Abstract
MUC14/Endomucin, a transmembrane mucin, is a potential prognostic biomarker in malignancies. This study aimed to elucidate the functional impact of MUC14 on tumor proliferation, migration, immune microenvironment modulation, and cisplatin response in lung adenocarcinoma (LUAD), and investigate its molecular mechanisms. LUAD cell lines with MUC14 overexpression (MUC14-OE) or silencing were constructed. Malignant behaviors were assessed via CCK-8, Transwell, and colony formation assays. Immune cell infiltration was quantified by CD3+/CD8 + immunohistochemistry. Subcutaneous xenograft and tail-vein metastasis murine models evaluated in vivo tumor progression and cisplatin responsiveness. Mechanisms were characterized using FRET and western blotting. Multiplatform bioinformatics analysis of public databases correlated MUC14 expression with clinical outcomes, immune infiltration, and chemotherapy response. MUC14-OE inhibited LUAD cell proliferation, migration, colony formation, and adhesion, while silencing promoted these phenotypes. MUC14 expression positively correlated with CD3+/CD8 + T-cell infiltration. In vivo, MUC14-OE suppressed subcutaneous tumor growth, lung metastasis, and enhanced cisplatin efficacy. Mechanistically, MUC14 inhibited integrin α8β6 clustering, suppressing PI3K/AKT and MAPK/ERK signaling. Cisplatin sensitization involved JNK/c-Jun pathway activation. This study establishes MUC14 as a multifunctional tumor suppressor in LUAD. It inhibits integrin α8β6-mediated PI3K/AKT and MAPK/ERK signaling to suppress tumor growth, promotes CD8+ T-cell infiltration, and augments cisplatin sensitivity via the JNK/c-Jun pathway. These findings nominate MUC14 as a prognostic biomarker and therapeutic target, suggesting combinatorial strategies integrating immunotherapy and chemotherapy.
Similar content being viewed by others
Introduction
Lung adenocarcinoma (LUAD) accounts for 40–55% of all lung cancer cases1. Despite recent advances in clinical management, the 5-year survival rate of LUAD patients remains below 20%, and 57–80% of patients eventually develop metastasis2. Cisplatin is still the standard first-line chemotherapeutic agent for advanced LUAD; however, its efficacy is severely limited by the development of drug resistance, which is primarily driven by aberrant DNA repair mechanisms, drug inactivation, and dysregulated apoptosis3,4. Thus, identifying novel therapeutic targets to overcome cisplatin resistance is of great clinical significance.
The transmembrane mucin MUC14 (also known as EMCN) is defined by its extracellular O-glycosylation modifications and cytoplasmic phosphorylation motifs5,6. Previous studies have demonstrated that MUC14 overexpression promotes cell proliferation and metastasis in colon cancer7. However, its broader oncogenic functions across different cancer types remain incompletely characterized. Specifically, the role of MUC14 in lung cancer—especially its association with cisplatin response—has not been fully elucidated, except for one study showing that endothelial MUC14 deficiency facilitates LUAD metastasis8. Notably, MUC14 has been reported to regulate cell-matrix interactions, as it can disrupt integrin-mediated cell adhesion by impairing the formation of focal adhesion complexes in fibroblasts9.
As well-established oncogenic drivers, integrins activate multiple pro-tumor signaling pathways, such as the FAK/Ras/MAPK and FAK/PI3K pathways10,11,12, High integrin α-5 expression correlates with poor prognosis and drives aggressive cellular behaviors in non-small cell lung cancer13. WISP-1-integrin ανβ3-STAT1 signaling mediates the suppressive effect of cancer-associated fibroblasts and apoptotic cancer cells on lung cancer cell proliferation14. In addition, integrins are directly linked to chemoresistance in LUAD: elevated expression of integrin α2β1 enhances migration and metastasis of non-small cell lung cancer (NSCLC), a category that includes LUAD15, and the binding of cancer cell-intrinsic PD-L1 to integrin β1 induces chemoresistance in NSCLC16, investigating whether MUC14 regulates cisplatin sensitivity through integrin-mediated pathways represents a promising strategy to overcome cisplatin resistance in LUAD.
Materials and methods
Cell lines
The A549, HCC827 and HEK293 cell lines were obtained from ATCC. We tested for mycoplasma contamination and verified cells by short tandem repeat (STR) test; morphology was confirmed by pathologist before the experiments. A549, HCC827 and HEK293 cells were maintained under standard culture conditions (37 °C, 5% CO2) using DMEM (for A549 and HEK293) and RPMI-1640 (for HCC827) media (Biosharp, China), supplemented with 10% fetal bovine serum (Biosharp, China), 100 U/mL penicillin, and 100 µg/mL streptomycin (Phygene, China).
Cell transfection
The MUC14 overexpression (MUC14-OE) plasmid was constructed by inserting the EMCN (NM_016242.4) coding sequence into the pcDNA3.1 + vector (FengHuiShengWu, China). Both the MUC14-OE and empty plasmids were purified from glycerol-preserved bacterial cultures using the plasmid extraction kit (Solarbio, China). Plasmid transfection into cells was performed with Lipofectamine 8000 (Beyotime, China), with empty vector transfection serving as the negative control.
Commercially synthesized small interfering RNAs (siRNAs) (Sango Biotech, China; sequences in Table 1) were transfected into cell lines using Lipofectamine 8000, followed by 24 h incubation at 37 °C. Transfection efficiency was validated by Quantitative Reverse Transcription Polymerase Chain Reaction (qRT-PCR) prior to functional assays, such as wound healing assay, CCK-8 assay, Transwell assay, colony formation assay and cell-adhesion assay. In order to ensure the effectiveness of siRNA, all the three siRNAs were transfected into the cells simultaneously in the experiments in vitro.
Wound healing assay
The A549-Vector cells/A549-MUC14 cells (5 × 105) were cultured in 6-well plates until they reached nearly 90% confluence. The same culture process was carried out for the HCC827 cells. A straight scratch was made in the center of each well using a P1000 pipette tip. Photographs were taken at 0 h and 24 h using a light microscope (EVOS, USA). Image processing and quantification were conducted using ImageJ software (Version 1.53t; National Institutes of Health, Bethesda, MD, USA; https://imagej.nih.gov/ij/). The same procedure was also performed for A549-siNC, A549-siMUC14, HCC827-siNC and HCC827-siMUC14 cells.
