Abstract
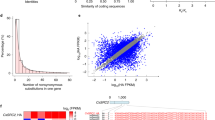
Tea is one of the oldest crops in the world and is of great economic value as a beverage. Its natural flavor and compounds related to health exhibit significant genetic diversity. This article reports the chromosome-scale genome assembly map of a new Camellia sinensis ‘Yuwan Xiaoye’ (YWXY) bred by our research center. The genome size is 3.18 Gb with a contig N50 of 181.8 Mb, and 93.43% of the assembled sequences were anchored to 15 chromosomes. The genome is predicted to contain 40,119 protein-coding genes, with 99.70% having functional annotations. Repeat elements account for approximately 82.21% of the genomic landscape. The completeness of YWXY genome assembly is highlighted by a BUSCO score of 99.07%. The assembled genome provides a critical resource for molecular breeding and functional studies in tea plants.
Similar content being viewed by others

Data availability
All software and pipelines were executed according to the manual and protocols of the published bioinformatic tools. All software used in this work is publicly available, with versions and parameters clearly described in Methods. If no detailed parameters were mentioned for a software, the default parameters suggested by the developer were used. No custom code was used during this study for the curation and/or validation of the datasets.
Code availability
All commands and pipelines used in data processing were executed according to the manual and protocols of the corresponding bioinformatics software. No specific code has been developed for this study.
References
Xia, E. et al. The Tea Tree Genome Provides Insights into Tea Flavor and Independent Evolution of Caffeine Biosynthesis. Mol. Plant. 10, 866–877 (2017).
Wei, K. et al. A coupled role for CsMYB75 and CsGSTF1 in anthocyanin hyperaccumulation in purple tea. The Plant Journal: For Cell and Molecular Biology. 97, 825–840 (2019).
Pastoriza, S., Mesías, M., Cabrera, C. & Rufián-Henares, J. A. Healthy properties of green and white teas: an update. Food Funct. 8, 2650–2662 (2017).
Zhang, Z. et al. Understanding the Origin and Evolution of Tea (Camellia sinensis [L.]): Genomic Advances in Tea. J. Mol. Evol. 91, 156–168 (2023).
Zhang, Q. et al. The Chromosome-Level Reference Genome of Tea Tree Unveils Recent Bursts of Non-autonomous LTR Retrotransposons in Driving Genome Size Evolution. pp. 935–938 (2020).
Zhang, W. et al. Genome assembly of wild tea tree DASZ reveals pedigree and selection history of tea varieties. Nat. Commun. 11, 3719 (2020).
Zhang, X. et al. Haplotype-resolved genome assembly provides insights into evolutionary history of the tea plant Camellia sinensis. Nat. Genet. 53, 1250–1259 (2021).
Kong, W., Yu, J., Yang, J., Zhang, Y. & Zhang, X. The high-resolution three-dimensional (3D) chromatin map of the tea plant (Camellia sinensis). Hortic. Res. 10, uhad179 (2023).
Chen, S. et al. Gene mining and genomics-assisted breeding empowered by the pangenome of tea plant Camellia sinensis. Nat. Plants. 9, 1986–1999 (2023).
Tariq, A. et al. In-depth exploration of the genomic diversity in tea varieties based on a newly constructed pangenome of Camellia sinensis. The Plant Journal: For Cell and Molecular Biology. 119, 2096–2115 (2024).
Yu, X. et al. Metabolite signatures of diverse Camellia sinensis tea populations. Nat. Commun. 11, 5586 (2020).
Jeyaraj, A. et al. Genome-wide identification of conserved and novel microRNAs in one bud and two tender leaves of tea plant (Camellia sinensis) by small RNA sequencing, microarray-based hybridization and genome survey scaffold sequences. Bmc Plant Biol. 17, 212 (2017).
Wang, X. et al. Population sequencing enhances understanding of tea plant evolution. Nat. Commun. 11, 4447 (2020).
Xia, E. et al. The Reference Genome of Tea Plant and Resequencing of 81 Diverse Accessions Provide Insights into Its Genome Evolution and Adaptation. Mol. Plant. 13, 1013–1026 (2020).
Kong, W. et al. Pan-transcriptome assembly combined with multiple association analysis provides new insights into the regulatory network of specialized metabolites in the tea plant Camellia sinensis. Hortic. Res. 9, uhac100 (2022).
Kong, W. et al. Genomic analysis of 1,325 Camellia accessions sheds light on agronomic and metabolic traits for tea plant improvement. Nat. Genet. 57, 997–1007 (2025).
Xiao, C. et al. MECAT: fast mapping, error correction, and de novo assembly for single-molecule sequencing reads. Nat. Methods. 14, 1072–1074 (2017).
Chin, C. et al. Phased diploid genome assembly with single-molecule real-time sequencing. Nat. Methods. 13, 1050–1054 (2016).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics. 30, 2114–2120 (2014).
Chen, Y. et al. SOAPnuke: a MapReduce acceleration-supported software for integrated quality control and preprocessing of high-throughput sequencing data. Gigascience. 7, 1–6 (2018).
Liu, B. et al. Estimation of genomic characteristics by analyzing k-mer frequency in de novo genome projects. Quant. Biol. (2013).
Cheng, H., Concepcion, G. T., Feng, X., Zhang, H. & Li, H. Haplotype-resolved de novo assembly using phased assembly graphs with hifiasm. Nat. Methods. 18, 170–175 (2021).
