Abstract
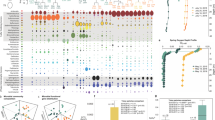
The pelagic environment represents a mosaic of biogeographical domains shaped by regional oceanographic processes. Here, a coastal-to-open ocean microbiome investigation was conducted from 64 water samples of the Santos Basin (SB), located in the subtropical South Atlantic Ocean. We combined shotgun metagenomics with a hybrid machine learning workflow to investigate the taxonomic diversity, community structure, and ecosystem functions of pelagic microbiomes. The workflow integrated self-organizing maps (unsupervised) for pattern discovery and Random Forest (supervised) for predictive modeling. Unsupervised machine learning revealed a clear spatial and vertical (light-driven) distribution, with indicator taxa reflecting biogeochemical patterns consistent with global surveys. Supervised learning identified phosphate, salinity, and nitrate, influenced by local upwelling and La Plata River plume, as the primary environmental drivers of microbial community structure. In terms of functionality, the SB microbiome displayed depth- and region-specific patterns: photoautotrophs and nitrogen fixers dominated photic waters (with differences between coastal and oceanic stations), whereas chemolithoautotrophs and mixotrophs prevailed in the aphotic zone. Notably, nitrification signatures were more frequent in northern mesopelagic communities, while sulfur-oxidation pathways were enriched toward the south. Genes for CO bio-oxidation and dimethylsulfoniopropionate (DMSP) degradation were present across all depths. Furthermore, potential non-cyanobacterial diazotrophs were detected in the deep waters, underscoring previous underappreciated to nitrogen cycling. Our findings indicated that the Santos Basin hosts a functionally diverse microbiome including putative novel lineages. The taxonomic and functional patterns observed in the SB might provide insights into potential ecological responses to shifts in nutrient dynamics and physical processes. This investigation provides an ecogenomic baseline for understanding the microbial ecosystem services in subtropical oceans and reveals the potential of machine learning to uncover ecological patterns in underexplored marine regions.
Similar content being viewed by others
Data availability
The sequencing reads dataset generated and analyzed during the current study are available in the National Centre for Biotechnology Information (NCBI) repositor under the BioProject PRJNA1191028. The genomic dataset generated during and analyzed during the current study are available in the Figshare repositor under https://doi.org/10.6084/m9.figshare.30043690. The iMESc and iMESc Savepoints used for the machine learning analyses are available at Github via https://github.com/DaniloCVieira/imesc_savepoints/tree/2c5a4441ad447c71cc8f5d3e11811fb5a76a3427/Bergo_2025/Microbial-signatures-define-the-ecosystem-functions-of-the-pelagic-microbiome-in-a-basin-scale-Southwest-Atlantic-Ocean
References
Stal, L. J. A Sea of Microbes: What’s So Special about Marine Microbiology. In The Marine Microbiome (eds Stal, L. J. & Cretoiu, M. S.) 1–44 (Springer, 2022).
Jiao, N. et al. The microbial carbon pump and climate change. Nat. Rev. Microbiol. 22, 408–419 (2024).
Li, W. K. W. Primary production of prochlorophytes, cyanobacteria, and eucaryotic ultraphytoplankton: measurements from flow cytometric sorting. Limnol. Oceanogr. 39, 169–175 (1994).
Jardillier, L. et al. Significant CO₂ fixation by small prymnesiophytes in the subtropical and tropical northeast Atlantic Ocean. ISME J. 4, 1180–1192 (2010).
Worden, A. Z. et al. Rethinking the marine carbon cycle: factoring in the multifarious lifestyles of microbes. Science 347, 1257594 (2015).
Buchan, A., LeCleir, G. R., Gulvik, C. A. & González, J. M. Master recyclers: features and functions of bacteria associated with phytoplankton blooms. Nat. Rev. Microbiol. 12, 686–698 (2014).
Baltar, F. & Gerhard, J. H. Ideas and perspectives: is dark carbon fixation relevant for oceanic primary production estimates?. Biogeosciences 16, 3793–3799 (2019).
Falkowski, P. G., Fenchel, T. & Delong, E. F. The microbial engines that drive Earth’s biogeochemical cycles. Science 320, 1034–1039 (2008).
