Abstract
Erysipelothrix rhusiopathiae is a pathogenic bacterium infecting both humans and animals and causing erysipeloid, a disease that can leads to osteoarticular conditions, abscesses, and septic arthritis. Here, we report the sequenced ancient genome of E. rhusiopathiae obtained from teeth of an Alan child who lived in the North Caucasus in 6th–7th centuries AD. Differential diagnosis using paleopathological methods suggested that the child suffered from tuberculosis. However, analysis of tooth pulp revealed no signs of M. tuberculosis but a massive presence of ancient E. rhusiopathiae DNA. A complete high-quality ancient E. rhusiopathiae genome was assembled and found to belong to clade II of the extant E. rhusiopathiae phylogeny tree, nested in a subclade of closely related modern isolates infecting wild boars and domestic pigs. The ancient genome belongs to serotype 5 and encodes the key E. rhusiopathiae virulence factor SpaA group 1 along with vancomycin resistance genes. We propose that infection with E. rhusiopathiae, alone or together with other pathogens, is the likely cause of skeletal pathologies observed.
Similar content being viewed by others
Introduction
Advances in ancient DNA (aDNA) recovery and high-throughput sequencing have made it possible to study not only the genomes of ancient host organisms but also their associated pathogens. For example, several complete ancient genomes of Yersinia pestis, Brucella melitensis, and Mycobacterium tuberculosis have been assembled, providing information on the epidemiology of the past as well as insights into microbial evolution1,2. However, for most modern bacterial pathogens our knowledge of ancient genomes is scarce or totally lacking. In the present work, we report an ancient genome of Erysipelothrix rhusiopathiae, a pathogen serendipitously discovered in the teeth of 1400-year-old human remains from the North Caucasus (Kabardino-Balkaria, Russian Federation) belonging to early medieval Alanian culture.
E. rhusiopathiae is a Gram-positive, non-sporulating, non-acid-fast rod-shaped bacterium belonging to the genus Erysipelothrix of the family Erysipelotrichaceae3. It causes erysipeloid, a zoonotic disease affecting a wide range of animals including terrestrial and aquatic mammals, birds, fish, and some invertebrates3,4,5. Transmission to humans occurs through direct contact with infected animals, their waste, and contaminated animal products derived from an infected animal5. In humans, three main clinical forms of erysipeloid are distinguished: cutaneous, generalized, and septic6. In recent years, new forms of erysipeloid have been described: osteoarticular conditions7, abscesses8, and septic arthritis9,10, including several case reports in children11,12. However, many cases of this infection stay undiagnosed due to similarities with other diseases and insufficient laboratory diagnostics13,14. The pathogenesis is also poorly understood6 although it is assumed that immunosuppressive conditions are an important factor in systemic infection in humans15.
The first complete genome of E. rhusiopathiae was sequenced in 201116. It consists of a single circular chromosome of approximately 1.8 million bp with an average GC content of 36.5%16,17,18. The E. rhusiopathiae genome lacks many biosynthetic pathways, reflecting reductive genome evolution typical of many pathogenic bacteria16,19. The main virulence factors of E. rhusiopathiae comprise neuraminidase, hemolysin, capsular polysaccharides, and surface adhesive proteins14,15,16,17,20. The key proteins responsible for pathogenesis are surface protective antigens SpaA or SpaB16. The SpaA protein mediates resistance to phagocytosis and is required for virulence16,21. The C-terminal region of SpaA includes GW-repeats and is responsible for adhesion to the host cells. It has been experimentally shown that the higher number of tandem repeats is positively correlated with virulence22. Also, the E. rhusiopathiae genome encodes several determinants of drug resistance. Intrinsic resistance to vancomycin is a characteristic feature of the species15,23.
An important characteristic for understanding strain epidemiology is its serotype – a unique complex of surface proteins. More than twenty serotypes have been described for E. rhusiopathiae24. We also employ Multilocus Sequence Typing (MLST), which determines the combination of alleles in bacterium’s core genes. This method is widely used to follow molecular epidemiology of pathogens25.
The population structure of E. rhusiopathiae includes three main clades: I, II, and III26,27. No strict geographic or host-specific associations of these clades have been identified so far27,28,29,30. However, clade II predominates in the Old World, while clade III – in the New World26,27,29,30. SpaA is common in clades II and III members26. The SpaB surface antigen protein is found exclusively in clade I, which is phylogenetically distant from clades II and III.
Frequent recombination involving more than half of core genes was shown to increase the genomic diversity of E. rhusiopathiae26, with the rate of recombination varying significantly between the clades26,30. The core genes are highly conserved, in contrast to the rest of the genome19,26.
To date, no complete paleogenomes of E. rhusiopathiae have been reported, although several genome fragments were identified in remains of human hunter-gatherers from Eastern Patagonia31 and of Chalcolithic horses from Mongolia32. In addition to genome reconstruction, we aimed to establish the key epidemiological characteristics of the ancient pathogen, namely, its key virulence factors, the Spa- and serotypes, and determinants of drug resistance. Phylogenetic relationships with modern strains were also investigated.
Materials and methods
Description of the archaeological site and biological profile of the individuals
The Zayukovo-3 burial ground is located near the village of Zayukovo in the Kabardino-Balkarian Republic, Russia (geographic coordinates 43° 36′ 06.1ʺ N 43° 12′ 57.1ʺ E). It is situated on a high rocky promontory at the confluence of the Baksan and Gundelen rivers (Fig. 1a). The multi-layered cemetery contains remains associated with three archaeological cultures: the Western Koban culture (8–4th centuries BC), a group of sites of the Podkumok–Khumara type (1–3rd centuries AD) and the Alan culture (5–8th centuries AD)33. The bearers of these cultures were engaged in agriculture, animal husbandry (primary cattle and small ruminants), and various handicrafts.
Archeological site, the Sk213 specimen, and material used for DNA extraction. (a) The geographical location of the archeological site is indicated by a red point. The grey area shows the territorial boundaries of the Alanian culture in the Northern Caucasus during the 5th – eighth centuries AD34. Figure 1a was generated using Cartopy v0.23.0 (https://scitools.org.uk/cartopy) with data from Natural Earth public domain datasets (https://www.naturalearthdata.com/). (b) Photo of the burial 213; the multifaceted bronze earring found in the burial site is shown at the down. Photo credit: A. Kadieva & S. Demidenko.
The excavations relevant to this study were conducted in 2021 by the Joint North Caucasian Expedition of the State Historical Museum, the Kabardino-Balkar Scientific Center RAS (Nalchik), and the Institute of Archaeology RAS (Moscow), under the leadership of A.A. Kadieva (License No. 1978–2021). All archaeological work was carried out in strict compliance with local legislation.
To establish the sex and age of the individuals an anthropological analysis was carried out. Sex and age at death was estimated by traditional methods of paleoanthropology35,36,37,38,39,40,41,42,43,44. Paleopathological manifestations were studied using high-resolution video microscopy HIROX RH-2000 (Hirox, Japan).
Radiocarbon dating
The accelerator mass spectrometry (AMS) radiocarbon dating was used to determine the age of the samples at the “AMS Golden Valley”, Novosibirsk State University, Budker Institute of Nuclear Physics of SB RAS (Novosibirsk, Russia. AMS Lab ID = GV). AMS was performed on a MICADAS-28 instrument with a δ13CVPDB value of − 25.7‰. Calibration of the radiocarbon age to calendar years was based on the IntCal20 calibration curve using the OxCal 4.4 software45 with atmospheric data from Reimer et al.46. The samples were assigned the code GV-4860 (remains Sk213) and GV-4861 (remains Sk258). For protocol details see Supplementary Methods.
Sample preparation and aDNA extraction
All the work was conducted in an ISO-9 positive-pressure rooms in the LLC “Biotech Campus” (Moscow, Russia) using appropriate measures to prevent contamination with modern DNA.
Prior to sampling the teeth were decontaminated following the guidelines of Keller et al.47. The pulp was prepared according to Neumann et al.48. Powder mass of the pulp was: first tooth of Sk213_42 (tooth 42, ID1) – 19 mg, second tooth of Sk213_12 (tooth 12, ID2) – 13 mg, Sk258_43 tooth (tooth 43, ID3) – 18 mg, and Sk226_13 tooth (tooth 13, ID53) – 29 mg. Ancient DNA was extracted using magnetic beads following Rohland et al.49 and Clavel et al.50 recommendations. The tooth powder was dissolved in the Lysis buffer for 1 h at 37 °C, then added to the Binding buffer D together with the magnetic beads. The beads were washed three times in the Protein Elution buffer. After washing, the beads were dried and the DNA was eluted with the Elution buffer. Detailed protocols and buffers compositions are presented in Supplementary Methods.
