Abstract
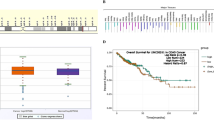
Triggering PI3K signaling cascade is most common in human cancers. This study aimed to screen the association of seven SNPs and two hotspot mutations, clinical outcome and therapeutic potential with susceptibility to colorectal carcinoma (CRC). In this case-control study of 495 CRC and 495 controls, seven SNPs were genotyped by ARMS-PCR. The mutant allele frequencies were significantly higher for five SNPs, E542K and E17K but lower for rs6443624 (P = 0.1480) and rs1883965 (P = 0.4105). DNA sequencing results revealed point mutations were more prevalent in CRC. Pairwise linkage disequilibrium analysis was performed and strong LD was observed between rs6443624 and rs10138227 (D’ = 0.953, r² = 0.435, LOD = 62.61) among CRC. Pearson’s chi-square test showed these variants were associated with CRC risk (P < 0.001). Kaplan Meier analysis revealed that six SNPs (P < 0.005) were strongly associated with clinicopathological parameters and drugs with OS in CRC but not for rs6443624 (P = 0.3). Breslow and Tarone-Ware Test showed significant survival differences but capecitabine revealed the highest survival estimates. Swiss model was used to analyze the changes in protein structure. QMEAND value for E542K is 0.88 ± 0.05 and for E17K is 0.78 ± 0.05 representing high quality structure, however, Ramachandaran structure 97.37% favoured for E542K and 95.23% for E17K. The current study suggests that PI3K pathway variants may be associated with colorectal cancer susceptibility and highlights their potential relevance as candidate biomarkers, pending further validation.
Similar content being viewed by others
Data availability
Data is provided in the supplementary information file.
References
Xi, Y. & Xu, P. Global colorectal cancer burden in 2020 and projections to 2040. Translational Oncol. 14 (10), 101174. https://doi.org/10.1016/j.tranon.2021.101174 (2021).
Morgan, E. et al. Global burden of colorectal cancer in 2020 and 2040: Incidence and mortality estimates from GLOBOCAN. Gut 72 (2), 338–344. https://doi.org/10.1136/gutjnl-2022-327736 (2023).
Keum, N. & Giovannucci, E. Global burden of colorectal cancer: emerging trends, risk factors and prevention strategies. Nat. Reviews Gastroenterol. Hepatol. 16 (12), 713–732. https://doi.org/10.1038/s41575-019-0189-8 (2019).
Johnson, S. M. et al. Novel expression patterns of PI3K/Akt/mTOR signaling pathway components in colorectal cancer. J. Am. Coll. Surg. 210 (5), 767–776. https://doi.org/10.1016/j.jamcollsurg.2009.12.008 (2010).
Muhammad, S. S., Shoaib, M. & Pervez, M. T. An Integrated Framework for Analysis and Prediction of Impact of Single Nucleotide Polymorphism Associated with Human Diseases. Evolutionary Bioinf. 20 (5), 1–14. https://doi.org/10.1177/11769343241249916 (2024).
Ge, Y. et al. Genetic variants in PI3K/Akt/mTOR pathway genes contribute to gastric cancer risk. Gene 670, 130–135. https://doi.org/10.1016/j.gene.2018.05.093 (2018).
Simons, C. C. et al. Polymorphisms in the mTOR-PI3K-Akt pathway, energy balance-related exposures and colorectal cancer risk in the Netherlands Cohort Study. BioData Min. 15, 1–20. https://doi.org/10.1186/s13040-021-00286-3 (2022).
Ertay, A. Altered PI3K/AKT/mTOR Signaling Pathway and Cancer Stem Cells. In Cancer Stem Cells and Cancer Therapy 131–158 (Springer Nature Switzerland, 2025). https://doi.org/10.1007/978-3-031-74842-4_5.
Mirzapoor Abbasabadi, Z. et al. KRAS, NRAS, BRAF, and PIK3CA mutation rates, clinicopathological association, and their prognostic value in Iranian colorectal cancer patients. J. Clin. Lab. Anal. 37 (5), 1–10. https://doi.org/10.1002/jcla.24868 (2023).
Hechtman, J. F. et al. AKT1 E17K in colorectal carcinoma is associated with BRAF V600E but not MSI-H status: a clinicopathologic comparison to PIK3CA helical and kinase domain mutants. Mol. Cancer Res. 13 (6), 1003–1008. https://doi.org/10.1158/1541-7786.MCR-15-0062-T (2015).
Chuang, J. et al. MAP2K1 Mutations in Advanced Colorectal Cancer Predict Poor Response to Anti-EGFR Therapy and to Vertical Targeting of MAPK Pathway. Clin. Colorectal Cancer. 20 (1), 72–78. https://doi.org/10.1016/j.clcc.2020.12.003 (2021).