Sunagawa, S. et al. Structure and function of the global ocean microbiome. Science 348, 1261359 (2015).
Tully, B. J., Graham, E. D. & Heidelberg, J. F. The reconstruction of 2,631 draft metagenome-assembled genomes from the global oceans. Sci. Data 5, 170203 (2018).
Yoshitake, K. et al. Development of a time-series shotgun metagenomics database for monitoring microbial communities at the Pacific coast of Japan. Sci. Rep. 11, 12222 (2021).
Acinas, S. G. et al. Deep ocean metagenomes provide insight into the metabolic architecture of bathypelagic microbial communities. Nat. Commun. 12, 604 (2021).
Junger, P. C. et al. Global biogeography of the smallest plankton across ocean depths. Sci. Adv. 9, eadg9763 (2023).
Sánchez, P. et al. Marine picoplankton metagenomes and MAGs from eleven vertical profiles obtained by the Malaspina Expedition. Sci. Data 11, 154 (2024).
Alves, J. N. et al. Microbial community diversity and physical–chemical features of the Southwestern Atlantic Ocean. Arch. Microbiol. 197, 165–179 (2015).
Coutinho, F. H. et al. Ecogenomics and metabolic potential of the South Atlantic Ocean microbiome. Sci. Total Environ. 765, 142758 (2021).
Bergo, N. M., Signori, C. N., Amado, A. M., Brandini, F. P. & Pellizari, V. H. The partitioning of carbon biomass among the pico- and nano-plankton community in the South Brazilian Bight during a strong summer intrusion of South Atlantic central water. Front. Mar. Sci. 4, 238 (2017).
Ribeiro, C. G. et al. Pico and nanoplankton abundance and carbon stocks along the Brazilian Bight. PeerJ 4, e2587 (2016).
Ribeiro, C. G. et al. Estimating microbial populations by flow cytometry: Comparison between instruments. Limnol. Oceanogr. Methods 14, 750–758 (2016).
Agostini, L. et al. Deep-sea plastisphere: long-term colonization by plastic-associated bacterial and archaeal communities in the Southwest Atlantic Ocean. Sci. Total Environ. 793, 148335 (2021).
Ferreira, J. C. N. et al. Abundance and microbial diversity from surface to deep water layers over the Rio Grande Rise. South Atlantic. Prog. Oceanogr. 201, 102736 (2022).
Moreira, J. C. F. et al. A Machine learning approach elucidates spatial patterns of environmental properties driving microbial composition over Santos Basin. South Atlantic bioRxiv https://doi.org/10.1101/2025.02.26.640418 (2025).
Moreira, D. L. et al. Multidisciplinary scientific cruises for environmental characterization in the Santos Basin – methods and sampling design. Ocean Coast. Res. 71, e23022 (2023).
Corrêa, A. M. S. et al. Hydrocarbon composition of oils spilt on the Brazilian coast in 2019 and 2022. Mar. Pollut. Bull. 207, 116821 (2024).
Sartori, É. et al. Trace metal concentration along the Brazilian coast: An assessment of the influence of the Doce River plume. Mar. Pollut. Bull. 188, 114640 (2023).
Senez-Mello, T. M. et al. Assessment of anthropogenic pollution in Guanabara Bay (SE Brazil) through biogeochemical data and stable isotope mixing models. Environ. Sci. Pollut. Res. 31, 32972–32997 (2024).
Ramírez, F. et al. Overfishing species on the move may burden seafood provision in the low-latitude Atlantic Ocean. Sci. Total Environ. 836, 155480 (2022).
Franco, B. C. et al. Climate change impacts on the atmospheric circulation, ocean, and fisheries in the southwest South Atlantic Ocean: a review. Clim. Change 162, 2359–2377 (2020).
Tara Ocean Foundation et al. Priorities for ocean microbiome research. Nat. Microbiol. 7, 937–947 (2022).
Asnicar, F., Thomas, A. M., Passerini, A., Waldron, L. & Segata, N. Machine learning for microbiologists. Nat. Rev. Microbiol. 22, 191–205 (2024).
