Figure 1
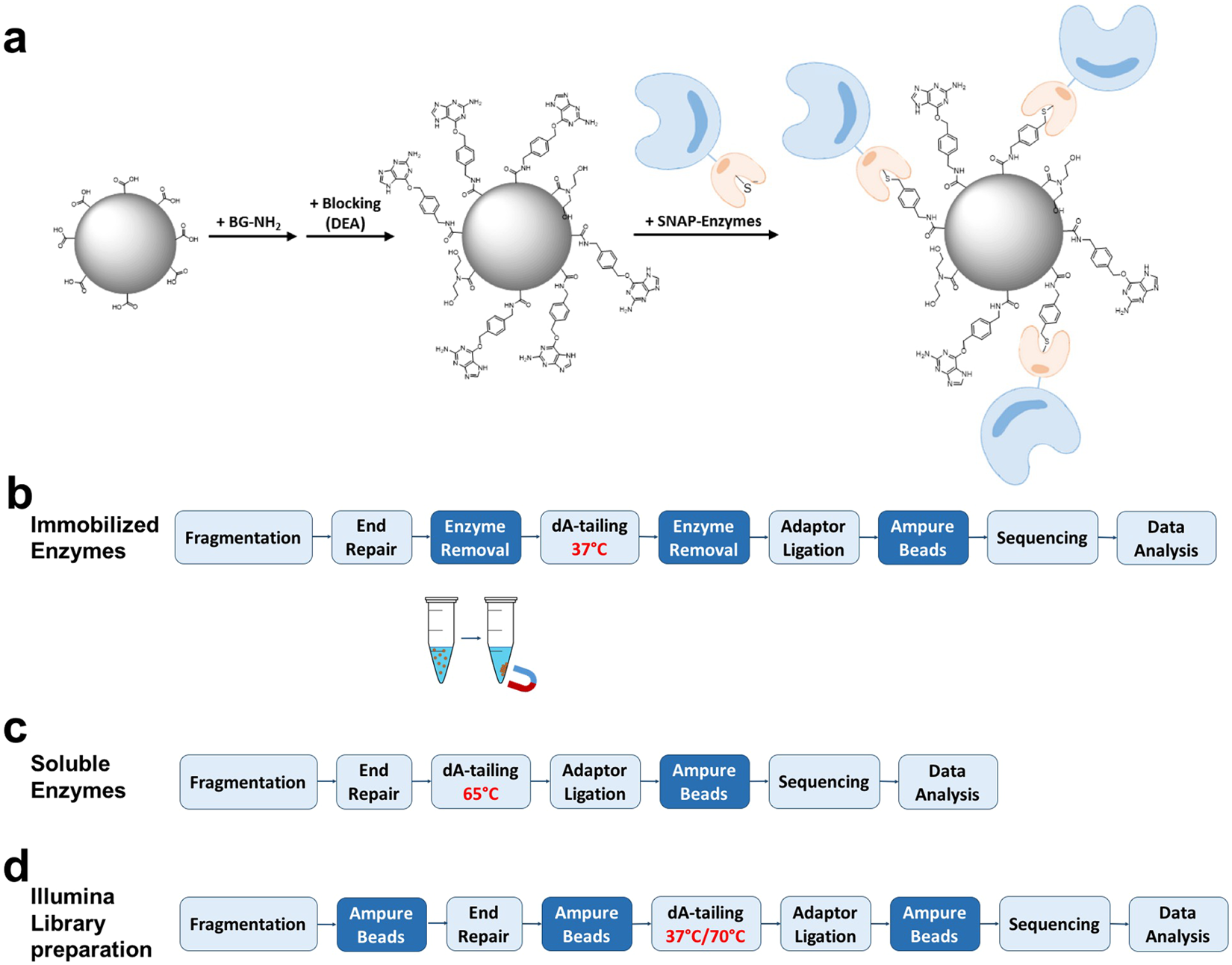
Enzyme immobilization and comparison of Illumina library preparation protocols. (a) A schematic of covalent conjugation of SNAP-tagged enzyme fusion proteins to magnetic beads functionalized with O6-benzylguanine (BG) moieties that specifically react with active site cysteine residues of SNAP-tag proteins, forming a stable covalent thioether bond15,16. (b) Workflow for library construction using immobilized enzymes for Illumina sequencing. A typical streamlined protocol for Illumina library construction is modified by employing immobilized enzymes to catalyze end repair and 3′ A-tailing. This method utilizes SNAP-tagged T4 DNA pol and PNK covalently conjugated to BG-functionalized magnetic beads to carry out end repair of fragmented DNA at 20°C (or 37 °C) for 30 min. The enzymes are removed by magnetic separation from the DNA pool, which is subjected to 3′ A-tailing at 37 °C for 30 min using immobilized Taq DNA pol. (c) Streamlined protocol for Illumina amplification-free library preparation using soluble enzymes. Typically, end repair and 3′ A-tailing of fragmented DNA are catalyzed by an enzyme mixture at 20 °C for 30 min, followed by heat treatment at 65 °C for 30 min. (d) The workflow of Illumina TruSeq DNA PCR-free LT Library Prep Kit with a purification step. End repair is performed for 30 min at 30 °C, followed by a bead-based step for clean up and size selection. 3′ A-tailing is carried out for 30 min at 37 °C with a subsequent treatment for 5 min at 70 °C. Each library was ligated to preannealed full-length paired-end Illumina adaptors, size-selected and analyzed, and sequenced on an Illumina sequencing platform.