Transwell assay
The A549-Vector cells/A549-MUC14 cells (1 × 105) were plated into the upper compartment of a 24-well Transwell plate (Corning, USA) filled with serum - free medium. The same operation was conducted for the HCC827 cells. Subsequently, medium supplemented with 10% FBS was added to the lower compartments. After an incubation period of 24 h, non-migrated cells on the upper surface of the membrane were gently wiped off using a cotton swab. The migrated cells adhering to the lower surface were fixed with methanol and then stained with 0.5% crystal violet (Phygene, China). Once dried, pictures were taken under a light microscope (EVOS, Thermo Fisher, USA) at 40× magnification. The ImageJ software was employed to quantify the stained areas. The identical process was carried out for A549-siNC, A549-siMUC14, HCC827-siNC, and HCC827-siMUC14 cells as well.
For the Matrigel invasion assay, the upper chamber membrane was pre-coated with 100 µL of Matrigel (Corning, USA) diluted 1:8 in serum-free medium and allowed to polymerize at 37 °C for 2 h. The subsequent steps were identical to those described above for the migration assay.
Cell counting Kit-8 (CCK-8) assay
Two experimental cell groups (the vector and the MUC14-OE) of A549 and HCC827 cells were plated in 96-well plates at 8 × 103 cells/well with complete medium. Cellular proliferation was assessed on consecutive days (Days 1–3) by adding 10% CCK-8 reagent (Beyotime, China) followed by 30 min incubation at 37 °C, with optical density measured at 450 nm using a microplate reader. The same performance was applied to A549-siNC, A549-siMUC14, HCC827-siNC, and HCC827-siMUC14 cells.
Cell-adhesion assay
Collagen I (100 µg/mL), fibronectin (20 µg/mL), poly-L-lysine (100 µg/mL), and laminin (100 µg/mL) were diluted in Phosphate Buffered Saline (PBS) and used to coat 24-well plates (300 µL/well), followed by 1 h incubation at 37 °C. Wells were blocked with 0.5% bovine serum albumin (BSA) in medium (300 µL/well) for 1 h. Subsequently, serum-free medium containing 2 × 105 cells/well (A549 variants: Vector, MUC14-OE, siNC, siMUC14) was added for 60 min of adhesion under standard culture conditions. Non-adherent cells were removed by gentle PBS washing, and adherent cells were sequentially fixed with methanol (100 µL/well, 10 min) and stained with 0.5% crystal violet (100 µL/well, 10 min). Plates were rinsed with PBS, air-dried, and analyzed by photomicrography for cell quantification. The HCC827 cells underwent identical experimental procedures.
Colony formation assay
Following transfection, cells were trypsinized, counted, and seeded into 6-well plates at optimized densities of 100 (A549) or 300 (HCC827) cells/well to avoid overcrowding, cultured for 10–14 days with medium replenishment every 3 days, fixed with 4% paraformaldehyde (10 min), stained with 0.5% crystal violet (10 min), washed with PBS, and colonies containing > 50 cells were quantified.
Determination of 50% inhibitory concentration (IC50)
A549 and HCC827 cells (separately transfected with Vector, MUC14-OE, siNC, and siMUC14) were seeded in 96-well plates (1 × 104 cells/well, 100 µL/well), incubated for 24 h, treated with cisplatin (1, 5, 10, 20, 40 µmol/L) or controls (negative: complete medium; blank: cell-free medium) for 24 h, incubated with CCK-8 reagent (10 µL/well), and absorbance (450 nm) was measured. Cell viability (%) was calculated as [(OD_negative − OD_blank)/(OD_sample − OD_blank)] × 100. Dose-response curves (logC vs. viability) were fitted using GraphPad Prism to determine IC50.
Apoptosis assay
A549 and HCC827 cells were seeded in 6-well plates and divided into four groups: Vector, MUC14-OE, Vector+Cisplatin (at IC50), and MUC14-OE+Cisplatin (at IC50). After 24 h incubation, cells were stained with 5 µL Hoechst 33342 and 5 µL propidium iodide (PI) in 1 mL staining buffer for 30 min on ice. Cells were visualized using a fluorescence microscope (Olympus DP74, Japan) under blue (Hoechst) and red (PI) channels.
Quantitative reverse transcription polymerase chain reaction (qRT-PCR)
Total RNA was isolated from A549 cells and HCC827 cells using Trizol reagent (Life Sciences, China) following the manufacturer’s recommended protocol. RNA concentration was performed using a NanoDrop spectrophotometer (Thermo Fisher Scientific, USA). Subsequently, RNA samples were reverse transcribed into cDNA with HiScript II Q RT SuperMix (Vazyme, R122-01, China).
Quantitative PCR analysis was performed using AceQ-qPCR SYBR Green Master Mix (Vazyme, Q111-02) on a LightCycler480 II platform (Roche, Switzerland). Relative gene expression was calculated using the 2−ΔΔCt method with GAPDH as the endogenous control.
Immunohistochemistry (IHC)
The study was approved by the Medical Ethics Committee of the Second Affiliated Hospital of Harbin Medical University (YJSKY2023-264). 112 surgically excised LUAD specimens were obtained from the Second Affiliated Hospital of Harbin Medical University (between June 2023 and June 2024). Histopathological evaluation stratified the specimens into three differentiation grades: well-differentiated (n = 30), moderately differentiated (n = 44), and poorly differentiated (n = 38). IHC analysis was conducted using EDTA-mediated antigen retrieval (pH 8.0, 120 s) followed by incubation with Endomucin polyclonal antibody (Thermo Fisher Scientific, PA5-21395; 1:3000 dilution). Staining intensity was categorized into four levels: 0 (negative/no staining), 1+ (weak/light yellow), 2+ (positive/brown yellow), and 3+ (strong/tan). Simultaneously, the proportion of positive cells was quantified using a four-level scale: 1 (≤ 25%), 2 (26–50%), 3 (51–75%), and 4 (> 75%). The composite immunohistochemical score was calculated by multiplying the intensity and proportion scores.
Western blot
Total protein was extracted from A549 and HCC827 cells (separately transfected with Vector, MUC14-OE, siNC and siMUC14) using RIPA lysis buffer (BioSharp, China). Lysates were mixed with 5× loading buffer (BioSharp, China), denatured (100 °C, 5 min), and resolved via 10% SDS-PAGE (Epizyme, China). Proteins were transferred onto PVDF membranes (0.45 μm), blocked with 5% nonfat milk for 1 h, then incubated overnight (12–16 h, 4 °C) with primary antibodies (Beyotime, China) against: PI3 Kinase p85α (1:1000), AKT1/2/3 (1:1000), Phospho-AKT1 (Thr308) (1:500), ERK1 (1:1000), Phospho-Erk1 (Thr202/Tyr204)/Erk2(Thr185/Tyr187) (1:1000), c-Jun (1:1000), JNK1 + JNK2+JNK3 (1:1000), Phospho-JNK1/JNK2/JNK3 (Thr183/Thr183/Thr221) (1:1000), Bcl-2 (1:1000), Vimentin (1:1000), N-Cadherin(1:1000), E-Cadherin (1:1000), SLUG (1:1000), and Endomucin (Thermo Fisher Scientific, 1:1000). GAPDH (ZsBio, China, 1:2000) served as loading control. After washing three times with TBST for 10 min each, membranes were incubated with HRP-conjugated secondary antibodies (1:1000, 1 h). Signals were developed using ECL (Beyotime, China) and quantified using ImageJ by normalizing band intensity to GAPDH on a ChemiDoc system (Bio-Rad, USA). Fresh paired tumor/adjacent tissues from 8 LUAD patients (The Second Affiliated Hospital of Harbin Medical University) were processed following identical experimental protocols. Primary antibodies used: Endomucin (Thermo, PA5-21395; 1:1000) and GAPDH.