Prabhu, A. et al. Machine learning and metagenomics identifies uncharacterized taxa inferred to drive biogeochemical cycles in a subtropical hypereutrophic estuary. ISME Commun. 4, 067 (2024).
Pinnaka, A. K. & Tanuku, N. R. S. Marine microbial diversity for sustainable development. In Microbial Diversity in Ecosystem Sustainability and Biotechnological Applications (eds Gupta, J. et al.) 117–158
Bowers, et al. Minimum information about a single amplified genome (MISAG) and a metagenome-assembled genome (MIMAG) of bacteria and archaea. Nat Biotechnol. 35, 725–731 (2017).
Sunagawa, S. et al. Tara Oceans: towards global ocean ecosystems biology. Nat. Rev. Microbiol. 18, 428–445 (2020).
Arístegui, J., Gasol, J. M., Duarte, C. M. & Herndl, G. J. Microbial oceanography of the dark ocean’s pelagic realm. Limnol. Oceanogr. 54, 1501–1529 (2009).
Duarte, 2015 – Malaspina
Zhang, Z. et al. Global biogeography of microbes driving ocean ecological status under climate change. Nat. Commun. 15, 4657 (2024).
Brandini, F. P., Turra, P. M. & Santos, P. P. G. M. Ecosystem responses to biogeochemical fronts in the South Brazil Bight. Prog. Oceanogr. 164, 52–62 (2018).
Xu, X. et al. (2024) Genome-based taxonomic classification of the genus Sulfitobacter along with the proposal of a new genus Parasulfitobacter gen. nov. and exploring the gene clusters associated with sulfur oxidation. BMC Genomics 25: 389.
Moran, M. A., González, J. M. & Kiene, R. P. Linking a bacterial taxon to sulfur cycling in the sea: studies of the marine Roseobacter group. Geomicrobiol. J. 20, 375–388 (2003).
Zeng, Y.-X. et al. Diversity of bacterial dimethylsulfoniopropionate degradation genes in surface seawater of Arctic Kongsfjorden. Sci. Rep. 6, 33031 (2016).
Sanz-Sáez, I. et al. Top abundant deep ocean heterotrophic bacteria can be retrieved by cultivation. ISME Commun. 3, 92 (2023).
Lappan, R. et al. Molecular hydrogen in seawater supports growth of diverse marine bacteria. Nat. Microbiol. 8, 581–595 (2023).
Dede, B. et al. Bacterial chemolithoautotrophy in ultramafic plumes along the Mid-Atlantic Ridge. ISME J. 18, wrae165 (2024).
Giebel, H. A. et al. Complementary energy acquisition via aerobic anoxygenic photosynthesis and carbon monoxide oxidation by Planktomarina temperata of the Roseobacter group. FEMS Microbiol. Ecol. 95. fiz050. (2019)
Cai, H. et al. Aerobic anoxygenic phototrophs play important roles in nutrient cycling within cyanobacterial Microcystisbloom microbiomes. Microbiome 12, 88 (2024).
Luo, H. & Moran, M. A. Evolutionary ecology of the marine Roseobacter clade. Microbiol. Mol. Biol. Rev. 78, 573–587 (2014).
Danovaro, R., Snelgrove, P. V. R. & Tyler, P. Challenging the paradigms of deep-sea ecology. Trends Ecol. Evol. 29, 465–475 (2014).
Xue, C.-X. et al. Insights into the vertical stratification of microbial ecological roles across the deepest seawater column on Earth. Microorganisms 8, 1309 (2020).
Li, J. et al. (2024) Arcobacteraceae are ubiquitous mixotrophic bacteria playing important roles in carbon, nitrogen, and sulfur cycling in global oceans. mSystems 9: 00513–24.
Stoecker, D. K. et al. Mixotrophy in the marine plankton. Annu. Rev. Mar. Sci. 9, 311–335 (2017).
Ward, B. A. & Follows, M. J. Marine mixotrophy increases trophic transfer efficiency, mean organism size, and vertical carbon flux. Proc. Natl Acad. Sci. USA 113, 2958–2963 (2016).
Swan, B. K. et al. Potential for chemolithoautotrophy among ubiquitous bacteria lineages in the dark ocean. Science 333, 1296–1300 (2011).
