. nepalensis and supports future studies on its biology and utilization.
Similar content being viewed by others
Data availability
The raw Illumina, PacBio, Hi-C, and RNA-seq sequencing data are available in the NCBI Sequence Read Archive (SRA) under accession number SRP679460. The final chromosome-level genome assembly is available in NCBI GenBank under accession number JBVQTV000000000. The genome annotation files are available in Figshare (https://doi.org/10.6084/m9.figshare.30416158.v1).
Code availability
All software and pipelines were implemented in full compliance with the manuals and protocols specified by the respective published bioinformatics tools. No custom programming or coding was employed.
References
Wu, Z. Y., Raven, P. H. & Hong, D. Y. (eds.) Flora of China, Vol. 10 (Fabaceae). Beijing: Science Press; St. Louis: Missouri Botanical Garden Press (2010).
Bhattarai, S. & Basukala, O. Antibacterial activity of selected ethnomedicinal plants of Sagarmatha region of Nepal. Int. J. Ther. Appl. 31, 27–31, https://doi.org/10.20530/IJTA_31_27-31 (2016).
Tamang, R. et al. Ethnomedicinal uses of plants and their antibacterial activities in Solukhumbu district, eastern Nepal. BMC Complement. Med. Ther. 20, 201, https://doi.org/10.1186/s12906-020-02986-9 (2020).
Paris, R. R., Faugeras, G. & Dobremez, J.-F. Isoflavones des rameaux de Piptanthus nepalensis. Planta Med. 29(1), 32–36, https://doi.org/10.1055/s-0028-1097625 (1976).
Sener, B. et al. Quinolizidine alkaloids in the Leguminosae: distribution and biological activities. Phytochem. Rev. 2, 123–136, https://doi.org/10.1023/A:1026064906835 (2003).
Wink, M. Evolution of secondary metabolites from an ecological and molecular phylogenetic perspective. Phytochemistry 64, 3–19, https://doi.org/10.1016/S0031-9422(03)00300-5 (2003).
Sun, H. et al. The complete chloroplast genome of Piptanthus nepalensis, a medicinal plant. Mitochondrial DNA B Resour. 7, 796–797, https://doi.org/10.1080/23802359.2021.1994897 (2022).
Chen, S. et al. Herbal genomics: examining the biology of traditional medicines. Science 347(6219), S27–S29 (2015).
Liu, X. et al. The genome of the medicinal plant Macleaya cordata provides new insights into benzylisoquinoline alkaloid metabolism. Mol. Plant 10, 975–989, https://doi.org/10.1016/j.molp.2017.05.007 (2017).
Doyle, J. J. & Doyle, J. L. A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochemical Bulletin 19, 11–15 (1987).
Chen, S., Zhou, Y., Chen, Y. & Gu, J. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34, i884–i890, https://doi.org/10.1093/bioinformatics/bty560 (2018).
Belton, J. M. et al. Hi-C: a comprehensive technique to capture the conformation of genomes. Methods 58, 268–276, https://doi.org/10.1016/j.ymeth.2012.05.001 (2012).
Marçais, G. & Kingsford, C. A fast, lock-free approach for efficient parallel counting of occurrences of k-mers. Bioinformatics 27, 764–770, https://doi.org/10.1093/bioinformatics/btr011 (2011).
Vurture, G. W. et al. GenomeScope: fast reference-free genome profiling from short reads. Bioinformatics 33, 2202–2204, https://doi.org/10.1093/bioinformatics/btx153 (2017).
Ranallo-Benavidez, T. R., Jaron, K. S. & Schatz, M. C. GenomeScope 2.0 and Smudgeplot for reference-free profiling of polyploid genomes. Nat. Commun. 11, 1432, https://doi.org/10.1038/s41467-020-14998-3 (2020).
Cheng, H., Concepcion, G. T., Feng, X., Zhang, H. & Li, H. Hifiasm: a haplotype-resolved assembler for accurate HiFi reads. Nat. Methods 18, 170–175, https://doi.org/10.1038/s41592-020-01056-5 (2021).
Li, H. Minimap2: pairwise alignment for nucleotide sequences. Bioinformatics 34, 3094–3100, https://doi.org/10.1093/bioinformatics/bty191 (2018).