Libraries construction and sequencing
Libraries were prepared from 15 µl of eluted DNA solution using the MGIEasy PCR-Free DNA Library Prep Set (MGI, China) with modifications. Adapters were pre-diluted fivefold prior to ligation, and post-ligation cleanup was performed with DNA Clean Beads at a 1.8X ratio to preserve short fragments. The eluted adapter-ligated libraries were amplified for 12 cycles using Q5 High-Fidelity DNA Polymerase (NEB, USA) and MGI-specific primers. The amplified libraries were purified, and their size distribution and concentration were assessed using an Agilent Tapestation System 4200 (Agilent, USA). No DNA contamination was detected in the negative controls. Subsequent library preparation and sequencing were conducted using the DNBSEQ-T7RS High-throughput Sequencing Set (MGI, China). To mitigate potential nucleotide distribution bias from overlapping adapter sequences, a custom sequencing recipe with reduced (to 60) cycle count was employed for paired-end 60-base reads (PE60) on the DNBSEQ platform. Detailed protocols are provided in the Supplementary Methods.
Primary processing and authentication of reads
Pair-end DNA reads were processed by fastp v0.23.251 for adapter removal (-length required 30). The metagenome was analysed with Kraken2 v2.1.3.52 using a dataset that included whole-genome nucleotide sequences of bacteria, archaea, protists and viruses from the NCBI RefSeq database (2023-08-12). To identify soil bacteria, we performed a cross-referencing check against a genomic catalogue of soil microbiomes53. Reads from both libraries prepared from Sk213 and assigned to E. rhusiopathiae were jointly used for further analysis. Mapping to the reference was performed using bowtie2 v2.5.1 (-n 1)54. The genome of E. rusiopathiae GCF_900637845.1 (strain NCTC8163, United Kingdom, 1950) was used as a reference sequence. This genome was chosen as the reference because, in contrast to the Fujisawa reference strain of the species that was isolated in Japan and represents an intermediate clade, it is representative of European strains and belongs to clade II of which the ancient genome is also a member. Duplicates were deleted by MarkDuplicates v4.0.11 (Picard)55. The resulting alignment was used to analyze DNA damage using mapDamage v2.0.1.56 and PyDamage v0.7057. Accordingly, the genome was identified as aDNA according to strict q values (= 0.011), with an accuracy of 1.0 for the test calculated by PyDamage v0.7057. We assigned “ERA_01” ID to the ancient genome.
Host genetics
Paired-end DNA reads were processed with fastp v0.23.251 for adapter removal and quality filtering (-length_required 30). The filtered reads were aligned to the human reference genome GRCh38/hg38 using bowtie2 v2.5.1 with the parameter -n 154. Duplicate reads were removed with MarkDuplicates v4.0.11 (Picard)55. Alignment quality metrics were assessed using Qualimap v.2.2.258 (bamqc module).
The mitochondrial haplogroup was determined with haplocheck v1.3.259, the Y-chromosomal haplogroup – using YLeaf v3.060 and Y-LineageTracker v1.3.061. Mitochondrial contamination was estimated with Schmutzi62 using the authors’ recommended pipeline for contamination assessment. Genetic analysis confirmed the female sex of Sk213 individual and identified her mitochondrial haplogroup as J1c + 16,261.
Screening for M. tuberculosis complex (MTC) and mycobacteria other than tuberculosis (MOTT)
The filtered paired-end DNA reads were aligned to the MTC-specific genomic island using bowtie2 v2.5.1 with a maximum of 1 allowed mismatch54. The MTC-specific genomic island (NCBI: NC_000962.3. Region: 3,119,185…3,123,576) contains the IS6110 mobile genetic element (NCBI: NC_000962.3. Region: 3,119,185…3,123,576), which is a specific marker of MTC but also other pathogenic MOTT63.
A dataset of assembled Erysipelothrix sequences
A dataset containing 506 genome sequences of representatives of the genus Erysipelothrix included 501 – E. rhusiopathiae isolates, 2 isolates of E. tonsillarum, and single isolates of E. piscisicarius, E. sp. 715, and E. sp. Pecs 56, for which the host, year and place of isolation were known. Various combinations of these sequences were previously used in phylogenetic studies: Dec et al.64, Forde et al.26,27,28, Groeschel et al.23, Huang et al.65, Ogawa et al.16, Shimoji et al.24, Soderlund et al.29, Webster et al.30 and Zautner et al.15. Data on some strains have been verified according to Kucsera66. Metadata including Sequence Read Archive (SRA) ID’s and other information is presented in Supplementary Table 1. SRA data was assembled de novo using SPAdes v3.13.0 genome assembler with default parameters67.
Phylogenetic analysis and temporal structure
The genomic dataset (506 assembled genomes, Supplementary Table 1) was processed with snippy-multi from the Snippy package v4.6.068 to identify SNPs with using E. rhusiopathiae GCF_900637845.1 (NCTC8163) as the reference genome. The resultant SNPs were filtered by snippy-core to obtain a genome-wide core alignment. Using the alignment, a maximum likelihood phylogeny was inferred with IQ-TREE v1.6.11 using the GTR + F + I + G4 substitution model and 1,000 rapid bootstrap inferences69.
The presence of a temporal signal in heterochronous sequences was checked using the TempEst v1.5.370.
Genotyping of spaA
The nucleotide sequence of the spaA gene was extracted from the FASTQ reads using the sequence of strain IMT23643 (NCBI: KR606239.1), which has the largest number of C-terminal repeats25 as a reference using SPAdes v3.13.0 genome assembler with default parameters67. Translation of the nucleotide sequence into an amino acid sequence was carried out with Expasy71 using standard genetic code. The nucleotide and amino acid sequences were aligned with MUSCLE72 integrated into Ugene v49.173. SpaA typing was performed according to the scheme proposed by Forde et al.27 for five variants at amino acid positions 55 (V/I), 70 (K/N), 178 (G/D), 195 (D/N), and 303 (G/E) relative to the reference sequence from the Fujisawa strain (NCBI: AP012027.1 Region: 112,931…114,811). The unique single nucleotide substitutions of the hypervariable region were also evaluated relative to the reference sequence of the NCTC8163 strain (NCBI: NZ_LR134439.1 Region: 793,538…795,418).
In silico serotyping and serotype-determination region reconstruction
In silico serotyping was performed using Ugene V49.173, by a BLAST74 search of all primers sets with 90% similarity for positive site identification, the PCR-based serotyping schemes by Shiraiwa et al.75 et Shimoji et al.24 were used. To address the Forde et al.27 comment about one nucleotide discrepancy in reverse primer specific for serotype 5, GAAATAATGCCAATAGATGGAGCACC primer was used. The reads were aligned to the chromosomal region defining the serotype of strain Pecs 67 genome (NCBI: LC380407.1) with bowtie2 v2.5.1 (−n 1)54.
Multi locus sequence typing
Multi Locus Sequence Typing (MLST) for ERA_01 was performed according to Janssen et al.25 and Webster et al.30 schemes.
Genomes reconstruction and functional analysis
Trimmed FASTQ reads were first mapped to the E. rhusiopathiae GCF_900637845.1 (NCTC8163) reference genome to extract organism-specific reads from the ancient DNA sample. Thus filtered reads were then assembled de novo using SPAdes v3.13.0 genome assembler with default parameters67.
Eggnog v2.1.1276 was used to identify clusters of orthologous genes. Prokka v1.14.677 and roary v3.11.278 were used for functional annotation. In addition, loci described in Zautner et al.15 as contributing to intrinsic or putative antibiotic resistance in E. rhusiopathiae were verified by targeted mapping. These included vex2 (ABC transporter, ATP-binding protein), vex3 (ABC transporter membrane-spanning permease), three MATE-family efflux transporters, and several β-lactamase-related genes (class C-like β-lactamase, penicillin-binding protein [PBP] superfamily, MBL-fold metallo-hydrolase, and Zn-dependent β-lactamase-like hydrolase). The same loci were also were also found in the ERA_01 assembly by a BLAST search. Completeness and contamination of the final assembly were identified by CheckM v1.2.279. The list of virulence factors was taken from Zautner et al.15.
To check the assembly quality, the raw reads were mapped to the assembled (SPAdes v3.13.067), annotated (Prokka v1.14.677) genome of the nearest neighbour – ERR3933026 (18ALD662, Sweden, 2018). Pysam v0.19.180 was used to identify genes with a coverage completeness of more than 80%. This number corresponds to the results obtained by the roary v3.11.278. The RAST server81 was used to predict coding DNA sequences (CDS).
Visualization
Proksee82 was used to visualize genome annotation. The aDNA-BAMPlotter v2.0.1 was used to visualize the completeness of coverage breadth83.