Baltar, F. et al. A ubiquitous gammaproteobacterial clade dominates expression of sulfur oxidation genes across the mesopelagic ocean. Nat. Microbiol. 8, 1137–1148 (2023).
Teira, E. et al. Distribution and activity of Bacteria and Archaea in the deep water masses of the North Atlantic. Limnol. Oceanogr. 51, 2131–2144 (2006).
Alonso-Sáez, L. et al. High bicarbonate assimilation in the dark by Arctic bacteria. ISME J. 4, 1581–1590 (2010).
La Cono, V. et al. Contribution of bicarbonate assimilation to carbon pool dynamics in the deep mediterranean sea and cultivation of actively nitrifying and CO₂-fixing bathypelagic prokaryotic consortia. Front. Microbiol. 9, 3 (2018).
Zhou, Z., Tran, P. Q., Kieft, K. & Anantharaman, K. Genome diversification in globally distributed novel marine Proteobacteria is linked to environmental adaptation. ISME J. 14, 2060–2077 (2020).
Chakraborty, S., Andersen, K. H., Merico, A. & Riemann, L. Particle-associated N2 fixation by heterotrophic bacteria in the global ocean. Sci. Adv. 11(8), 4693 (2025).
Yuan, J., Lai, Q., Zheng, T. & Shao, Z. Novosphingobium indicum sp. nov., a polycyclic aromatic hydrocarbon-degrading bacterium isolated from a deep-sea environment. Int. J. Syst. Evol. Microbiol. 59, 2084–2088 (2009).
Acids Res. 44, D590-D594 (2016).
Wood, D. E. & Salzberg, S. L. Kraken: ultrafast metagenomic sequence classification using exact alignments. Genome Biol. 15, R46 (2014).
Wood, D. E., Lu, J. & Langmead, B. Improved metagenomic analysis with Kraken 2. Genome Biol. 20, 257 (2019).
Wilke, A. et al. The MG-RAST metagenomics database and portal in 2015. Nucleic Acids Res. 44, D590–D594 (2016).
Keegan, K. P., Glass, E. M. & Meyer, F. MG-RAST, a metagenomics service for analysis of microbial community structure and function. In Microbial Environmental Genomics (MEG) 207–233.
McMurdie, P. J. & Holmes, S. phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. PLoS ONE 8, e61217 (2013).
Wickham, H. ggplot2. Wiley Interdiscip. Rev. Comput. Stat. 3, 180–185 (2011).
Oksanen, J. et al. vegan: Community Ecology Package. R package version 2.9 (2013).
Fonseca, G. & Vieira, D. C. Overcoming the challenges of data integration in ecosystem studies with machine learning workflows: an example from the Santos project. Ocean Coast. Res. 71, e23021 (2023).
Vieira, D. C. et al. iMESc - an interactive machine learning app for environmental sciences. Front. Environ. Sci. 13, 1533292 (2025).
Kohonen, T. The self-organizing map. Proc. IEEE 78, 1464–1480 (1990).
Vesanto, J. & Alhoniemi, E. Clustering of the self-organizing map. IEEE Trans. Neural Netw. 11, 586–600 (2000).
Debeljak, P. et al. Seasonal patterns in microbial carbon and iron transporter expression in the Southern Ocean. Microbiome 11, 187 (2023).
Wu, Z. et al. Metagenomic insights into microbial adaptation to the salinity gradient of a typical short residence-time estuary. Microbiome 12, 115 (2024).
Li, D. et al. MEGAHIT v1. 0: a fast and scalable metagenome assembler driven by advanced methodologies and community practices. Methods 102, 3–11 (2016).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Eren, A. M. et al. Anvi’o: an advanced analysis and visualization platform for ’omics data. PeerJ 3, e1319 (2015).
Hyatt, D. et al. Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinformatics 11, 119 (2010).
Finn, R. D., Clements, J. & Eddy, S. R. HMMER web server: interactive sequence similarity searching. Nucleic Acids Res. 39, W29–W37 (2011).
Galperin, M. Y., Makarova, K. S., Wolf, Y. I. & Koonin, E. V. Expanded microbial genome coverage and improved protein family annotation in the COG database. Nucleic Acids Res. 43, D261–D269 (2015).