Camacho, C. et al. BLAST+: architecture and applications. BMC Bioinformatics 10, 421, https://doi.org/10.1186/1471-2105-10-421 (2009).
Guan, D. et al. Identifying and removing haplotypic duplication in primary genome assemblies. Bioinformatics 36, 2896–2898, https://doi.org/10.1093/bioinformatics/btaa025 (2020).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760, https://doi.org/10.1093/bioinformatics/btp324 (2009).
Zeng, X. et al. Chromosome-level scaffolding of haplotype-resolved assemblies using Hi-C data without reference genomes. Nat. Plants 10, 1184–1200, https://doi.org/10.1038/s41477-024-01755-3 (2024).
Xu, M. et al. TGS-GapCloser: a fast and accurate gap closer for large genomes with third-generation sequencing reads. GigaScience 9, giaa067, https://doi.org/10.1093/gigascience/giaa067 (2020).
Hu, J. et al. NextPolish2: A repeat-aware polishing tool for genomes assembled using HiFi long reads. Genomics Proteomics Bioinformatics 22(1), qzad009, https://doi.org/10.1093/gpbjnl/qzad009 (2024).
Rhie, A. et al. Merqury: reference-free quality, completeness, and phasing assessment for genome assemblies. Genome Biol. 21, 245, https://doi.org/10.1186/s13059-020-02134-9 (2020).
Ou, S. et al. Assessing genome assembly quality using the LTR Assembly Index (LAI). Nucleic Acids Res. 46, e126, https://doi.org/10.1093/nar/gky730 (2018).
Simão, F. A., Waterhouse, R. M., Ioannidis, P., Kriventseva, E. V. & Zdobnov, E. M. BUSCO: assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics 31, 3210–3212, https://doi.org/10.1093/bioinformatics/btv351 (2015).
Xu, Z. & Wang, H. LTR_FINDER: an efficient tool for the prediction of full-length LTR retrotransposons. Nucleic Acids Res. 35, W265–W268, https://doi.org/10.1093/nar/gkm286 (2007).
Price, A. L., Jones, N. C. & Pevzner, P. A. De novo identification of repeat families in large genomes. Bioinformatics 21, i351–i358, https://doi.org/10.1093/bioinformatics/bti1018 (2005).
Flynn, J. M. et al. RepeatModeler2 for automated genomic discovery of transposable element families. Proc. Natl Acad. Sci. USA 117, 9451–9457, https://doi.org/10.1073/pnas.1921046117 (2020).
Tarailo-Graovac, M. & Chen, N. Using RepeatMasker to identify repetitive elements in genomic sequences. Curr. Protoc. Bioinformatics 25, 4.10.1–4.10.14, https://doi.org/10.1002/0471250953.bi0410s25 (2009).
Jurka, J. et al. Repbase Update, a database of eukaryotic repetitive elements. Cytogenet. Genome Res. 110, 462–467, https://doi.org/10.1159/000084979 (2005).
Benson, G. Tandem repeats finder: a program to analyze DNA sequences. Nucleic Acids Res. 27, 573–580, https://doi.org/10.1093/nar/27.2.573 (1999).
Kim, D., Langmead, B. & Salzberg, S. L. HISAT2: graph-based alignment of next-generation sequencing reads to a population of genomes. Nat. Methods 12, 357–360, https://doi.org/10.1038/nmeth.3317 (2015).
Danecek, P. et al. Twelve years of SAMtools and BCFtools. GigaScience 10, giab008, https://doi.org/10.1093/gigascience/giab008 (2021).
Kovaka, S. et al. Transcriptome assembly from long-read RNA-seq alignments with StringTie2. Genome Biol. 20, 278, https://doi.org/10.1186/s13059-019-1910-1 (2019).
Brůna, T., Hoff, K. J., Lomsadze, A., Stanke, M. & Borodovsky, M. BRAKER2: automatic eukaryotic genome annotation with GeneMark-EP+ and AUGUSTUS supported by a protein database. NAR Genomics Bioinformatics 3, lqaa108, https://doi.org/10.1093/nargab/lqaa108 (2021).
Stanke, M. et al. AUGUSTUS: ab initio prediction of alternative transcripts. Nucleic Acids Res. 34, W435–W439, https://doi.org/10.1093/nar/gkl200 (2006).