Phylogenetic reconstructions were visualized and managed using the iTOL web server84. Heatmaps were constructed using custom python scripts (https://github.com/melibrun/ancient-genome-of-Erysipelothrix-rhusiopathiae-). The Venn diagram and pie chart were created using the matplotlib library v3.7.1 (python). The serotype-determination region was visualized using pyGenomeViz v0.4.485.
Results
Pathological manifestations of an early medieval Alanian skeleton
This study examined six medieval individuals from the Zayukovo-3 site. Burials 226 and 258 contained 3 and 2 persons, respectively; burial 213 was a single grave. According to the archeological data, burial 226 dates back to the 5–6th century AD it contained remains of two children (9–18 months and 9–12 years old) and an adult male of 40–45 years. Burial 258 dates back to the 5–7th century AD and contained remains of two young (18–20 years and 16–20 years old) individuals. Burial 213 contained the remains of a 10–11 year old child.
Here, we studied an early medieval Alanic skeleton of a 10–11 year old child from burial 213 (Sk213) (Fig. 1b). Genetic analysis confirmed the female sex of the individual and identified her mitochondrial haplogroup as J1c + 16,261.
The grave goods consisted solely of a single earring, which is of the polyhedral type (Fig. 1b) common in the North Caucasus between 5 and 7th centuries AD86. The diameter of the earring is 2.5 cm, the dimensions of the 14-sided extension at the end are 0.6 × 0.6 cm.
AMS radiocarbon dating of Sk213 (AMS lab ID: GV-4860) indicated an age of 1415 ± 40 years before present (corresponding to 574–668 AD), Supplementary Fig. 1. Morphological examination revealed multiple pathological changes in the thoracic region of the spine and ribs. On the cervical (hereinafter C) vertebrae (C2–C7), the upper thoracic (hereinafter T) vertebrae (T1, T2), and the upper lumbar (hereinafter L) vertebrae, enlarged nutrient openings are observed on the anterior surface of vertebral bodies. The thoracic region is not fully represented with thoracic vertebrae T3, T11, and T12 missing. Multiple lytic lesions are noted on the anterior surface of T4–T5 vertebral bodies. The anterior surface of both vertebral bodies has a coarsely cellular disorganized structure; the size of these lytic lesions varies from 2 to 3 to 10 mm (Fig. 2a). On the anterior surface of T6, periostitis consistent with an inflammatory process is noted. On the anterior part of the vertebral body of T7 there are no pathological alterations, the surface shows normal variability. On the lateral surfaces of T4–T7, there are traces of an inflammatory process of the periosteum; a similar process is detected on the posterior part of vertebral bodies around the vascular hole. The anterior and inferior body parts of T8 and the body of T9 are completely absent due to a destructive lytic process. Only the upper endplate is preserved in the body of T10 (Fig. 2b). The preserved parts of the vertebra have a shape of a broad wedge, covered with bone and strands and small lytic cavities. On the preserved lateral sides of vertebral bodies T8–10, small-cell periostitis is visible. Ankylosis of the inferior articular processes of T9 and the superior articular processes of T10 is noted.
Pathological changes in Sk213 vertebrae and ribs. (a) 1 – anterior, 2 – posterior, 3 – right side, and 4 – left side views of T4–T7 vertebrae. The black arrows show lytic lesions of vertebral bodies. (b) 1 – left side, 2 – posterior, 3 – anterior views of T8 – T9 vertebrae; 4 – upper, 5 – lower surfaces of T10. (c) High resolution microscopy of 6th right-side rib; periostitis located on the internal surface in close proximity to the angulus costae (vertebrae end) is visible.
The Sk213 ribs were also involved in the pathological process (Fig. 2c). On the inner surface of the heads and necks of right ribs, from the 5th to the 10th, there is a fine-meshed layered periostitis. Some rib heads are visibly enlarged. The highest degree of pathology is found in the 6th and 7th right ribs. On the left side, the inflammatory processes reached the bodies of ribs 3–5; in addition, there is periostitis on the heads of ribs 4 and 6–8. The periostitis is dense, uniform, well-defined, and similar in degree of formation both on the heads and on the bodies of the ribs. Thus, the greatest degree of periostitis development is observed in the middle of the thoracic region, corresponding to extensive pathology of vertebrae T4–T10. Signs of a possible pathological process were detected in the pronounced woven surface of the posterior part of the sternum.
Metagenomic analysis of ancient DNA from Sk213 teeth reveals abundant E. rhusiopathiae sequences
Spinal pathologies of Sk213 are consistent with the M. tuberculosis infection. To provide support for this notion, we sequenced metagenomic DNA isolated from the remains of tooth pulp taken from a well-preserved Sk213 tooth. As a control, libraries prepared from tooth pulp from remains of individuals Sk258_43 (ID3) and Sk226_13 (ID53) from nearby burial sites but without the signs of severe skeletal pathology were sequenced. Taxonomic classification of reads was carried out using Kraken2 v2.1.352 with the NCBI RefSeq database (version from 2023-08-12). In libraries prepared from Sk258_43 and Sk226_13, only reads matching soil bacteria were detected (Supplementary Table 2). In the library prepared from Sk213_42 (ID1), 226,506 reads (0.11% of the total) matched the Erysipelothrix rhusiopathiae genome. Reads matching E. rhusiopathiae were the most abundant, followed by reads corresponding to environmental Enhydrobacter sp. (0.08%) (Supplementary Table 2). The pyDamage57, mapDamage56, and aDNA-BAMplotter83 analysis revealed strong deamination patterns at the ends of E. rhusiopathiae reads, a characteristic of ancient DNA (Fig. 3a).
Authentication and phylogeny of the ancient E. rhusiopathiae genome. (a) Genome-wide coverage and alignment quality metrics are shown for the reference (GCF_900637845). The x-axis in the coverage plot represents nucleotide positions along the reference, and the y-axis shows read depth. Coverage from all mapped reads is shown in grey, while reads with mapping quality ≥ 30 are shown in green. Genomic regions where reads have mapping quality equal to zero are highlighted as red points, indicating low-confidence alignments. The edit distance distribution is shown as a bar plot, where the x-axis represents the number of mismatches and small indels per read, and the left y-axis indicates the fraction of reads at each edit distance. Grey bars correspond to all mapped reads, while green bars indicate reads with mapping quality ≥ 30. The right y-axis demonstrates the frequencies of nucleotide misincorporations at the 5′ and 3′ ends of reads, based on mapDamage256 output. Specifically, it indicates the frequency of C → T (at 5′ ends) and G → A (at 3′ ends) substitutions, which are characteristic of ancient DNA damage. (b) A dendrogram representing the ML tree reconstructed for modern and the ancient E. rhusiopathiae genomes. E. tonsillarum was used as an outgroup. The reference genome GCF_900637845 is marked with a blue asterisk. On the outer circle, strains belonging to Spa types are indicated (SpaA – blue, SpaB – violet). The inner circle shows isolation sources (hosts); the circle in the middle shows a country of isolation in accordance with ISO 3166–1 alpha-2 codes. A fragment of the dendrogram containing the ancient genome (marked with an arrow) is zoomed in the inset. Modern genomes used to reconstruct local phylogeny of the ancient genome are marked with yellow asterisks. Bootstrap values are indicated.
To further confirm the systemic presence of E. rhusiopathiae in Sk213_12, a second library (ID2) was prepared from another tooth of this individual and sequenced. Consistent with results obtained with the first tooth, 153,600 reads (0.08% of the total) matched E. rhusiopathiae sequences and showed the deamination pattern typical of ancient DNA. No significant quantities of a sequences corresponding to M. tuberculosis we found in either library. Since Kraken2 read classification is prone to high levels of false positives87, we conducted additional screening for pathogenic MTC and MOTT specific genetic marker – the mobile element IS6110. The search returned no results.
The ancient genome belongs to E. rhusiopathiae clade II
Alignment of ancient E. rhusiopathiae reads to the GCF_900637845.1 reference genome resulted in an average coverage depth of 21x. Full alignment statistics is presented in the Supplementary Table 3. Several regions of the reference genome are not covered by aDNA reads and are thus either absent or highly diverged in the ancient genome.
To reconstruct the phylogeny of the ancient genome, we used data from 501 modern E. rhusiopathiae strains available in the Sequence Read Archive (SRA) as of 2023-09-25. The SRA contains unassembled raw sequencing data and its subsets were previously used to study E. rhusiopathiae phylogeny16,23,24,26,27,28,29,30,65. To identify the core set of E. rhusiopathiae SNPs, reads from all strains, including the ancient one, were mapped to the GCF_900637845.1 reference. In total, 4,687 SNPs were identified and used to build a Maximum Likelihood tree (Supplementary Table 4). Mapping of the ID1 and ID2 reads resulted in the same set of SNPs, showing that the same strain was present in both Sk213 teeth and thus validating the joint analysis of both libraries. The ancient genome is located inside E. rhusiopathiae clade II (Fig. 3b) currently dominant in the Old World26,27,29,30. Thus, strains from this clade were already present in Europe in the early Middle Ages. On a finer scale, the ancient genome is clustered with 18 modern isolates of which 17 were isolated from pigs and wild boars in Europe (predominantly from Sweden, Italy, and the UK) and one – from caribou in the Canadian Arctic (Fig. 3b, Supplement Table 1, List 2). Ancient E. rhusiopathiae is phylogenetically closest to the modern genome ERR3933026, strain 18ALD662 isolated from a wild boar in Sweden (Fig. 3b).