Fluorescence resonance energy transfer (FRET) assay
HEK293 cells were co-transfected with plasmids encoding MUC14-EGFP (donor) and ITGA8/B6-dsRED(acceptor). FRET imaging was performed on a Zeiss LSM 900 confocal microscope equipped with a 63× oil-immersion objective. Excitation was carried out using a 488 nm laser (the excitation range of EGFP is 400–568 nm). Emission collection: (1) Donor channel: 500–550 nm (EGFP); (2) Acceptor channel: 568–700 nm (dsRED); (3) FRET channel: The emission of dsRED was collected under EGFP excitation to detect energy transfer.
Survival related data mining of transmembrane mucins
Kaplan–Meier Plotter (https://kmplot.com/) was used to analyze the expression of transmembrane mucins in lung cancer to correlate with overall survival (OS) and free progressions (FPS). Statistical significance was determined using the log-rank test (P ≤ 0.05), identifying prognostic biomarkers.
Gene expression profiling interactive analysis (GEPIA)
GEPIA (http://gepia.cancer-pku.cn)17, a pan-cancer analysis platform integrating The Cancer Genome Atlas (TCGA) RNA-seq and clinical data, was used to assess transmembrane mucin expression differences between LUAD tumors and adjacent tissues (P ≤ 0.05). It also quantified MUC14 transcript levels across LUAD clinical stages ranked by transcripts per million (TPM), using F-statistics for intergroup heterogeneity analysis.
Pan-cancer data of MUC14
The expression profiles of MUC14 across multiple cancer types were analyzed using the Tumor Immune Estimation Resource (TIMER) database (https://cistrome.shinyapps.io/timer/)18. Tumor and normal tissues from TCGA pan-cancer cohort were compared. MUC14 expression levels were quantified as log2 TPM. Visualization was performed via box plots. Statistical significance between tumor and normal groups was evaluated using the Wilcoxon rank-sum test (P ≤ 0.05).
Identification of MUC14-associated integrin subunits
Transcriptomic data from the TCGA-LUAD cohort were analyzed to assess correlations between MUC14 expression and all integrin subunit genes. Pearson correlation coefficients were calculated for each subunit. The Benjamini-Hochberg (BH) method controlled the false discovery rate (FDR). Subunits with |r| > 0.3 and FDR < 0.05 were considered significant.
Cisplatin response analysis of MUC14
The expression of MUC14 association with cisplatin response in LUAD was assessed by Genomics of Drug Sensitivity in Cancer (GDSC) datasets (GDSC1/GDSC2, https://www.cancerrxgene.org/)19. The receiver operating characteristic (ROC) curve was accessed and the Area Under Curve (AUC) and corresponding P-value were calculated.
MUC14 correlation with immune cell infiltration analysis
We utilized TIMER database analysis to assess the correlation between MUC14 expression (log2 TPM values) and T cell subsets via Spearman rank correlation and multiple regression modeling. Additionally, scatter plots were generated to visualize relationships between MUC14 expression (log2 TPM) and tumor purity, as well as infiltration levels of cancer-associated fibroblasts, NK cells, Tregs, and CD8 + T cells, with Spearman’s Rho coefficients (-1 to 1) used to quantify monotonic associations. All analyses were performed in the R environment using standardized data.
Animal experiments
The animal experiments were approved under ethics protocol YJSDW2023-233. Male BALB/c nude mice (6-week-old, weighing 18–20 g, SPF grade, Vital River Laboratories, Beijing, China) were randomized into two groups (n = 5 per group). After 1 week of acclimatization, 2 × 106 A549-Vector or A549-MUC14 cells suspended in 100 µL PBS were subcutaneously injected into the right inguinal region of mice. Tumor dimensions were measured every 3 days using a digital caliper, with volume calculated as V = 0.5 × L × W2. On day 18 post-inoculation, mice were euthanized. Prior to euthanasia, mice were not anesthetized, as approved by our animal ethics protocol (YJSDW2023-233). Euthanasia was performed by rapid cervical dislocation carried out by trained personnel, which ensures immediate loss of consciousness. Death was confirmed by the absence of respiration, heartbeat, and corneal reflex. Tumors were excised, photographed, and weighed.
Twenty-four male BALB/c nude mice (6-week-old, SPF grade; Vital River Laboratories, China) were randomized into four groups (n = 6 per group): (1) A549-Vector + Cisplatin; (2) A549-MUC14 + Cisplatin; (3) A549-Vector + Saline; (4) A549-MUC14 + Saline. Mice received tail vein injections of 1 × 106 cells (A549-Vector or A549-MUC14) in 200 µL PBS. Starting from day 14 post-inoculation, cisplatin groups were administered 5 mg/kg cisplatin (200 µL in saline, i.p.) weekly for 5 weeks; controls received saline equally. Body weight was recorded weekly. On day 49, mice were euthanized by cervical dislocation. Lungs were fixed in 4% paraformaldehyde, paraffin-embedded, sectioned, and HE-stained. Metastatic foci were quantified by two pathologists blinded to group allocation. Additionally, the liver and kidney from one randomly selected mouse per group were collected, similarly processed into paraffin sections, and subjected to HE staining for histopathological observation.
Statistical analysis
Two groups were analyzed using GraphPad Prism software (Version 9.4.1; San Diego, CA, USA; https://www.graphpad.com) with a two-tailed unpaired t-test. Comparisons among three or more groups were performed using one-way ANOVA followed by Tukey’s post hoc test. All data are expressed as mean ± SEM. Differences were considered statistically significant when P ≤ 0.05.