Lomsadze, A., Ter-Hovhannisyan, V., Chernoff, Y. O. & Borodovsky, M. Gene identification in novel eukaryotic genomes by self-training algorithm. Nucleic Acids Res. 33, 6494–6506, https://doi.org/10.1093/nar/gki937 (2005).
O’Leary, N. A. et al. Reference sequence (RefSeq) database at NCBI: current status, taxonomic expansion, and functional annotation. Nucleic Acids Res. 44, D733–D745, https://doi.org/10.1093/nar/gkv1189 (2016).
The UniProt Consortium. UniProt: the Universal Protein Knowledgebase in 2023. Nucleic Acids Res. 51, D523–D531, https://doi.org/10.1093/nar/gkac1052 (2023).
Huerta-Cepas, J. et al. eggNOG 5.0: a hierarchical, functionally and phylogenetically annotated orthology resource. Nucleic Acids Res. 47, D309–D314, https://doi.org/10.1093/nar/gky1085 (2019).
Tatusov, R. L. et al. The COG database: an updated version includes eukaryotes. BMC Bioinformatics 11, 41, https://doi.org/10.1186/1471-2105-11-41 (2010).
Eddy, S. R. Accelerated profile HMM searches. PLoS Comput. Biol. 7, e1002195, https://doi.org/10.1371/journal.pcbi.1002195 (2011).
Mistry, J. et al. Pfam: the protein families database in 2021. Nucleic Acids Res. 49, D412–D419, https://doi.org/10.1093/nar/gkaa913 (2021).
Ashburner, M. et al. Gene ontology: tool for the unification of biology. Nat. Genet. 25, 25–29, https://doi.org/10.1038/75556 (2000).
Kanehisa, M. & Goto, S. KEGG: Kyoto Encyclopedia of Genes and Genomes. Nucleic Acids Res. 28, 27–30, https://doi.org/10.1093/nar/28.1.27 (2000).
Seemann, T. Barrnap 0.9: rapid ribosomal RNA prediction. https://github.com/tseemann/barrnap (2014).
Chan, P. P. & Lowe, T. M. tRNAscan-SE: searching for tRNA genes in genomic sequences. Nucleic Acids Res. 47, 1–14, https://doi.org/10.1093/nar/gky1048 (2019).
Ontiveros-Palacios, N. et al. Rfam 15: RNA families database in 2025. Nucleic Acids Res. 53(D1), D258–D267, https://doi.org/10.1093/nar/gkae1023 (2025).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:SRP679460 (2026).
Zhang, C. et al. Piptanthus nepalensis Genome sequencing and assembly. Genbank https://identifiers.org/ncbi/insdc:JBVQTV000000000 (2026).
Zhang, J. et al. The genome annotation files of Piptanthus nepalensis. Figshare. https://doi.org/10.6084/m9.figshare.30416158.v1 (2026).
Acknowledgements
The computations in this research were performed using the CFFF platform of Fudan University. This work was supported by the Science and Technology Projects of Xizang Autonomous Region, China (Project Nos. XZ202402ZD0005, XZ202402ZY0023, XZ202402JX0003, XZ202401ZR0028, and XZ202303ZY0002G), and by the Open Project of the Key Laboratory of Biodiversity and Environment on the Qinghai-Tibet Plateau, Ministry of Education (Project No. KLBE2025003).
Author information
Authors and Affiliations
Contributions
Y.W., J.W., and L.Q. conceived and designed the research framework. J.Z., and Z.Z. collected the samples, prepared experimental materials, and conducted laboratory work. N.B., N.N., N.T., and X.T. contributed to data acquisition, curation, and preliminary analyses. Z.Z. and J.Z. performed the formal data analysis and drafted the initial manuscript, including figures and tables. Y.W., J.W., and L.Q. critically revised the manuscript for intellectual content and provided guidance throughout the study. All authors contributed to improving the manuscript and approved the final version for publication.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhang, J., Zeng, Z., Bonjor, N. et al. Chromosome-Level Genome Assembly and Annotation of Piptanthus nepalensis (Hook.) Sweet. Sci Data (2026). https://doi.org/10.1038/s41597-026-07134-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41597-026-07134-1