Reconstruction, comparative analysis, and functional annotation of the ancient E. rhusiopathiae genome
To reconstruct the ancient genome, we assembled the genome of the closest modern isolate ERR3933026 from reads deposited in SRA, and used the resulting assembly to map the ancient E. rhusiopathiae reads. Next, the mapped reads were re-assembled with SPAdes67. The reconstructed ancient genome, named ERA_01, consists of 259 contigs with N50 of 32 kbp and a total length of 1,713,412 bp, which is comparable to the sizes of modern E. rhusiopathiae genomes (Supplementary Fig. 2). The genome has completeness of 100.00% and contamination of 2.65% by CheckM79, which corresponds to a high-quality draft genome by MIMAG criteria88 and allows further reliable comparative analysis with modern genomes.
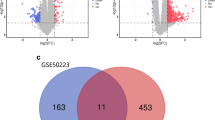
The ERA_01 genome was next compared to the ERR3933026 genome and the GCF_900637845.1 reference. ERA_01 contained 1,561 coding DNA sequences (CDS) predicted with the RAST server81. 1,520 genes are shared by all three strains (Fig. 4a).
Comparative analysis of the ancient E. rhusiopathiae genome ERA_01. (a) Numbers of genes shared between ERA_01, ERR3933026, and GCF_900637845.1 genomes. (b) Comparison of structural variations in the ERA_01, GCF_900637845.1, and ERR3933026 genomes.
97 ERR3933026 genes are unique (Fig. 4a. Supplementary Table 5). 43 of these genes are part of a 32.7 kbp prophage (ERR3933026 genome positions: 378,815…412,373, Fig. 4b) that is sporadically present in E. rhusiopathiae isolates (e.g., in strain Fujisawa). 47 unique ERR3933026 genes are organized in two clusters. Cluster 1 (genome positions: 8,390…72,580, Fig. 4b) contains a terminase and transposase genes and likely defines a mobile genetic element. Cluster 2 (genome positions: 847,640…865,687, Fig. 4b) is a three-gene ara phosphotransferase (PTS) system. 7 standalone unique ERR3933026 genes code for hypothetical proteins/domains of unknown functions.
Of 34 genes unique to GCF_900637845.1, 8 (genome positions: 304,293…339,739, Fig. 4b) are organized in a putative virulence cluster, which encodes hypothetical proteins, internalin, and an ig-like domain-containing protein. 4 genes are serotype-specific (note that GCF_900637845.1 and ERA_01 are of different serotypes, see below). 6 genes are organised into a cluster (genome positions: 535,023…539,880, Fig. 4b) containing a gene that codes for an AIPR phage defense protein. A closely adjacent unique 8-gene cluster (genome positions: 539,982…546,592, Fig. 4b) contains a gene coding for a GNAT family N-acetyltransferase gene. 11 genes are grouped into a radical SAM protein cluster (genome positions: 1,369,602…1,390,305, Fig. 4b). The remaining 5 unique GCF_900637845.1 genes are standalone and code for an LPXTG cell wall anchor domain-containing protein, an AAA family ATPase, a FtsX-like permease family protein, and two hypothetical proteins.
Loci previously described as potential determinants of antibiotic resistance in E. rhusiopathiae15 – including vex2 (ABC transporter, ATP-binding protein), vex3 (ABC transporter membrane-spanning permease), three MATE-family efflux transporters, and several β-lactamase-related genes (class C-like β-lactamase, penicillin-binding protein [PBP] superfamily, MBL-fold metallo-hydrolase, and Zn-dependent β-lactamase-like hydrolase) – were identified by targeted mapping. All loci showed complete coverage, confirming their presence in the ERA_01 genome (Supplementary Table 6). Genetic determinants associated with resistance to tetracycline and fluoroquinolones found in some modern E. rhusiopathiae genomes are absent from the ancient genome.
Of the 83 E. rhusiopathiae genes associated with virulence15, 78 are present in the ancient genome (Supplementary Table 7). Since the five genes missing from ERA_01 encode supplementary virulence factors with no clear functional role, and absence in several modern virulent strains (Table 1, Supplementary Table 7), the ancient microbe was likely virulent.
The ancient E. rhusiopathiae genome contains the spaA gene, belongs to serotype 5, and defines a novel MLST group
The key E. rhusiopathiae virulence factor is SpaA, a protein responsible for adhesion to host cells and protection of bacteria from phagocytosis. We used discriminatory amino acid positions in the hypervariable immunoprotective domain of SpaA (amino acid positions 30–413) to classify the ancient SpaA27 (Fig. 5a). The ERA_01 spaA has substitutions in nucleotide positions 163, 210, 533, 583, and 908, which lead to non-synonymous amino acid substitutions that position the ancient SpaA in group 1 according to the Forde classification scheme. The same substitutions are found in the ERA_01 closest relative ERR3933026 and in the GCF_900637845.1 reference (Fig. 5b, Supplementary Fig. 4). Compared to the GCF_900637845.1 reference, two synonymous single nucleotide substitutions C- > T and T- > C are found, respectively, at positions 712 and 861 of the ancient spaA (Supplementary Fig. 4). Only one of these, at position 712, is present in spaA from ERR3933026.
The spaA genotyping and serotype identification of ERA_01. (a) A schematic representation of the SpaA protein according to Forde et al.27 with discriminatory amino acid positions and amino acid changes they introduce indicated. GW repeats (at a typical number of 8) are shown, additionally, the C-terminal GS motif containing domain is indicated. (b) Results of spaA genotyping of ERA_01 and its close modern relatives using the Forde et al.27 scheme. (c) Serotype-determining region of the ERA_01 genome. The scheme shows the structure of a serotype-determining gene cluster split into three contigs. Contig names and predicted gene products are labeled. Locations of synonymous and non-synonymous SNPs (with amino acid substitutions for the latter labeled) are shown with, respectively, green and red vertical lines.
The C-terminal region of SpaA contains a number of GW tandem repeats responsible for adhesion. The typical number of repeats in modern strains (e.g., 18ALD662 and members of its subclade, NCTC8163, and Fujisawa) is 8 (Fig. 5b, Table 2). Some group 1 SpaA proteins contain 11 GW repeats25. Interestingly, the ERA_01 SpaA has 13 repeats. The only known modern genome with 13 GW repeats, IMT23643 from a domestic pig in Germany, is incomplete and thus it is impossible to establish the degree of its relatedness to ERA_01.
According to in silico PCR with primers specific to gene clusters responsible for capsule biosynthesis and antigenicity24,75, ERA_01 belongs to serotype 5. We reconstructed the serotype-determining region of ERA_01, split between 3 contigs in our assembly, using the genome of Pecs 67 strain, which is a reference for serotype 5 strains. The alignment showed that the entire serotype-determining region from Pecs 67 was covered by ERA_01 reads without deletions or insertions. Compared to Pecs 67, six out of eleven serotype determining genes in ERA_01 carry SNPs, some of which are nonsynonymous (Fig. 5c, Supplementary Table 8). Modern isolates that cluster with ERA_01 into a subclade inside clade II (Fig. 3b) belong to serotypes 2 (13 genomes) and 5 (4 genomes, including the closest modern genome ERR3933026), with a single genome (ERR3932964 – strain 07BKT12931 Sweden, pig, 2007) belonging to serotype 21 (Table 2). While this variability indicates, on the one hand, high evolutionary plasticity of the serotype-determining region, the fact that the ERA_01 serotype is preserved in 5 of the 15 closest extant relatives also shows stability over hundreds of years.
Standard MLST analysis of E. rhusiopathiae is carried out using polymorphic loci in seven housekeeping genes, with different combinations of alleles forming multiple sequence types for individual strains. According to in silico MLST analysis of ERA_01, six of these genes (gpsA, purA, pta, prsA, galK, and ldhA) belong to allele 1, as in the reference GCF_900637845.1. However, the seventh gene, recA, has an additional synonymous G- > A SNP in position 912 that is unique and not found in any modern E. rhusiopathiae, thus defining a novel MLST type. The allelic profile of the ancient genome in the PubMLST format is as follows: “gpsA:1, recA: Novel, purA:1, pta:1, prsA:1, galK:1, ldhA:1. ST = Novel” (Supplementary Table 9).