Results
MUC14 is down-expressed in LUAD tissue and associated with favorable clinical outcomes
According to Kaplan-Meier survival analysis, the expression levels of transmembrane mucins were evaluated for their associations with OS and FPS in lung cancer patients. Patients with high MUC14 expression exhibited significantly longer OS (P = 0.0018) and delayed FPS (P = 0.00098) compared to the low-expression group (Fig. 1A,B). Consistently, analyses from the GEPIA database revealed that MUC14 expression was markedly reduced in cancer vs. normal (Fig. 1C, P < 0.05). Venn diagram analysis uniquely pinpointed MUC14 as the sole transmembrane mucin overlapping across all three criteria (OS, FPS, and cancer-normal expression), highlighting its distinctive role in LUAD (Fig. 1D). Additionally, GEPIA-based analysis of p-TNM staging in LUAD revealed statistically significant differences in MUC14 expression levels (Fig. 2B, F = 2.41, P = 0.0197). We further analyzed the expression of MUC14 in pan-cancers in the TCGA database, and the results showed that MUC14 was suppression across multiple malignancies, strongly supporting its tumor-suppressive potential (Fig. 2A).
Association of MUC14 with clinical outcomes in LUAD. (A,B) The Kaplan–Meier survival analysis for overall survival (OS) and free progressions (FPS) for different transmembrane mucins in lung cancer. (C) Expression of transmembrane mucins between cancer and normal in LUAD (GEPIA). (D) Venn diagram identifying MUC14 as the only transmembrane mucin associated with OS, FPT and cancer/normal expression. *P < 0.05; HR, Hazard ratio.
MUC14 expression in LUAD vs. adjacent normal tissues. (A) Pan-cancer MUC14 expression profile (TIMER). (B) A violin plot of MUC14 expression in LUAD pTNM stage (GEPIA). Pr (> F), the association between the significance probability value and the F value. (C,D) Representative IHC of MUC14 in LUAD vs. adjacent normal tissues. (E) The IHC staining score of MUC14 stratified by cancer differentiation grade. (F,G) Western blot showing MUC14 expression in eight LUAD patients (cancer vs. normal). *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001; ns, not significant; C, Cancer; N, Normal; TPM, Transcripts Per Kilobase Million.
Validation of clinical LUAD samples via IHC confirmed lower MUC14 expression levels in cancer tissues compared to normal tissues (100/112) (Fig. 2C,D, P < 0.0001). However, IHC scoring revealed no significant correlation between MUC14 expression and tumor differentiation grades (Fig. 2E). Western blot analysis was performed on 8 pairs of fresh LUAD tissues, revealing that the expression of MUC14 in cancer tissues was lower than that in normal tissues (8/8) (Fig. 2F,G, P < 0.001).
MUC14 inhibits the malignant phenotype of LUAD cells
To investigate MUC14’s role in LUAD cell biology, we transfected A549 and HCC827 cells with the MUC14 plasmid. The Transwell assay evidenced that MUC14-OE attenuated cell migration under starvation conditions (A549: P < 0.001, HCC827: P < 0.01) (Fig. 3A). Consistently, Matrigel-based Transwell invasion assays demonstrated that MUC14-OE attenuated cell invasion (A549: P < 0.0001, HCC827: P < 0.001) (Supplementary Fig. 1A,B). Furthermore, Western blot showed that MUC14-OE led to upregulated E-cadherin and downregulated N-cadherin, Vimentin, and SLUG expression, indicating the suppression of Epithelial-Mesenchymal Transition (EMT) (Supplementary Fig. 1C,D). Similarity, the wound healing assay exhibited delayed wound closure of MUC14-OE cells at 24 h (A549: P < 0.01, HCC827: P < 0.05) (Fig. 3B). The number of cells adhering to the four extracellular matrices (ECM) in MUC14-OE group was significantly less than that in vector group over the same time period (Fibronectin: A549, P < 0.01; HCC827, P < 0.001; Collagen I: A549, P < 0.01; HCC827, P < 0.01; Poly-L-lysine: A549, P < 0.01; HCC827, P < 0.001; Laminin: A549, P < 0.001; HCC827, P < 0.001) (Fig. 3C,D). Cell proliferation assay showed that MUC14-OE cells proliferated less than the vector cells on day3 (A549: P < 0.01, HCC827: P < 0.001) (Fig. 3E). Clonogenic assay revealed that MUC14-OE cells exhibited significantly reduced colony formation capacity compared to vector controls (A549: P < 0.001; HCC827: P < 0.01) (Fig. 3F). In vivo subcutaneous xenograft models using MUC14-OE A549 cells demonstrated significant suppression of tumor growth compared to control xenografts (5/5), with statistically significant decreases in tumor volume and weight (P < 0.0001, Fig. 3G–I).
Tumor-suppressive effects of MUC14-OE in LUAD. (A) Transwell assay showing suppressed migration in MUC14-OE cells under serum starvation after 24 h. (B) Wound healing assay exhibiting delayed wound closure of MUC14-OE cells after 24 h (yellow lines demarcate wound edges). (C,D) Impaired adhesion of extracellular matrix components (fibronectin, collagen I, poly-l-lysine, laminin) in MUC14-OE cells. (E) CCK-8 assay demonstrating reduced proliferation in MUC14-OE cells on day 2–3. (F) MUC14-OE attenuating colony formation capacity in A549 and HCC827 cells. (G–I) MUC14-OE inhibiting subcutaneous tumor growth in BALB/c nude mice (5/5). *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001; ns, not significant.
To conduct a comprehensive analysis of the impact of MUC14 on LUAD cells, siRNAs were employed to silence MUC14 expression in both A549 and HCC827 cell lines. The knockdown effect of the three siRNAs on MUC14 was verified by qPCR (A549: siMUC14-428, ns; siMUC14-377, P < 0.01; siMUC14-325, ns. HCC827: siMUC14-428, ns; siMUC14-377, P < 0.001; siMUC14-325, P < 0.0001) (Fig. 4A). CCK-8 assay suggested that siMUC14 cells exhibited significantly higher proliferation rates compared to controls on day 2–3 (A549: P < 0.05, HCC827: P < 0.01) (Fig. 4B). Furthermore, siMUC14 significantly augmented migratory potential in LUAD cells subjected to 24 h serum starvation, as determined by Transwell migration assay (A549: P < 0.05, HCC827: P < 0.001) (Fig. 4C). And the accelerated wound closure was observed in siMUC14 cells at 24 h (A549: P < 0.01, HCC827: P < 0.01) (Fig. 4D). SiMUC14 cells exhibited greater adherence to multiple ECM components when compared to siNC cells at identical time intervals (Fibronectin: A549, P < 0.001; HCC827, ns; Collagen I: A549, P < 0.01; HCC827, ns; Poly-L-lysine: A549, ns; HCC827, P < 0.001; Laminin: A549, ns; HCC827, P < 0.01) (Fig. 4E,F). Clonogenic assay revealed that siMUC14 cells exhibited significantly enhanced colony formation capacity compared to siRNA controls (A549: P < 0.01; HCC827: P < 0.001) (Fig. 4G,H).
siMUC14 aggravating LUAD malignancy. (A) Efficiency of MUC14 knockdown by three siRNAs at the RNA level. (B) CCK-8 assay revealing enhanced proliferative capacity in siMUC14 cells on day 2–3. (C) Transwell assay exhibiting augmented siMUC14 cells migration under serum deprivation after 24 h. (D) Wound healing assay exhibiting accelerated wound closure of siMUC14 cells after 24 h (yellow lines demarcate wound edges). (E,F) siMUC14 enhancing adhesion of A549 cells to fibronectin and collagen I; similarly, HCC827 cells showing increased binding to poly-L-lysine and laminin. (G,H) siMUC14-OE promoting colony formation capacity in A549 and HCC827 cells. *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001; ns, not significant.