Murugan, A. K., Alzahrani, A. & Xing, M. Mutations in critical domains confer the human mTOR gene strong tumorigenicity. J. Biol. Chem. 288 (9), 6511–6521. https://doi.org/10.1074/jbc.M112.399485 (2013).
Bando, H., Ohtsu, A. & Yoshino, T. Therapeutic landscape and future direction of metastatic colorectal cancer. Nat. Reviews Gastroenterol. Hepatol. 20 (5), 306–322. https://doi.org/10.1038/s41575-022-00736-1 (2023).
Domingo, E. et al. Evaluation of PIK3CA mutation as a predictor of benefit from nonsteroidal anti-inflammatory drug therapy in colorectal cancer. J. Clin. Oncol. 31 (34), 4297–4305. https://doi.org/10.1200/JCO.2013.50.0322 (2013).
Bahrami, A. et al. Therapeutic potential of targeting PI3K/AKT pathway in treatment of colorectal cancer: rational and progress. J. Cell. Biochem. 119 (3), 2460–2469. https://doi.org/10.1002/jcb.25950I (2018).
Iqbal, M. J., Shakoori, F. R., Muneer, B. & Shakoori, A. R. Mutation Profiling of PI3K/AKT1/MTOR Pathway Genes in Breast Cancer Patients of Pakistan. Pakistan J. Zool. 56 (1), 273. https://doi.org/10.17582/journal.pjz/20221006131025 (2024).
Tufail, M. & Wu, C. Cancer statistics in Pakistan from 1994 to 2021: data from cancer registry. JCO Clin. Cancer Inf. 7, e2200142. https://doi.org/10.1200/CCI.22.00142 (2023).
Dairawan, M. & Shetty, P. J. The evolution of DNA extraction methods. Am. J. Biomedical Sci. Res. 8 (1), 39–45. https://doi.org/10.34297/AJBSR.2020.08.001234 (2020).
Ahlawat, S., Sharma, R., Maitra, A., Roy, M. & Tantia, M. S. Designing, optimization and validation of tetra-primer ARMS PCR protocol for genotyping mutations in caprine Fec genes. Meta Gene. 2, 439–449. https://doi.org/10.1016/j.mgene.2014.05.004 (2014).
Caetano-Anolles, D. Polymerase Chain Reaction, Editor(s): Stanley Maloy, Kelly Hughes, Brenner’s Encyclopedia of Genetics (Second Edition), Academic Press, 392–395. (2013).
Li, Q. et al. Signaling pathways involved in colorectal cancer: pathogenesis and targeted therapy. Signal. Transduct. Target. Therapy. 9 (1), 1–48. https://doi.org/10.1038/s41392-024-01953-7 (2024).
Quaye, L. et al. Tagging single-nucleotide polymorphisms in candidate oncogenes and susceptibility to ovarian cancer. Br. J. Cancer. 100 (6), 993–1001. https://doi.org/10.1038/sj.bjc.6604947 (2009). Ovarian Cancer Association Consortium, Easton,.
Wang, Y., Zhang, H., Lin, M. & Wang, Y. Association of FGFR2 and PI3KCA genetic variants with the risk of breast cancer in a Chinese population. Cancer Manage. Res. 10 (5), 1305–1311. https://doi.org/10.2147/CMAR.S164084 (2018).
Slattery, M. L. et al. Genetic variation in a metabolic signaling pathway and colon and rectal cancer risk: mTOR, PTEN, STK11, RPKAA1, PRKAG2, TSC1, TSC2, PI3K and Akt1. Carcinogenesis 31 (9), 1604–1611. https://doi.org/10.1093/carcin/bgq142 (2010).
Kim, J., Yum, S., Kang, C. & Kang, S. J. Gene-gene interactions in gastrointestinal cancer susceptibility. Oncotarget 7 (41), 67612–67625. https://doi.org/10.18632/oncotarget.11701 (2016).
Bizhani, F. et al. Association between single nucleotide polymorphisms in the PI3K/AKT/mTOR pathway and bladder cancer risk in a sample of Iranian population. EXCLI J. 17, 3–13. https://doi.org/10.17179/excli2017-329 (2018).
Ulu, E., Yaylım, İ., Arıkan, S. & Cacına, C. The evaluation of PIK3CA gene variation and serum PI3K level in breast cancer risk and prognosis in Turkish population. Turkish J. Biochem. 47 (3), 317–324. https://doi.org/10.1515/tjb-2021-0072 (2022).
Allam, L. et al. AKT1 Polymorphism (rs10138227) and Risk of Colorectal Cancer in Moroccan Population: A Case Control Study. Asian Pac. J. cancer prevention: APJCP. 21 (11), 3165–3170. https://doi.org/10.31557/APJCP.2020.21.11.3165 (2020).