Discussion
In this work, we reconstructed a ~ 1400 year old genome of E. rhusiopathiae, the first complete ancient genome for the species. The ERA_01 paleogenome belongs to E. rhusiopathiae clade II, which is predominantly distributed in the Old World26,27,29,30. Further, ERA_01 belongs to subclade containing 17 strains isolated between 1970 and 2018 from wild boars and farm pigs in Europe (Sweden, Italy, Great Britain, Denmark, Hungary) and one strain from caribou in the Canadian Arctic (2197i, SRR2085513) (Fig. 3b, Supplementary Table 1). Based on SNP polymorphisms, the ERR3933026 isolated from a wild boar in Sweden in 2018, is the closest modern relative of ERA_01. Thus, the phylogenetic lineage to which the ancient genome belongs exists at least since the seventh century AD. Circulation of strains belonging to ancient phylogenetic lineages in the modern times has been described for other zoonotic pathogens, for example, the causative agent of plague (Yersinia pestis)89, brucellosis (Brucella melitensis)90, and tuberculosis (Mycobacterium tuberculosis complex)91. Determination of whether the ERA_01 lineage bacteria were common in the past and tended to infect wild boars will require additional analysis of ancient E. rhusiopathiae genomes.
The diversity of serotypes within a phylogenetically close subclade to which ERA_01 belongs must be due to high frequency of horizontal gene transfer in E. rhusiopathiae27,28,29. The ancient genome belongs to serotype 5. The etiological agents of erysipeloid in farm animals are most often strains of serotypes 1 and 227.Serotype 5 is more common among wild animals, which infect farm animals upon contact with them75,92. For example, 28.5% strains isolated from wild animals had serotype 5, compared to just 6.3% for farm animals27. In an independent study of Soderlund et al. these numbers were 18.7% for wild boars compared to 5% in farm pigs29. While by no means definitive, these observations may be consistent with a scenario that Sk213 was infected with ERA_01 from a wild boar.
Though ERA_01 is phylogenetically closest to ERR3933026 based on SNP typing, it is actually more similar to GCF_900637845.1, a E. rhusiopathiae reference strain that falls into a different subclade of clade II, based on structural variations. Like GCF_900637845.1, ERA_01 lacks both prophages present in ERR3933026. It also lacks the virulence, GNAT, SAM clusters present in GCF_900637845.1. In addition, a three-gene cluster containing a CHAP-domain protein gene that is present in both modern strains is absent from ERA_01. While the ancient genome may have contained its own unique genes, they can not be identified because the genome was reconstructed by mapping on existing references.
Forde et al.26 failed to observe a temporal structure for the E. rhusiopathiae species. The absence of temporal structure in the species as a whole but its presence in some lineages was previously described for some pathogens, for example M. tuberculosis93 and Y. pestis94. We revisited the temporal structure of the E. rhusiopathiae species by analysing a set that included the ancient genome and all currently available E. rhusiopathiae genomes or just the lineage comprising the ancient genome and its closest relatives. The “root-to-tip” regression did not reveal a temporal signal (Supplementary Fig. 3). This is consistent with the finding of Forde et al26, who showed that the substitution rates in E. rhusiopathiae are poorly represented by a simple linear model, and implies that there is no strict molecular clock within the species as a whole.
Out of 83 confirmed and putative E. rhusiopathiae virulence factors belonging to five functional groups14,15,16,17,20, 78 were found in the ancient genome (Supplementary Table 7, List 2). Importantly, main clade II virulence factors, spaA, neuraminidase, hemolysin are encoded by ERA_01. The putative virulence factors missing from ERA_01 are patchily represented in modern pathogenic strains and thus their contribution to pathogenicity, if any, is minor. It is thus highly likely that ERA_01 was pathogenic and could have caused erysipeloid.
Differential diagnosis using paleopathological methods of spinal pathologies of Sk213 is consistent with tuberculosis95. However, the presence of MTC and MOTT could not be confirmed by molecular methods. Nevertheless, based on archeological case of tuberculosis described by Palfi et al.96, which is morphologically similar to our case, we cannot exclude the presence of infection. Bone lesions caused by bacteria of other genera of the Mycobacteriaceae family (e.g., Mycolicibacterium, Mycolicibacter, Mycolicibacillus, Mycobacteroides) are morphologically indistinguishable from lesions caused by M. tuberculosis. Therefore, the presence of infection by MOTT cannot be ruled out. In addition to more or less typical changes known for bone tuberculosis, some changes in Sk213 are induced by probable inflammation of the posterior ligaments of the spine and are not typical of tuberculosis.
Among the contemporary clinical cases in which the link between the presence of E. rhusiopathiae and the pathological process in the bone tissue has been confirmed, there are several cases involving the spine. Several studies reported pathological manifestations in the lumbar region, for example, an extensive disc disease as well as a destructive lesion of the left L3 pedicle caused by bacterial osteomyelitis97, psoas abscess and discitis together with L2-3 osteomyelitis8 and prevertebral abscess and L5-S1 osteomyelitis98,99. Clinical cases of the upper spine pathology, though less common are not exceptionally rare99. Thus, destructive changes in vertebral bodies, arches, and spinous processes in the thoracic segment (T5 – T6) caused by E. rhusiopathiae infection have been described. Wedge-shaped destruction of the vertebral arches, vertebral joints, and spinous processes were noted100.
All clinical cases described above have one important factor in common—immunosuppression associated with another background disease such as chronic arthritis, cerebral infection, osseous necrosis4, HIV, endocarditis with acute renal failure, septicaemia, lupus nephritis, pneumonia101, osteomyelitis97,102. Human E. rhusiopathiae infection is most commonly associated with the occupational exposure to animals or their contaminated products and represents a zoonotic disease103. In most cases the pathway of the bacterial infection is through damaged skin when handling an infected animal. This results in localised cutaneous cellulitis. However, there are some cases when the infection takes on more aggressive forms. E. rhusiopathiae is known to establish chronic foci of osteomyelitis due to its ability to evade host immune responses and persist in tissues104,105. Such foci serve as reservoirs that can lead to bacteremia and subsequent sepsis long after the initial infection104,105. It is known that endocarditis and sepsis can be caused by E. rhusiopathiae colonisation in humans6,13,106,107. Moreover, the mortality rate of this endocarditis is almost twice as high as that of the same pathology caused by other organisms103.
The presence of E. rhusiopathiae DNA in the pulp chamber of Sk213 teeth indicates its intravital circulation in the blood, i.e., a septic form of the disease1,2,108. According to Principe et al.,109 septic forms cause mortality in 12.5% of cases in humans (35–40% in cases associated with endocarditis110). At the same time, comorbidity increases the risk of developing fatal forms of erysipeloid111,112.
Cases discussed above are primarily associated with farmers, abattoir workers, butchers, fishermen, etc.4,5,13,14. For the Alans, since the beginning of the 1st millennium A.D., large settlements with areas reaching up to one and a half square kilometres are known113. In these settlements, sedentary inhabitants were actively engaged in farming and herding, as evidenced by the finds of hand millstones and the composition of herds reconstructed from animal bone remains33. The herds included cattle, goats, sheep, horses, and pigs114. A zoonotic disease in a member of a community whose main speciality was pastoralism is clearly an expected event115,116,117. Judging by the fact that the individual in burial 213 is buried in a pit rather than in a catacomb, she likely belonged to a low social class. This is supported by the scarcity of grave goods, represented by a single bronze earring. The fact that the grave was dug carelessly and contained only a single, simple artifact strongly suggests that the girl was also of a low social status, which could have indirectly compromised her immune system and overall health, potentially increasing her susceptibility to infectious diseases.
The high abundance of E. rhusiopathiae DNA in Sk213 clearly indicates human erysipeloid and observed osteopathology may be a direct consequence of this infection. Bony manifestations of erysipeloid are known from modern clinical studies but are largely overlooked during analysis of ancient skeletons. Very few studies describe osteopathological changes supposedly caused by erysipeloid in ancient remains of humans118 and farm animals119,120. Skeletal lesions ascribed to erysipeloid include arthritic changes of the sacrum118, arthropathy of the limb joints119 and ankylosis of the spine118,120. In no case have these observations been confirmed by genetic methods. Though differential diagnosis based on morphological manifestations of skeletal pathology does not allow us to exclude bone tuberculosis in Sk213, given the increased aggressiveness of E. rhusiopathiae disease in the presence of other chronic diseases, co-infection of the child from grave 213 with Mycobacterium sp. and E. rhusiopathiae would by a conservative reason to explain her early depth. Additional paleogenetic studies will be needed to determine the significance of the impact of E. rhusiopathiae on ancient herders.