These findings established MUC14 as a tumor suppressor in LUAD, directly accounting for the observed survival advantage in LUAD patients with higher MUC14 expression.
MUC14 prevents integrin α8β6 clustering, leading to the Inhibition of PI3K/AKT and MAPK/ERK signaling pathways
Since MUC14 has been confirmed impairing integrin-mediated cell-matrix interactions in the fibroblast, we investigated its interaction with integrins in LUAD to delve deeper into the molecular underpinnings. Given integrins’ structural complexity from different α and β subunit combinations, we performed genome-wide correlation analysis using TCGA dataset to identify MUC14-associated integrin subunits. The results showed that among all integrin subunits, integrin α8 and β6 exhibited the strongest correlation with MUC14 expression (Integrin α8: statistic = 0.616821371380505; P = 2.62657224505408e-55; FDR (BH) = 6.24999119454059e−53; Integrin β6: statistic = 0.311077958654011; P = 5.11206343099489e−13; FDR (BH) = 5.10899619293629e−12) (Fig. 5A). This finding clarified that the integrin α8β6 represented the most probable integrin subtype that directly interacts with MUC14. Since MUC14 and integrin are transmembrane proteins, we next confirmed direct physical interaction using FRET assays in live cells. We first tagged MUC14 plasmids with EGFP and integrin α8β6 plasmids with dsRED. Subsequently, co-transfection of MUC14-EGFP and integrin α8β6-dsRED into HEK293 cells yielded significant energy transfer (EGFP excitation → dsRED emission), demonstrating sub-10 nm proximity between MUC14 and α8β6 on the plasma membrane (Fig. 5B,C). It suggested that MUC14-OE could disrupt integrin α8β6 clustering at cell membrane.
MUC14 inhibiting PI3K/AKT and MAPK/ERK signaling pathways by preventing integrin α8β6 clustering. (A) Correlation analysis between MUC14 and different integrin subunits (TCGA): integrin α8 (ITGA8) and integrin β6 (ITGB6) exhibiting the strongest correlation with MUC14 expression. (B) Schematic of FRET methodology. (C) MUC14-EGFP and ITGA8/ITGB6-dsRED plasmids were co-transfected into HEK293 cells. Excitation with EGFP’s light yielded dsRED’s emission, suggesting an energy transition in FRET assay. (D,E) Western blot analysis revealing attenuated AKT and ERK phosphorylation in MUC14-OE cells; (F,G) siMUC14 enhancing phosphorylation of AKT and ERK. *P < 0.05, **P < 0.01, ***P < 0.001; ns, not significant.
Given the established role of PI3K/AKT and MAPK/ERK signaling as canonical oncogenic pathways downstream of integrins, we evaluated phosphorylation status of key components (p-AKT, p-ERK) in these cascades via western blot analysis. MUC14-OE reduced phosphorylation levels of AKT and ERK (Fig. 5D,E), whereas siMUC14 promoted the phosphorylation (Fig. 5F,G). These data establish that MUC14 directly binds integrin α8β6, prevents its clustering, and consequently inactivates oncogenic PI3K/AKT and MAPK/ERK signaling in LUAD cells.
MUC14 is associated with the immune microenvironment in LUAD
Tumor immune microenvironment (TIME) is a key determinant of tumor development and chemotherapy response. Enhanced peritumoral infiltration of cytotoxic T lymphocytes can notably boost chemotherapeutic response, contributing to a positive prognosis20,21. Therefore, we explore the correlation between MUC14 and TIME. We conducted a comprehensive analysis of immune cell correlations using the TIMER database. Our analysis revealed a statistically significant positive correlation between MUC14 expression level and effector T cell infiltration (P = 2.1e−14, R = 0.34). However, no significant associations were observed with effector Treg cells (P = 0.75, R = − 0.015) or exhausted T cell populations (P = 0.49, R = 0.032) (Fig. 6A–C). Furthermore, we identified significant correlations between MUC14 expression and multiple immune cell subsets, including cancer-associated fibroblasts (P = 1.27e−18, Rho = 0.382), NK cells (P = 3.58e−03, Rho = − 0.131), Treg cells (P = 1.11e−7, Rho = 0.236), and CD8 + T cells (P = 1.74e−2, Rho = 0.107) (Fig. 6D–G).
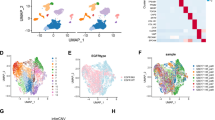
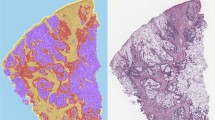
Correlation between MUC14 and immune microenvironment in LUAD. (A–C) TIMER analysis of the correlation between MUC14 expression and T cell subsets (effector T - cell, effector Treg T cell, exhausted T cell) via Spearman correlation of log2-transformed TPM values of MUC14 with three effector T-cell markers (CX3CR1, FGFBP2, FCGR3A), four effector Treg T cell markers (FOXP3, CTLA4, CCR8, TNFRSF9), and five exhausted T cell markers (HAVCR2, TIGIT, LAG3, PDCD1, CXCL13, LAYN). R quantifies the strength of association in multiple regression models, ranging from 0 to 1. (D–G) Scatter plots showing the relationships between tumor purity (percentage of tumor cells) and MUC14 expression (log2 TPM), as well as cancer-associated fibroblast/NK cell/T cell regulatory (Tregs)/T cell CD8 + infiltration levels and MUC14 expression (log2 TPM) in tumor samples. Rho is used to assess the monotonic relationship between variables, ranging from − 1 to 1. (H) Consecutive section IHC displaying relationships between MUC14, CD3 and CD8. (I) Correlation of MUC14 IHC score with CD3+/CD8 + T cells in the same field of view. P < 0.05 is considered statistically significant and r is used to assess the linear relationship between two variables, ranging from − 1 to 1.