Alneberg, J. et al. CONCOCT: clustering contigs on coverage and composition. arXiv preprint https://abs/arXiv.org/1312.4038 (2013).
Eren, A. M. et al. Community-led, integrated, reproducible multi-omics with anvi’o. Nat. Microbiol. 6, 3–6 (2021).
Parks, D. H. et al. CheckM: assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes. Genome Res. 25, 1043–1055 (2015).
Chaumeil, P.-A., Mussig, A. J., Hugenholtz, P. & Parks, D. H. GTDB-Tk: a toolkit to classify genomes with the Genome Taxonomy Database. Bioinformatics 36, 1925–1927 (2020).
Shaffer, M. et al. DRAM for distilling microbial metabolism to automate the curation of microbiome function. Nucleic Acids Res. 48, 8883–8900 (2020).
Yu, X. et al. SCycDB: a curated functional gene database for metagenomic profiling of sulphur cycling pathways. Mol. Ecol. Resour. 21, 924–940 (2021).
Acknowledgements
We are grateful to PETROBRAS for the PCR-BS project planning and management. We would like to thank Daniel Moreira for coordinating the PCR-BS project and Dr. Frederico Pereira Brandini for coordinating the “Pelagic realm project”. We would like to thank members of the LECOM team for their scientific support on board. We are deeply grateful for the invaluable guidance and support provided by Rosa Carvalho Gamba, your legacy wisdom continues to inspire us even in her absence. A special thanks to the following PCR-BS project’s researchers and their institutions: UFRJ, UFF, PUC-Rio, UERJ, FIRJAN/SENAI, SALT, INPE, USP, UNIFESP, UFPR, UNESP, IP-SP, FURG and Socioambiental. To the University of São Paulo Foundation (FUSP) for financial administrative management. We also thank the RV Ocean Stalwart and RV Seward Johnson crew, and OceanPact for the scientific expeditions. We thank Barbara Resende Silva for providing valuable support in designing the conceptual diagram. We thank Alexandra Lehmkuhl Gerber and Ana Paula C. Guimarães from UGCDFA/LNCC for sequencing the samples. We acknowledge the anonymous reviewers’ valuable comments that helped to improve this manuscript.
Funding
This study was funded by Petróleo Brasileiro S.A. (PETROBRAS), through the RD&I investments clauses of Brazilian National Agency of Petroleum, Natural Gas, and Biofuels (ANP), under grant numbers 5850.0109317.18.9 and 21167–2, respectively. The project “Caracterização química e biológica do sistema pelágico da Bacia de Santos/PCR-BS” is a national collaboration between PETROBRAS and Universidade de São Paulo. NMB and LNL was financed by a Pos Postdoctoral ‘s fellowship from the PETROBRAS. ATRV is supported by FAPERJ (E-26/201.046/2022) and CNPq (307145/2021–2).
Author information
Authors and Affiliations
Contributions
N.M.B., V.H.P., D.L.M. and C.R.J. conceived and designed the study. J.C.F.M., A.M.A., A.M.E., R.G.R., D.C.D.C., F.S.P. and W.S.G.B. was responsible for onboard sampling and extracting DNA from water samples. M.G.C and F.P.B. was responsible for onboard sampling and generating environmental dataset. A.T.R.V. was responsible for generating the sequencing dataset. N.M.B., L.N.L., F.V.P., R.G.M.L., D.C.V. and F.M. carried out the bioinformatics and statistical analysis. N.M.B., R.G.R., and F.V.P., performed analysis of genomic data. D.C.V. and N.M.B. performed oceanographic analyses; N.M.B., F.V.P., D.C.V. and F.M. prepared the figures and tables and wrote the first draft of the manuscript. A.G.B. and G.F. provided helpful discussions and technical help. All authors discussed the results, commented, and reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Consent for publication
All authors have read and approved the paper for submission.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Bergo, N.M., Peres, F.V., Vieira, D.C. et al. Microbial signatures define the ecosystem functions of the pelagic microbiome in a basin-scale, Southwest Atlantic Ocean. Sci Rep (2026). https://doi.org/10.1038/s41598-026-37419-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-026-37419-9