Methodological limitations
Although our reconstruction achieved a near-complete genome coverage, it is inherently limited by the reference-guided assembly approach used, which could have resulted in the loss of highly divergent or unique genomic regions. Therefore, our analysis cannot rule out the presence in the ancient genome of genes missing from the reference/modern genomes. Further studies using reference-free assembly strategies will be required to capture the full genetic diversity of ancient E. rhusiopathiae.
Data availability
Data is provided within the manuscript and supplementary files. The raw sequence data was uploaded in NCBI, under the Bioproject: PRJNA1119393, BioSample: SAMN41652521, SRA: SRR29344859. The assembled genome ERA_01 have NCBI accession number JBEFKQ000000000, spaA gene: PP928806 and serotype-determining region: PP993147. Direct links: Bioproject: https://www.ncbi.nlm.nih.gov/bioproject/PRJNA1119393/. BioSample: https://www.ncbi.nlm.nih.gov/biosample/SAMN41652521/. SRA: https://www.ncbi.nlm.nih.gov/sra/SRR29344859/. WGS: https://www.ncbi.nlm.nih.gov/nuccore/JBEFKQ000000000/. spaA gene: https://www.ncbi.nlm.nih.gov/nuccore/PP928806.1/. serotype-determining region: https://www.ncbi.nlm.nih.gov/nuccore/PP993147.1/
References
Duchêne, S., Ho, S. Y., Carmichael, A. G., Holmes, E. C. & Poinar, H. The recovery, interpretation and use of ancient pathogen genomes. Curr. Biol. 30, R1215–R1231 (2020).
Malyarchuk, A. B. et al. Genomics of ancient pathogens: First advances and prospects. Biochem. Mosc. 87, 242–258 (2022).
Brooke, C. J. & Riley, T. V. Erysipelothrix rhusiopathiae: Bacteriology, epidemiology and clinical manifestations of an occupational pathogen. J. Med. Microbiol. 48, 789–799 (1999).
Reboli, A. C. & Farrar, W. E. Erysipelothrix rhusiopathiae: An occupational pathogen. Clin. Microbiol. Rev. 2, 354–359 (1989).
Wang, Q., Chang, B. J. & Riley, T. V. Erysipelothrix rhusiopathiae. Vet. Microbiol. 140, 405–417 (2010).
Rostamian, M., Rahmati, D. & Akya, A. Clinical manifestations, associated diseases, diagnosis, and treatment of human infections caused by Erysipelothrix rhusiopathiae: A systematic review. Germs 12, 16 (2022).
Kheng, M. et al. Sacro-iliite à Erysipelothrix rhusiopathiae révélant une endocardite tricuspide, premier cas rapporté sur le bouclier des guyanes: Cas clinique et revue de la littérature. [Sacroiliitis with Erysipelothrix Rhusiopathiae revealing tricuspid endocarditis, the first case reported on the Guiana Shield: Clinical case and review of the literature]. Médecine Tropicale et Santé Internationale 2, mtsi-v2i3 (2022).
Upapan, P. & Chayakulkeeree, M. Erysipelothrix rhusiopathiae bacteremia without endocarditis associated with psoas abscess: The first case report in Thailand. J. Med. Assoc. Thai. 97, S232–S236 (2014).
Neumann, D., Hafner, M. & Dorn, U. Septic arthritis caused by Erysipelotrix rhusiopathiae: Case report and review of the literature. Zeitschrift fur Orthopadie und Unfallchirurgie 147, 740–742 (2009).
Goossens, J. et al. Polyarticular septic arthritis caused by Erysipelothrix rhusiopathiae infection. Intern. Med. J. 52, (2022).
Mukhopadhyay, C., Shah, H., Vandana, K., Munim, F. & Vijayan, S. A child with Erysipelothrix arthritis–beware of the little known. Asian Pac. J. Trop. Biomed. 2, 503–504 (2012).
Alawdah, L. S., Campbell, J. N., Pollock, N. & Watnick, P. I. Erysipelothrix rhusiopathiae suppurative arthritis in a 12-year-old boy after an unusual fresh water exposure. Pediatr. Infect. Dis. J. 36, 431–433 (2017).
Veraldi, S., Girgenti, V., Dassoni, F. & Gianotti, R. Erysipeloid: A review. Clin. Exp. Dermatol. 34, 859–862 (2009).
Spiteri, M. & Taylor-Robinson, A. Erysipelothrix rhusiopathiae: An important cause of bacterial disease in farmed pigs and an occupational pathogen of humans. Int. J. Clin. Med. Microbiol. https://doi.org/10.15344/2456-4028/2018/134 (2018).
Zautner, A. E. et al. Human Erysipelothrix rhusiopathiae infection via bath water–case report and genome announcement. Front. Cell. Infect. Microbiol. 12, 981477 (2022).
Ogawa, Y. et al. The genome of Erysipelothrix rhusiopathiae, the causative agent of swine erysipelas, reveals new insights into the evolution of Firmicutes and the organism’s intracellular adaptations. J. Bacteriol. 193, 2959–2971 (2011).
Kwok, A. H., Li, Y., Jiang, J., Jiang, P. & Leung, F. C. Complete genome assembly and characterization of an outbreak strain of the causative agent of swine erysipelas–Erysipelothrix rhusiopathiae SY1027. BMC Microbiol. 14, 1–12 (2014).
Tang, H., Xie, J., Wang, L., Liu, F. & Wu, J. Complete genome sequence of Erysipelothrix rhusiopathiae strain GXBY-1 isolated from acute swine erysipelas outbreaks in South China. Genomics Data 8, 70–71 (2016).
Grazziotin, A. L. et al. Comparative genomics of a novel clade shed light on the evolution of the genus Erysipelothrix and characterise an emerging species. Sci. Rep. 11, 3383 (2021).
Li, Y. et al. Proteomic and transcriptomic analyses of swine pathogen Erysipelothrix rhusiopathiae reveal virulence repertoire. PLoS ONE 11, e0159462 (2016).
Borrathybay, E., Gong, F., Zhang, L. & Nazierbieke, W. Role of surface protective antigen A in the pathogenesis of Erysipelothrix rhusiopathiae strain C43065. J. Microbiol. Biotechnol. 25, 206–216 (2015).
Wu, C. et al. The C-terminal repeat units of SpaA mediate adhesion of Erysipelothrix rhusiopathiae to host cells and regulate its virulence. Biology (Basel) 11, 1010. https://doi.org/10.3390/biology11071010 (2022).
Groeschel, M. et al. An unusual case of Erysipelothrix rhusiopathiae prosthetic joint infection from the Canadian arctic: Whole genome sequencing unable to identify a zoonotic source. BMC Infect. Dis. 19, 1–7 (2019).
Shimoji, Y. et al. Development of a multiplex PCR-based assay for rapid serotyping of Erysipelothrix species. J. Clin. Microbiol. 58, e00315-e320 (2020).
Janßen, T. et al. A combinational approach of multilocus sequence typing and other molecular typing methods in unravelling the epidemiology of Erysipelothrix rhusiopathiae strains from poultry and mammals. Vet. Res. 46, 1–15 (2015).
Forde, T. et al. Genomic analysis of the multi-host pathogen Erysipelothrix rhusiopathiae reveals extensive recombination as well as the existence of three generalist clades with wide geographic distribution. BMC Genomics 17, 1–15 (2016).
Forde, T. L. et al. Genomic and immunogenic protein diversity of Erysipelothrix rhusiopathiae isolated from pigs in Great Britain: Implications for vaccine protection. Front. Microbiol. 11, 418 (2020).
Forde, T. L. et al. Bacterial genomics reveal the complex epidemiology of an emerging pathogen in arctic and boreal ungulates. Front. Microbiol. 7, 1759 (2016).
Söderlund, R. et al. Comparative genome analysis of Erysipelothrix rhusiopathiae isolated from domestic pigs and wild boars suggests host adaptation and selective pressure from the use of antibiotics. Microbial Genomics 6, e000412 (2020).
Webster, J. et al. Population structure and genomic characteristics of Australian Erysipelothrix rhusiopathiae reveals unobserved diversity in the Australian pig industry. Microorganisms 11, 297 (2023).
López, M. J. B. Estudio paleogenómico de patógenos humanos en población prehispánica y colonial de México. PhD Thesis. In [Paleogenomic study of human pathogens in the pre-Hispanic and colonial population of Mexico] (Universidad Nacional Autónoma de México, Ciudad de México, México, 2021).
Kusliy, M. A. 2022. Geneticheskoe raznoobrazie drevnikh i sovremennykh loshadey Altaya i sopredel’nykh territoriy. PhD Thesis. In [Genetic diversity of ancient and modern horses of Altai and adjacent territories] (Novosibirsk, Russia, 2022).