To validate the results of database analysis, IHC staining targeting MUC14, CD3 and CD8 was performed from 52 LUAD patients. Representative IHC images displayed the spatial distribution and the IHC score of MUC14, along with consecutive sections stained for CD3 and CD8 markers within the same tumor microenvironment (Fig. 6H). Quantitative analysis revealed a significant positive correlation between MUC14 IHC score and the density of CD3 + and CD8 + T cells in proximity to MUC14-positive tumor cells (CD3: P = 1.6e−16, r = 0.87; CD8: P = 2.1e−8, r = 0.69) (Fig. 6I). The results collectively indicated that MUC14 expression was positively associated with the infiltration of CD3 + and CD8 + T cells in the tumor microenvironment of LUAD, nominating MUC14 as a biomarker candidate for LUAD immune modulation and cisplatin response.
MUC14 enhances cisplatin response via the JNK/c-Jun signaling pathway
Comprehensive analysis of the GDSC database demonstrated significantly elevated MUC14 levels in cisplatin-responsive LUAD patients compared to non-responders, the ROC curve showed an AUC = 0.626, with P = 1.9e−2 (GDSC1, Fig. 7A), and an AUC = 0.582, with P = 0.065 (GDSC2, Fig. 7B). CCK-8 assays revealed significantly lower IC50 values of cisplatin in MUC14-OE cells vs. vector cells: 18.93 µM vs. 45.67 µM in A549 cells, and 37.53 µM vs. 50.05 µM in HCC827 cells, suggesting that MUC14-OE enhances cisplatin sensitivity (Fig. 7C). Conversely, siMUC14 increased IC50 values: 63.74 µM vs. 41.46 µM in A549 cells, and 73.69 µM vs. 48.78 µM in HCC827 cells (Fig. 7D). To further elucidate whether the modulation of cisplatin sensitivity by MUC14 is coupled with drug-induced cell death, we evaluated cellular mortality through Hoechst and PI staining. The Hoechst and PI staining revealed that 24 h cisplatin-treated MUC14-OE groups exhibited higher cell mortality compared to 24 h cisplatin-treated Vector groups (A549 and HCC827: P < 0.01). In contrast, no significant difference in cell mortality was observed between untreated MUC14-OE and Vector groups. The above results indicate that MUC14-OE enhances the cytotoxic efficacy of cisplatin but does not affect cell apoptosis (Fig. 7E,F).
MUC14 enhancing cisplatin sensitivity via JNK/c-JUN activation. (A,B) GDSC1 and GDSC2 datasets showing MUC14 expression in cisplatin responders vs. non-responders and the Receiver operating characteristic (ROC) curves, respectively. The plot displays true positive rate (TPR, y-axis) against false positive rate (1 − TNR, x-axis). The area under the curve (AUC) reflects overall accuracy, while the strongest cut-off (dashed line) denotes the optimal threshold balancing sensitivity and specificity. (C,D) MUC14-OE reduces cisplatin resistance (lower IC50) in A549 and HCC827 cells, whereas siMUC14 increases resistance (higher IC50) in their cisplatin dose-response curves. (E,F) The Hoechst and PI staining of A549 and HCC827 cells under cisplatin treatment (24 h) or untreated conditions. (G,H) Western blot analysis revealing enhanced JNK phosphorylation and c-Jun expression, along with reduced BCL-2 expression in MUC14-OE cells. (I,J) siMUC14 suppressing JNK phosphorylation and c-Jun expression while upregulating BCL-2. *P < 0.05, **P < 0.01, ***P < 0.001. ****P < 0.0001.
As JNK/c-Jun signaling promotes cisplatin-induced apoptosis in different cancers (e.g., esophageal and colon cancer) and is a downstream pathway regulated by integrins22,23,24,25, we hypothesized that MUC14 enhances cisplatin sensitivity via this pathway. Therefore, we separately detected the changes of JNK/c-Jun pathway in MUC14-OE and siMUC14 cells after treatment with cisplatin, and the results showed that MUC14-OE activated JNK signaling (phospho-JNK/c-Jun upregulation) (Fig. 7G,H), whereas siMUC14 suppressed the pathway (Fig. 7I,J). These data confirmed that MUC14 enhanced cisplatin sensitivity in LUAD cells via JNK/c-Jun signaling pathway.
To further investigate the impact of MUC14 on tumor dissemination and its influence on cisplatin chemotherapy in vivo, we established tail vein xenograft tumor models in BALB/c nude mice (Fig. 8A). The tumor formation rates were as follows: 6/6 in the vector group, 4/6 in the vector group treated with cisplatin, 2/6 in the MUC14-OE group, and 1/6 in the MUC14-OE group treated with cisplatin (Fig. 8B). Tumor diameters across all four groups had statistically significant differences (P = 0.0091, Fig. 8C). Furthermore, histopathological examination of livers and kidneys from randomly selected mice across all groups revealed no necrosis, or fibrosis (Supplementary Fig. 1E), confirming the absence of cisplatin-induced hepatorenal toxicity. A schematic diagram summarized the molecular mechanisms: MUC14 suppresses LUAD progression by antagonizing integrin α8β6-mediated activation of PI3K/AKT and MAPK/ERK oncogenic signaling pathways. Besides, MUC14 reprograms the tumor immune microenvironment by promoting CD8 + T cell infiltration and potentiates cisplatin efficacy through activation of the JNK/c-Jun pathway (Fig. 8D).
MUC14 enhancing cisplatin response in vivo. (A) Schematic representation of tail vein xenograft model and cisplatin treatment. (B) Pathological lesions in HE-stained sections of entire lung lobes from BALB/c nude mice (four groups, n = 6 per group) are highlighted by triangular arrows. (C) Quantification and statistical analysis of microscopically measured lesion diameter. (D) Schematic diagram of the molecular mechanism of MUC14-Integrin α8β6 mediated regulation of PI3K/AKT, MAPK/ERK and JNK/c-Jun signaling pathways.
Discussion
This study establishes MUC14 as a pivotal tumor suppressor in LUAD. Its downregulation correlates with poor patient prognosis, and we identify the functional suppression of the integrin α8β6 axis as the core mechanism for its anti-tumor effects. While prior work highlighted integrins in NSCLC progression13,26,27,28,29,30,31 and fibroblast-derived MUC149, we demonstrate a novel, tumor-intrinsic role for MUC14 in inhibiting integrin α8β6 clustering. This inhibition concurrently attenuates key oncogenic pathways (PI3K/AKT, MAPK/ERK)29,30,31, suppresses the EMT phenotype, and enhances cisplatin sensitivity via JNK-mediated apoptosis32,33,34,35,36. Furthermore, it is associated with increased CD8 + T cell infiltration.