Wang, M. Y. et al. A Functional Polymorphism (rs2494752) in the AKT1 Promoter Region and Gastric Adenocarcinoma Risk in an Eastern Chinese Population. Sci. Rep. 6 (1), 1–8. https://doi.org/10.1038/srep20008 (2016).
Vuorinen, S. I. et al. SDC4-rs1981429 and ATM-rs228590 may provide early biomarkers of breast cancer risk. J. Cancer Res. Clin. Oncol. 149 (8), 4563–4578. https://doi.org/10.1007/s00432-022-04236-2 (2023).
Zubair, H., Khan, Z. & Imran, M. Impact of AKT1 polymorphism on DNA damage, BTG2 expression, and risk of colorectal cancer development. Radiol. Oncol. 56 (3), 336–345. https://doi.org/10.2478/raon-2022-0031 (2022).
Pérez-Ramírez, C., Cañadas-Garre, M., Molina, M. Á., Barrera, C., Faus-Dáder, M. J. & J., & Impact of single nucleotide polymorphisms on the efficacy and toxicity of EGFR tyrosine kinase inhibitors in advanced non-small cell lung cancer patients. Mutat. Res. Reviews Mutat. Res. 781, 63–70. https://doi.org/10.1016/j.mrrev.2019.04.001 (2019).
Li, Y. et al. Association of inflammation-related gene polymorphisms with susceptibility and radiotherapy sensitivity in head and neck squamous cell carcinoma patients in northeast China. Front. Oncol. 11 (6), 1–25. https://doi.org/10.3389/fonc.2021.651632 (2021).
Kim, J. G. et al. Prostaglandin synthase 2/cyclooxygenase 2 (PTGS2/COX2) 8473T > C polymorphism associated with prognosis for patients with colorectal cancer treated with capecitabine and oxaliplatin. Cancer Chemother. Pharmacol. 64, 953–960. https://doi.org/10.1007/s00280-009-0947-3 (2009).
Ilozumba, M. N. et al. mTOR pathway candidate genes and energy intake interaction on breast cancer risk in Black women from the Women’s Circle of Health Study. Eur. J. Nutr. 62 (6), 2593–2604. https://doi.org/10.1007/s00394-023-03176-y (2023).
Piao, Y. et al. Association of MTOR and AKT gene polymorphisms with susceptibility and survival of gastric cancer. PloS one. 10 (8), 1–19. https://doi.org/10.1371/journal.pone.0136447 (2015).
He, J. et al. Genetic variations of mTORC1 genes and risk of gastric cancer in an Eastern Chinese population. Mol. Carcinog. 52 (S1), 70–79. https://doi.org/10.1002/mc.22013 (2013).
Zhu, J. et al. Associations of PI3KR1 and mTOR polymorphisms with esophageal squamous cell carcinoma risk and gene-environment interactions in Eastern Chinese populations. Sci. Rep. 5 (1), 1–11. https://doi.org/10.1038/srep08250 (2015).
Devaney, J. M. et al. AKT1 polymorphisms are associated with risk for metabolic syndrome. Hum. Genet. 129 (2), 129–139. https://doi.org/10.1007/s00439-010-0910-8 (2011).
Gibbs, R. A. et al. The international HapMap project. Nature 426, 789–796. https://doi.org/10.1038/nature02168 (2003).
Acknowledgements
We gratefully acknowledge Fatima Jinnah Women University Rawalpindi and healthy controls/patients for their contribution to this research. We are also grateful to Benazir Bhutto, Holy Family, District Headquarter and PIMS (Pakistan) hospitals for their support and cooperation.
Funding
This research did not receive any specific grant from funding agencies in the public, commercial, or not-for-profit sectors.
Author information
Authors and Affiliations
Contributions
HP contributed to conceptualizing, methodology, writing-original draft, visualization. NM contributed to supervision, validation, writing-review and editing. PAM, JSK, MAK and HM contributed to resources. All authors have read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
This study was approved by ethical committee of Fatima Jinnah Women University Rawalpindi (Approval No: FJWU/EC/2024/102, Date: 05-01-2025), Rawalpindi Medical University RMU (Holy Family, Benazir Bhutto, District Headquarter hospitals; Approval No: Ref. No. 531/IREF/RMU/2023, Date: 30-9-2023) and Shaheed Zulfiqar Ali Bhutto Medical University, PIMS, Islamabad (Approval No: F. 1–1/2015/ERB/SZABMU/1203, Date: 30-11-2023). All participants provided their written informed consent before enrollment in this research, and all the methods are carried out in accordance with World Medical Association Declaration of Helsinki.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Pervaiz, H., Masood, N., Malik, P.A. et al. Genetic analysis of key players in PI3K signaling cascade of colorectal carcinoma. Sci Rep (2026). https://doi.org/10.1038/s41598-026-42006-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-026-42006-z