Kadieva, A. A. & Demidenko, S. V. Pogrebenie rannesarmatskogo vremeni iz mogil’nika Zayukovo-3 (Kabardino-Balkarskaya respublika) [Burial of the early sarmatian time from the Zayukovo-3 burial ground (Kabardino-Balkaria)]. Caucasology 2, 33–40 (2017).
Korobov, D.S. Modelirovanie sel’skokhozyajstvennyj ugodij alan Kislovodskoj kotloviny V–VIII vv. In [Modeling the agricultural lands of the Alanians in the Kislovodsk Basin in the 5th–8th centuries] pp 249–268 (In the circle of ideas: Models and technologies for historical reconstruction, Russia, Novosibirsk, 2010).
Massler, M., Schour, I. & Poncher, H. Developmental pattern of the child as reflected in the calcification pattern of the teeth. Am. J. Dis. Child. 62, 33–67 (1941).
Alekseev, V. P. & Debec, G. F. Kraniometrija: Metodika Antropologicheskich Issledovanij [Craniometry. Methods of Anthropological Studies]. (Nauka, Moscow, 1964).
Alekseev, V. P. Osteometrija: Metodika Antropologicheskih Issledovanij [Osteometry: Methodology of Anthropological Research] (Nauka, 1966).
Stloukal, M. & Hanáková, H. Die länge der längsknochen altslawischer bevölkerungen unter besonderer berücksichtigung von wachstumsfragen. Homo 29, 53–69 (1978).
Ubelaker, D. H. Estimating age at death from immature human skeletons: An overview. J. Forensic Sci. 32, 1254–1263 (1987).
Brothwell, D. R. Digging up bones: The excavation, treatment, and study of human skeletal remains (Cornell University Press, New York, 1981).
Lovejoy, C. O., Russell, K. F. & Harrison, M. L. Long bone growth velocity in the Libben population. Am. J. Hum. Biol. 2, 533–541 (1990).
Szilvássy, J., Kritscher, H. & Hauser, G. Eine urnenfelderzeitliche mehrfachbestattung in stillfried an der march, NÖ. [An Urnfield period multiple burial in Stillfried an der March, Lower Austria] in Stillfried: Archäologie—anthropologie (eds. Felgenhauer, F., Szilvássy, J., Kritscher, H. & Hauser, G.) vol. 3 9–76 (Veröffentlichungen des Museums für Ur- und Frühgeschichte Stillfried, Stillfried, 1988).
Rose, J. C. et al. Skeletal database committee recommendations. Paleopathology newsletter supplement (1991).
Cox, M. & Mays, S. Human osteology: In archaeology and Forensic science (Cambridge University Press, Cambridge, 2000).
Bronk Ramsey, C. Radiocarbon calibration and analysis of stratigraphy: The OxCal program. Radiocarbon 37, 425–430 (1995).
Reimer, P. J. et al. The IntCal20 Northern hemisphere radiocarbon age calibration curve (0–55 cal kBP). Radiocarbon 62, 725–757 (2020).
Keller, M. & Scheib, C. L. Decontamination of tooth roots/petrous bone cores for ancient DNA extraction. (2023).
Neumann, G. U., Valtueña, A. A., Fellows Yates, J., Stahl, R. & Brandt, G. Tooth sampling from the inner pulp chamber for ancient DNA extraction v.1. (2020).
Rohland, N., Glocke, I., Aximu-Petri, A. & Meyer, M. Extraction of highly degraded DNA from ancient bones, teeth and sediments for high-throughput sequencing. Nat. Protoc. 13, 2447–2461 (2018).
Clavel, P. et al. Improving the extraction of ancient Yersinia pestis genomes from the dental pulp. Iscience 26, (2023).
Chen, S. Ultrafast one-pass FASTQ data preprocessing, quality control, and deduplication using fastp. Imeta 2, e107 (2023).
Wood, D. E., Lu, J. & Langmead, B. Improved metagenomic analysis with Kraken 2. Genome Biol. 20, 1–13 (2019).
Ma, B. et al. A genomic catalogue of soil microbiomes boosts mining of biodiversity and genetic resources. NatureCommunications 14, 7318 (2023).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Picard toolkit. Broad Institute, GitHub repository (2019).
Jónsson, H., Ginolhac, A., Schubert, M., Johnson, P. L. & Orlando, L. mapDamage2.0: Fast approximate Bayesian estimates of ancient DNA damage parameters. Bioinformatics 29, 1682–1684 (2013).
Borry, M., Hübner, A., Rohrlach, A. B. & Warinner, C. PyDamage: Automated ancient damage identification and estimation for contigs in ancient DNA de novo assembly. PeerJ 9, e11845 (2021).
Okonechnikov, K., Conesa, A. & García-Alcalde, F. Qualimap 2: Advanced multi-sample quality control for high-throughput sequencing data. Bioinformatics 32, 292–294 (2016).
Weissensteiner, H. et al. Contamination detection in sequencing studies using the mitochondrial phylogeny. Genome Res. 31, 309–316 (2021).
Ralf, A., Montiel González, D., Zhong, K. & Kayser, M. Yleaf: Software for human Y-chromosomal Haplogroup inference from next-generation sequencing data. Mol. Biol. Evol. 35, 1291–1294 (2018).
Chen, H., Lu, Y., Lu, D. & Xu, S. Y-LineageTracker: A high-throughput analysis framework for Y-chromosomal next-generation sequencing data. BMC Bioinform. 22, 114 (2021).
Renaud, G. et al. Schmutzi: Estimation of contamination and endogenous mitochondrial consensus calling for ancient DNA. Genome Biol. 16, 224 (2015).
Müller, R., Roberts, C. A. & Brown, T. A. Complications in the study of ancient tuberculosis: Non-specificity of IS6110 PCRs. STAR Sci. Technol. Archaeol. Res. 1, 1–8 (2015).
Dec, M., Nowak, T., Webster, J. & Wódz, K. Serotypes, antimicrobial susceptibility, and potential mechanisms of resistance gene transfer in Erysipelothrix rhusiopathiae strains from waterfowl in Poland. Int. J. Mol. Sci. 25, 12192 (2024).
Huang, W. et al. First identification of human infection with Erysipelothrix piscisicarius by metagenomic next-generation sequencing. Emerg. Microbes Infect. 11, 2781–2784 (2022).
Kucsera, G. Proposal for standardization of the designations used for serotypes of Erysipelothrix rhusiopathiae (Migula) buchanan. Int. J. Syst. Evol. Microbiol. 23, 184–188 (1973).
Bankevich, A. et al. SPAdes: A new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 19, 455–477 (2012).
Seemann, T. Snippy: Fast bacterial variant calling from NGS reads. (2015).
Haeseler, A. von, Schmidt, H. A., Bui, M. Q. & Nguyen, L. T. IQ-TREE: A fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. (2014).
Rambaut, A., Lam, T. T., Max Carvalho, L. & Pybus, O. G. Exploring the temporal structure of heterochronous sequences using TempEst (formerly path-o-gen). Virus Evol. 2, vew007 (2016).
Duvaud, S. et al. Expasy, the swiss bioinformatics resource portal, as designed by its users. Nucleic Acids Res. 49, W216–W227 (2021).
Edgar, R. C. MUSCLE: Multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004).
Okonechnikov, K., Golosova, O., Fursov, M. & Team, U. Unipro UGENE: A unified bioinformatics toolkit. Bioinformatics 28, 1166–1167 (2012).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Shiraiwa, K., Ogawa, Y., Nishikawa, S., Eguchi, M. & Shimoji, Y. Identification of serovar 1a, 1b, 2, and 5 strains of Erysipelothrix rhusiopathiae by a conventional gel-based PCR. Vet. Microbiol. 225, 101–104 (2018).
Cantalapiedra, C. P., Hernández-Plaza, A., Letunic, I., Bork, P. & Huerta-Cepas, J. eggNOG-mapper v2: Functional annotation, orthology assignments, and domain prediction at the metagenomic scale. Mol. Biol. Evol. 38, 5825–5829 (2021).
Seemann, T. Prokka: Rapid prokaryotic genome annotation. Bioinformatics 30, 2068–2069 (2014).
Page, A. J. et al. Roary: Rapid large-scale prokaryote pan genome analysis. Bioinformatics 31, 3691–3693 (2015).
Parks, D. H., Imelfort, M., Skennerton, C. T., Hugenholtz, P. & Tyson, G. W. CheckM: Assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes. Genome Res. 25, 1043–1055 (2015).
Developers, P. Pysam: A python module for reading, manipulating, and analyzing genomic data. (2009).
Aziz, R. K. et al. The RAST server: Rapid annotations using subsystems technology. BMC Genomics 9, 1–15 (2008).
Grant, J. R. et al. Proksee: In-depth characterization and visualization of bacterial genomes. Nucleic Acids Res. 51, W484–W492 (2023).
Guellil, M. aDNA-BAMPlotter. (2023).