The convergence of enhanced chemosensitivity and immunomodulation confers profound therapeutic significance. This dual action directly addresses two core limitations in NSCLC: intrinsic chemoresistance and the restricted efficacy of immunotherapy due to inadequate T-cell infiltration37,38. By simultaneously increasing tumor cell vulnerability and fostering an immune-permissive microenvironment, MUC14 activation provides a rationale for synergistic chemo-immunotherapy. This adds an integrin-focused dimension to the concept that targeting tumor-intrinsic pathways can reprogram the tumor immune microenvironment, akin to strategies modulating lactate metabolism39 or utilizing non-coding RNAs40. Recent combinatorial successes support this rationale; for instance, ATM inhibition in colorectal cancer enhanced radiosensitivity, activated STING, increased CD8 + T cell infiltration, and synergized with anti-PD-L1 therapy41. Similarly, targeting the MUC14-integrin axis may disrupt both tumor viability and immune evasion. Conversely, MUC14 loss might sustain a treatment-resistant, immunosuppressive state, potentially involving mechanisms like altered macrophage metabolism observed in other contexts42.
Investigations into MUC14 in cancer remain preliminary. It has been proposed as a candidate prognostic biomarker in gastric, hepatic, and renal cancers43,44,45. Notably, the tumor-suppressive role we report contrasts with a reported oncogenic function in colorectal cancer7, underscoring the context-dependent duality of molecular signals, which necessitates interpretation within specific tissue environments.
In summary, MUC14 emerges as a valuable clinical biomarker. High expression identifies tumors with a favorable prognosis, potentially responsive to standard therapies, while low expression characterizes an aggressive, treatment-resistant subtype requiring combinatorial strategies targeting the MUC14-integrin α8β6 axis. We acknowledge that a limitation of this study is the lack of validation in an independent cohort of primary LUAD tissues. Future validation with clinical samples is needed to strengthen the translational impact of our results. Therefore, future research must first correlate MUC14 expression directly with patient survival, immune infiltration, and chemotherapy response in clinical specimens. Subsequent efforts should then validate the integrin-immune crosstalk in vivo, decipher the precise mechanistic interactions, and prospectively evaluate MUC14’s predictive value in therapeutic trials to advance targeted strategies for MUC14-low tumors.
Conclusion
Our study establishes MUC14 as a master regulatory hub in LUAD, deploying a coordinated tumor-suppressive program that concurrently disrupts integrin α8β6-driven PI3K/AKT-MAPK/ERK oncogenic signaling, reprograms immunosuppressive landscapes via CD8+ T cell recruitment, and overcomes cisplatin resistance through JNK/c-Jun-mediated apoptosis potentiation. This functional convergence—spanning cell-intrinsic pathway blockade, immune contexture remodeling, and chemosensitization—defines a unified defense framework against LUAD pathogenesis. Therapeutically, MUC14 activation represents a transformative axis for precision combination strategies, simultaneously targeting tumor plasticity, dismantling immune evasion barriers, and reversing chemoresistance to pioneer integrated therapeutic paradigms for advanced disease.
Data availability
The datasets generated and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Xu, J. Y. et al. Integrative proteomic characterization of human lung adenocarcinoma. Cell. 182 (1), 245–261e17 (2020).
Potter, A. L. et al. Recurrence after complete resection for non-small cell lung cancer in the national lung screening trial. Ann. Thorac. Surg. 116 (4), 684–692 (2023).
Suzuki, T. et al. Characterization of the nucleotide excision repair pathway and evaluation of compounds for overcoming the cisplatin resistance of non-small cell lung cancer cell lines. Oncol. Rep. 47 (4), – (2022).
Yu, X. et al. Regulation of cisplatin resistance in lung cancer by epigenetic mechanisms. Clin. Epigenet. 17 (1), 145 (2025).
Zhang, G., Yang, X. & Gao, R. Research progress on the structure and function of Endomucin. Anim. Model. Exp. Med. 3 (4), 325–329 (2020).
Li, X. et al. Understanding of endomucin: a multifaceted glycoprotein functionality in vascular inflammatory-related diseases, bone diseases and cancers. Adv. Biol. (Weinh). 8 (10), e2400061 (2024).
Huang, Q. et al. Tumor-derived Endomucin promotes colorectal cancer proliferation and metastasis. Cancer Med. 12 (3), 3222–3236 (2023).
Zhang, G. et al. Loss of endothelial EMCN drives tumor lung metastasis through the premetastatic niche. J. Transl. Med. 20 (1), 446 (2022).
Kinoshita, M. et al. Identification of human endomucin-1 and – 2 as membrane-bound O-sialoglycoproteins with anti-adhesive activity. FEBS Lett. 499 (1–2), 121–126 (2001).
Liddington, R. C. Structural aspects of integrins. Adv. Exp. Med. Biol. 819 (null), 111–126 (2014).
Humphries, J. D., Byron, A. & Humphries, M. J. Integrin ligands at a glance. J. Cell. Sci. 119 (Pt 19), 3901–3903 (2006).
Soe, Z. Y., Park, E. J. & Shimaoka, M. Integrin regulation in immunological and cancerous cells and exosomes. Int. J. Mol. Sci. 22 (4), null (2021).
Ka, M. et al. Integrin-α5 expression and its role in non-small cell lung cancer progression. Cancer Sci. 116 (2), 406–419 (2025).
Kim, S. et al. The interplay of cancer-associated fibroblasts and apoptotic cancer cells suppresses lung cancer cell growth through WISP-1-integrin ανβ3-STAT1 signaling pathway. Cell. Commun. Signal. 23 (1), 98 (2025).
Huang, W. et al. Exosomes with low miR-34c-3p expression promote invasion and migration of non-small cell lung cancer by upregulating integrin α2β1. Signal. Transduct. Tar. 5 (1), 39 (2020).
Li, J. et al. USP51/PD-L1/ITGB1-deployed juxtacrine interaction plays a cell-intrinsic role in promoting chemoresistant phenotypes in non-small cell lung cancer. Cancer Commun. (Lond). 43 (7), 765–787 (2023).
Tang, Z. et al. GEPIA: a web server for cancer and normal gene expression profiling and interactive analyses. Nucleic Acids Res. 45 (W1), W98–w102 (2017).
Li, T. et al. TIMER: A web server for comprehensive analysis of Tumor-Infiltrating immune cells. Cancer Res. 77 (21), e108–e110 (2017).
Yang, W. et al. Genomics of drug sensitivity in cancer (GDSC): a resource for therapeutic biomarker discovery in cancer cells. Nucleic Acids Res. 41 (Database issue), D955–D961 (2013).
Wang, Z. et al. Tumor immune microenvironment analysis in different pathologic responses to neoadjuvant immunotherapy combined with chemotherapy in non-small cell lung cancer. Transl. Lung Cancer Res. 14 (9), 3975–3987 (2025).
Phan, T. G. et al. Chemotherapy and the extra-tumor immune microenvironment: EXTRA-TIME. Cancer Discov. 14 (4), 643–647 (2024).