Dudchenko, O. et al. De novo assembly of the Aedes aegypti genome using Hi-C yields chromosome-length scaffolds. Science (New York, N.Y.). 356, 92–95 (2017).
Burton, J. N. et al. Chromosome-scale scaffolding of de novo genome assemblies based on chromatin interactions. Nat. Biotechnol. 31, 1119–1125 (2013).
Flynn, J. M. et al. RepeatModeler2 for automated genomic discovery of transposable element families. Proc. Natl. Acad. Sci. USA. 117, 9451–9457 (2020).
Bao, Z. & Eddy, S. R. Automated de novo identification of repeat sequence families in sequenced genomes. Genome Res. 12, 1269–1276 (2002).
Price, A. L., Jones, N. C. & Pevzner, P. A. De novo identification of repeat families in large genomes. Bioinformatics. 21(Suppl 1), i351–i358 (2005).
Xu, Z. & Wang, H. LTR_FINDER: an efficient tool for the prediction of full-length LTR retrotransposons. Nucleic. Acids. Res. 35, W265–W268 (2007).
Benson, G. Tandem repeats finder: a program to analyze DNA sequences. Nucleic. Acids. Res. 27, 573–580 (1999).
Jurka, J. et al. Repbase Update, a database of eukaryotic repetitive elements. Cytogenet. Genome Res. 110, 462–467 (2005).
Jurka, J. Repbase update: a database and an electronic journal of repetitive elements. Trends in Genetics: Tig. 16, 418–420 (2000).
Stanke, M. & Morgenstern, B. AUGUSTUS: a web server for gene prediction in eukaryotes that allows user-defined constraints. Nucleic. Acids. Res. 33, W465–W467 (2005).
Stanke, M. et al. AUGUSTUS: ab initio prediction of alternative transcripts. Nucleic. Acids. Res. 34, W435–W439 (2006).
Burge, C. & Karlin, S. Prediction of complete gene structures in human genomic DNA. J. Mol. Biol. 268, 78–94 (1997).
Majoros, W. H., Pertea, M. & Salzberg, S. L. TigrScan and GlimmerHMM: two open source ab initio eukaryotic gene-finders. Bioinformatics (Oxford, England). 20, 2878–2879 (2004).
Slater, G. S. C. & Birney, E. Automated generation of heuristics for biological sequence comparison. Bmc Bioinformatics. 6, 31 (2005).
Pertea, M. et al. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat. Biotechnol. 33, 290–295 (2015).
Haas, B. J. et al. Improving the Arabidopsis genome annotation using maximal transcript alignment assemblies. Nucleic. Acids. Res. 31, 5654–5666 (2003).
Cantarel, B. L. et al. MAKER: an easy-to-use annotation pipeline designed for emerging model organism genomes. Genome Res. 18, 188–196 (2008).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Camacho, C. et al. BLAST+: architecture and applications. Bmc Bioinformatics. 10, 421 (2009).
Kanehisa, M., Goto, S., Sato, Y., Furumichi, M. & Tanabe, M. KEGG for integration and interpretation of large-scale molecular data sets. Nucleic. Acids. Res. 40, D109–D114 (2012).
Ashburner, M. et al. Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat. Genet. 25, 25–29 (2000).
Boeckmann, B. et al. The SWISS-PROT protein knowledgebase and its supplement TrEMBL in 2003. Nucleic. Acids. Res. 31, 365–370 (2003).
Bairoch, A. & Apweiler, R. The SWISS-PROT protein sequence database and its supplement TrEMBL in 2000. Nucleic. Acids. Res. 28, 45–48 (2000).
Mitchell, A. et al. The InterPro protein families database: the classification resource after 15 years. Nucleic. Acids. Res. 43, D213–D221 (2015).
Lowe, T. M. & Eddy, S. R. tRNAscan-SE: a program for improved detection of transfer RNA genes in genomic sequence. Nucleic. Acids. Res. 25, 955–964 (1997).
Lagesen, K. et al. RNAmmer: consistent and rapid annotation of ribosomal RNA genes. Nucleic. Acids. Res. 35, 3100–3108 (2007).
Nawrocki, E. P., Kolbe, D. L. & Eddy, S. R. Infernal 1.0: inference of RNA alignments. Bioinformatics (Oxford, England). 25, 1335–1337 (2009).
Griffiths-Jones, S. et al. Rfam: annotating non-coding RNAs in complete genomes. Nucleic. Acids. Res. 33, D121–D124 (2005).
NCBI Sequence Read Archiv https://www.ncbi.nlm.nih.gov/sra/SRP679934 (2026).
Zhang, W. Camellia sinensis isolate YWXY, whole genome shotgun sequencing project. Genebank https://identifiers.org/ncbi/insdc:JBVOCX000000000 (2026).
CNCB Genome Warehouse https://ngdc.cncb.ac.cn/gwh/Assembly/98825/show (2026).
Manni, M., Berkeley, M. R., Seppey, M., Simão, F. A. & Zdobnov, E. M. BUSCO Update: Novel and Streamlined Workflows along with Broader and Deeper Phylogenetic Coverage for Scoring of Eukaryotic, Prokaryotic, and Viral Genomes. Mol. Biol. Evol. 38, 4647–4654 (2021).
Acknowledgements
This work was supported by the Henan Province Central Leading Local Science and Technology Development Fund Project Funding (Z20231811160).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhang, W., Chen, Y. Chromosome level genome assembly of Camellia sinensis ‘Yuwan Xiaoye’. Sci Data (2026). https://doi.org/10.1038/s41597-026-07142-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41597-026-07142-1