Letunic, I. & Bork, P. Interactive tree of life (iTOL) v5: An online tool for phylogenetic tree display and annotation. Nucleic Acids Res. 49, W293–W296 (2021).
Yachida, W. PyGenomeViz web. (2023).
Mastykova A.V. Zhenskiy kostyum zentral’nogo i zapadnogo predkavkaz’ja v kontse IV - seredine VI v.n.e. In [Female Costume of the Central and Western Ciscaucasia in the Late 4th - Mid-6th Centuries A.D.] 500 (Institute of Archaeology RAS, Moscow, 2009).
Guinet, B. et al. Ancient host-associated microbes obtained from mammoth remains. Cell 2, S0092-8674(25)00917-1 (2025).
Bowers, R. M. et al. Minimum information about a single amplified genome (MISAG) and a metagenome-assembled genome (MIMAG) of bacteria and archaea. Nat. Biotechnol. 35, 725–731 (2017).
Eroshenko, G. A. et al. Yersinia pestis strains of ancient phylogenetic branch 0. ANT are widely spread in the high-mountain plague foci of Kyrgyzstan. PLoS ONE 12, e0187230 (2017).
Long, G. S. et al. A 14th century CE Brucella melitensis genome and the recent expansion of the western Mediterranean clade. PLoS Pathog. 19, e1011538 (2023).
Orgeur, M., Sous, C., Madacki, J. & Brosch, R. Evolution and emergence of Mycobacterium tuberculosis. FEMS Microbiol. Rev. 48, fuae006 (2024).
Shimoji, Y. et al. Wild boars: A potential source of Erysipelothrix rhusiopathiae infection in Japan. Microbiol. Immunol. 63, 465–468 (2019).
Menardo, F., Duchêne, S., Brites, D. & Gagneux, S. The molecular clock of Mycobacterium tuberculosis. PLoS Pathog. 15, e1008067 (2019).
Eaton, K. et al. Plagued by a cryptic clock: Insight and issues from the global phylogeny of Yersinia pestis. Commun. Biol. 6, 23 (2023).
Ortner, D. J. Identification of pathological conditions in human skeletal remains Vol. 645 (Academic Press, Amsterdam, 2003).
Pálfi, G. et al. Unusual spinal tuberculosis in an Avar age skeleton (csongrád-felgyő, Ürmös-tanya, hungary): A morphological and biomolecular study. Tuberculosis 95, S29–S34 (2015).
Romney, M., Cheung, S. & Montessori, V. Erysipelothrix rhusiopathiae endocarditis and presumed osteomyelitis. Can. J. Dis. Med. Microbiol. 12, 254–256 (2001).
Lorenz, M. L., Bouton, T. C. & Caliendo, A. M. First reported case of vertebral osteomyelitis due to Erysipelothrix rhusiopathiae. IDCases 11, 3–5 (2018).
Maillard, A. et al. Osteoarticular infections caused by Erysipelothrix rhusiopathiae: Case report and literature review. In Open forum infectious diseases vol. 8 ofab461 (Oxford University Press US, 2021).
Andrychowski, J., Jasielski, P., Netczuk, T. & Czernicki, Z. Empyema in spinal canal in thoracic region, abscesses in paravertebral space, spondylitis: In clinical course of zoonosis Erysipelothrix rhusiopathiae. Eur. Spine J. 21, 557–563 (2012).
Meric, M. & Keceli Ozcan, S. Erysipelothrix rhusiopathiae pneumonia in an immunocompetent patient. J. Med. Microbiol. 61, 450–451 (2012).
Denes, E., Camilleri, Y., Fiorenza, F. & Martin, C. First case of osteomyelitis due to Erysipelothrix rhusiopathiae: Pubic osteomyelitis in a gored farmer. Int. J. Infect. Dis. 30, 133–134 (2015).
Umana, E. Erysipelothrix rhusiopathiae: An unusual pathogen of infective endocarditis. Int. J. Cardiol. 88, 297–299 (2003).
Shimoji, Y. Pathogenicity of Erysipelothrix rhusiopathiae: Virulence factors and protective immunity. Microbes Infect. 2, 965–972 (2000).
Seru, L. V. et al. Genomic characterization and virulence gene profiling of Erysipelothrix rhusiopathiae isolated from widespread muskox mortalities in the Canadian arctic archipelago. BMC Genomics 14, 691 (2024).
Challa, H. R., Tayade, A. C., Venkatesh, S. & Nambi, P. S. Erysipelothrix bacteremia; Is endocarditis a rule?. J. Global Infect. Dis. 15, 31–34 (2023).
Volpicelli, L., Oliva, A. & Venditti, M. Did the updated duke criteria missed Erysipelothrix rhusiopathiae from the list of typical microorganisms causing infective endocarditis?. Infez. Med. 31, 425 (2023).
Mai, B. H. A., Drancourt, M. & Aboudharam, G. Ancient dental pulp: Masterpiece tissue for paleomicrobiology. Mol. Genet. Genomic Med. 8, e1202 (2020).
Principe, L. et al. Erysipelothrix rhusiopathiae bacteremia without endocarditis: Rapid identification from positive blood culture by MALDI-TOF mass spectrometry. A case report and literature review. Infect. Dis. Rep. 8, 6368 (2016).
Nielsen, J. J., Blomberg, B., Gaı̈ni, S. & Lundemoen, S. Aortic valve endocarditis with Erysipelothrix rhusiopathiae: A rare zoonosis. Infect. Dis. Rep. 10, 7770 (2018).
Hua, P. et al. Erysipelothrix rhusiopathiae-induced aortic valve endocarditis: Case report and literature review. Int. J. Clin. Exp. Med. 8, 730 (2015).
Kobayashi, K. et al. Erysipelothrix rhusiopathiae bacteremia following a cat bite. IDCases 18, e00631 (2019).
Artamonov M.I. Istorija Hazar. [The History of the Khazars]. 523 (Published by the Hermitage Museum, Leningrad 1962).
Korobov D.S. Alany Severnogo Kavkaza: Etnos, Archeologija, Paleogenetika. [Alans of the North Caucasus: Ethnos, Archeology, Paleogenetics]. 156 (Nestor-Istoriya, Saint Petersburg, 2019).
Recht, J., Schuenemann, V. J. & Sánchez-Villagra, M. R. Host diversity and origin of zoonoses: The ancient and the new. Animals 10, 1672 (2020).
Bendrey, R. & Martin, D. Zoonotic diseases: New directions in human–animal pathology. Int. J. Osteoarchaeol. 32, 548–552 (2022).
Weiss, R. A. & Sankaran, N. Emergence of epidemic diseases: Zoonoses and other origins. Faculty Rev. 11, 2 (2022).
Hudson, C., Butler, R. & Sikes, D. Arthritis in the prehistoric southeastern United States: Biological and cultural variables. Am. J. Phys. Anthropol. 43, 57–62 (1975).
O’Connor, T. P. & O’Connor, T. The archaeology of animal bones (Texas A&M University Press, Texas, 2000).
Teegen, W. R. & Schultz, M. Trauma in Starigard/Oldenburg (Northern Germany). In San Antonio meeting report: 27th annual meeting of the paleopathology association (San Antonio, Texas, 2000).
Acknowledgements
Morphological research was carried out under partial support of MSU Shared Research Equipment Center “Technologies for obtaining new nanostructured materials and their complex study”, National Project “Science” and MSU Program of Development. This work was funded by the Russian Science Foundation (Project No. 25-18-00322).
Funding
Russian Science Foundation, 25-18-00322.
Author information
Authors and Affiliations
Contributions
A.A.K.—Conceptualisation, Methodology, Writing the original draft, Formal analysis, Investigation, Data curation, Review and editing, N.Y.B.—Writing the original draft, Investigation, Data curation, A.O.I.—Writing the original draft, Investigation, Review, Formal analysis, Visualisation, Software, D.A.S—Writing the original draft, Formal analysis, Investigation, Software, E.I.B.—Methodology, Writing the original draft, Investigation, A.A.P.—Writing the original draft, Investigation, A.A.K.—Resources, Review and editing, T.R.T. – Writing the original draft, Investigation, M.V.K.—Writing the original draft, Investigation, Software, Review, A.V.P.—Writing the original draft, Investigation, S.V.D.—Resources, Review and editing, E.I.K.—Resources, Review and editing, A.P.B.—Conceptualisation, Supervision, Resources, Review and editing, K.V.S.—Conceptualisation, Supervision, Resources, Review and editing.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Kritsky, A.A., Berezina, N.Y., Ivanova, A.O. et al. An ancient Erysipelothrix rhusiopathiae genome recovered from 1400-year-old human remains in the Northern Caucasus. Sci Rep 16, 7097 (2026). https://doi.org/10.1038/s41598-026-37742-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-026-37742-1