Lian, Q. et al. Enhancing radiosensitivity of osteosarcoma by ITGB3 knockdown: a mechanism linked to enhanced osteogenic differentiation status through JNK/c-JUN/RUNX2 pathway activation. J. Exp. Clin. Cancer Res. 44 (1), 159 (2025).
Zhan, Z. et al. Chronic alcohol consumption aggravates acute kidney injury through integrin β1/JNK signaling. Redox Biol. 77, 103386 (2024).
Zhang, Y. et al. RIPK1 contributes to cisplatin-induced apoptosis of esophageal squamous cell carcinoma cells via activation of JNK pathway. Life Sci. 269, 119064 (2021).
Hong, L. et al. Isodeoxyelephantopin inactivates thioredoxin reductase 1 and activates ROS-Mediated JNK signaling pathway to exacerbate cisplatin effectiveness in human colon cancer cells. Front. Cell. Dev. Biol. 8, 580517 (2020).
Ando, T. et al. Integrin α11 in non-small cell lung cancer is associated with tumor progression and postoperative recurrence. Cancer Sci. 111 (1), 200–208 (2020).
Matsumoto, Y. et al. Integrin alpha 2 is associated with tumor progression and postoperative recurrence in non-small cell lung cancer. Jpn. J. Clin. Oncol. 53 (1), 63–73 (2023).
Yasuda, M. et al. Stimulation of beta1 integrin down-regulates ICAM-1 expression and ICAM-1-dependent adhesion of lung cancer cells through focal adhesion kinase. Cancer Res. 61 (5), 2022–2030 (2001).
Wu, J. L. et al. Fibronectin promotes tumor progression through integrin αvβ3/PI3K/AKT/SOX2 signaling in non-small cell lung cancer. Heliyon. 9 (9), e20185 (2023).
Zhang, M. et al. Lipopolysaccharide and lipoteichoic acid regulate the PI3K/AKT pathway through osteopontin/integrin β3 to promote malignant progression of non-small cell lung cancer. J. Thorac. Dis. 15 (1), 168–185 (2023).
Yan, P. et al. Integrin αvβ6 promotes lung cancer proliferation and metastasis through upregulation of IL-8-Mediated MAPK/ERK signaling. Transl. Oncol. 11 (3), 619–627 (2018).
Hayakawa, J. et al. The activation of c-Jun NH2-terminal kinase (JNK) by DNA-damaging agents serves to promote drug resistance via activating transcription factor 2 (ATF2)-dependent enhanced DNA repair. J. Biol. Chem. 278 (23), 20582–20592 (2003).
Hayakawa, J. et al. Identification of promoters bound by c-Jun/ATF2 during rapid large-scale gene activation following genotoxic stress. Mol. cell. 16 (4), 521–535 (2004).
Zhang, Y. et al. RIPK1 contributes to cisplatin-induced apoptosis of esophageal squamous cell carcinoma cells via activation of JNK pathway. Life Sci. 269, 119064 (2021).
Hong, L. et al. Isodeoxyelephantopin inactivates thioredoxin reductase 1 and activates ROS-mediated JNK signaling pathway to exacerbate cisplatin effectiveness in human colon cancer cells. Front. Cell. Dev. Biol. 8, 580517 (2020).
Mansouri, A. et al. Sustained activation of JNK/p38 MAPK pathways in response to cisplatin leads to Fas ligand induction and cell death in ovarian carcinoma cells. J. Biol. Chem. 278 (21), 19245–19256 (2003).
Wei, W. & Su, Y. Function of CD8+, conventional CD4+, and regulatory CD4 + T cell identification in lung cancer. Comput. Biol. Med. 160, 106933 (2023).
Dasari, S. & Tchounwou, P. B. Cisplatin in cancer therapy: molecular mechanisms of action. Eur. J. Pharmacol. 740, 364–378 (2014).
Cui, Y. et al. Baicalin attenuates the immune escape of oral squamous cell carcinoma by reducing lactate accumulation in tumor microenvironment. J. Adv. Res. 77, 721–732 (2025).
Du, W. et al. Regulation of tumor metastasis and CD8 + T cells infiltration by circRNF216/miR-576-5p/ZC3H12C axis in colorectal cancer. Cell. Mol. Biol. Lett. 29 (1), 19 (2024).
Xie, Y. et al. Targeting ATM enhances radiation sensitivity of colorectal cancer by potentiating radiation-induced cell death and antitumor immunity. J. Adv. Res. 74, 513–530 (2025).
Qin, H. et al. The fatty acid receptor CD36 promotes macrophage infiltration via p110γ signaling to stimulate metastasis. J. Adv. Res. 74, 237–253 (2025).
Zhang, Q. et al. Weighted correlation gene network analysis reveals a new stemness index-related survival model for prognostic prediction in hepatocellular carcinoma. Aging (Albany NY). 12 (13), 13502–13517 (2020).
Dai, W. et al. Systematical analysis of the cancer genome atlas database reveals EMCN/MUC15 combination as a prognostic signature for gastric cancer. Front. Mol. Biosci. 7, 19 (2020).
Meng, H. et al. A mucins expression signature impacts overall survival in patients with clear cell renal cell carcinoma. Cancer Med. 10 (17), 5823–5838 (2021).
Acknowledgements
We thank all who help us in writing the manuscript.
Funding
This study was supported by 2023 Basic scientific research expenses for scientific research projects of universities of Heilongjiang province (No.2023-KYYWF-0182).
Author information
Authors and Affiliations
Contributions
X.L. performed the literature search, in vitro experiments and drafted the manuscript. S.H. was responsible for the in vivo studies. M.L., Z.Z. and C.X. assisted with specific in vitro assays. S.Y. and G.H. designed concept of the study and revised the whole manuscript. All authors have read and agreed to the published version of the manuscript.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
This study was approved by the Medical Ethics Committee of the Second Affiliated Hospital of Harbin Medical University (YJSKY2023-264). Informed consent was obtained from all individual participants and/or their legal guardians. All procedures followed were in accordance with the ethical standards approved by the Institutional Ethics Committee on human experimentation and with the Helsinki Declaration. The animal study protocol (YJSDW2023-233) was approved by the same Medical Ethics Committee of the Second Affiliated Hospital of Harbin Medical University, and was conducted in accordance with the National Institutes of Health guide for the care and use of Laboratory animals. This study is reported in accordance with the ARRIVE guidelines 2.0.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Li, X., Li, M., Huang, S. et al. MUC14 suppresses lung adenocarcinoma via integrin α8β6/PI3K/AKT/MAPK modulating cisplatin response and immunity. Sci Rep 16, 7784 (2026). https://doi.org/10.1038/s41598-026-39019-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-026-39019-z