Abstract
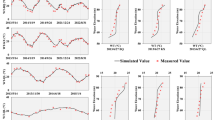
Accurate monitoring and prediction of Chlorophyll-a (Chl-a) concentrations are critical for protecting desalination systems from algal blooms. This study presents an advanced framework employing the Graph Neural Network Transformer Model (GNN-T) to predict the spatiotemporal dynamics of Chl-a in semi-enclosed marine environments, such as the Persian Gulf, a key region for desalination facilities in the Middle East. The GNN-T model integrates critical environmental variables, utilizing MODIS/Aqua and ERA5 datasets with 300,000 observations. Demonstrating robust generalizability, the test model achieves an R² of 0.906 in the Gulf of Mexico, outperforming conventional deep learning approaches, including CNN-LSTM, BiLSTM, Temporal-Relational GNN, and AGTCNSD. Statistical error metrics confirm the GNN-T’s superior predictive accuracy and lower error rates. Global sensitivity and uncertainty analysis (GSUA) highlights sea surface temperature, normalized fluorescence line height, and particulate organic carbon as key drivers. Convergent Cross-Mapping (CCM) elucidates nonlinear causal relationships, distinguishing correlation from mechanistic causality. Additionally, a causality-driven ablation study, guided by CCM and Sobol sensitivity analyses, streamlined the model by selecting the top 13 influential variables, achieving a test R² of 0.882 with 25% reduced computational costs, enhancing operational efficiency without substantial loss in predictive accuracy. Uncertainty quantification, performed using Monte Carlo dropout, provides 95% confidence intervals. Quartile analysis establishes bloom thresholds at the 50th (bloom), 75th (intense bloom), and 90th percentiles (extreme bloom) for probabilistic risk assessments. This model serves as an effective operational tool for detecting algal bloom onset and mitigating associated economic impacts.
Similar content being viewed by others
Data availability
The datasets used or analysed during the current study available from the corresponding authors on reasonable request.
Abbreviations
- Chl-a:
-
Chlorophyll-a
- HABs:
-
Harmful Algal Blooms
- GNN-T:
-
Graph Neural Network Transformer
- MODIS:
-
Moderate Resolution Imaging Spectroradiometer
- ERA5:
-
ECMWF Reanalysis v5
- AGTCNSD:
-
Adaptive Graph Convolutional Network with Series Decomposition
- CCM:
-
Convergent Cross-Mapping
- ROMS:
-
Regional Ocean Modeling System
- Variables:
-
(In Table 1, e.g., NFLH, POC, SST, etc.)
- \(\varvec{\rho}\) :
-
Correlation
References
Chen, J. et al. Common fate of sister lakes in Hulunbuir Grassland: Long-term harmful algal bloom crisis from multi-source remote sensing insights. J. Hydrol. 594, 125970. https://doi.org/10.1016/J.JHYDROL.2021.125970 (2021).
Yu, H. Climate change unveils hidden microbial dangers. Environ. Sci. Ecotechnology. 24, 100544. https://doi.org/10.1016/J.ESE.2025.100544 (2025).
Cho, H., Choi, U. J. & Park, H. Deep learning application to time-series prediction of daily chlorophyll-a concentration. WIT Trans. Ecol. Environ. 215, 157–163 (2018).
Dwivedi, V. P. & Bresson, X. A generalization of transformer networks to graphs. ArXiv Preprint ArXiv:2012.09699. (2020).
Ying, C. et al. Do transformers really perform badly for graph representation? Adv. Neural. Inf. Process. Syst. 34, 28877–28888 (2021).
Barzegar, R., Aalami, M. T. & Adamowski, J. Short-term water quality variable prediction using a hybrid CNN–LSTM deep learning model. Stoch. Env. Res. Risk Assess. 34 (2), 415–433 (2020).
Hu, C., Lee, Z. & Franz, B. Chlorophyll aalgorithms for oligotrophic oceans: A novel approach based on three-band reflectance difference. J. Geophys. Research: Oceans, 117(C1), C01011. https://doi.org/10.1029/2011JC007395 (2012).
Zarbipour, P., Nikoo, M. R., Akbari, H., Nazari, R. & Karimi, M. Bridging Causality and Deep Learning for Harmful Algal Bloom Prediction. Water Res. 125492. https://doi.org/10.1016/j.watres.2026.125492 (2026b).
Huang, H. & Zhang, J. Prediction of chlorophyll a and risk assessment of water blooms in Poyang Lake based on a machine learning method. Environ. Pollut. 347, 123501. https://doi.org/10.1016/J.ENVPOL.2024.123501 (2024).
Choi, J. H., Kim, J., Won, J. & Min, O. Modelling chlorophyll-a concentration using deep neural networks considering extreme data imbalance and skewness. 2019 21st Int. Conf. Adv. Communication Technol. (ICACT). 631, 634 (2019).
Firozjaei, M. R., Hajebi, Z., Naeeni, S. T. O. & Akbari, H. Discharge performance of a submerged seawater intake in unsteady flows: Combination of physical models and decision tree algorithms. J. Water Process. Eng. 60, 105198 (2024).
Rasolpour, S. & Akbari, H. Development of desalination plants within the Semi-enclosed Persian Gulf (Applied Water Science, 2024).
Zarbipour, P. & Akbari, H. Reliability design of seawater desalination outfalls based on a novel probabilistic environmental assessment. Ocean Eng. 313, 119465. https://doi.org/10.1016/j.oceaneng.2024.119465 (2024).
Boyer, J. N., Kelble, C. R., Ortner, P. B. & Rudnick, D. T. Phytoplankton bloom status: Chlorophyll a biomass as an indicator of water quality condition in the southern estuaries of Florida, USA. Ecol. Ind. 9 (6), S56–S67. https://doi.org/10.1016/J.ECOLIND.2008.11.013 (2009).
Moloto, T. M. et al. Remote sensing of phytoplankton community composition in the northern Benguela upwelling system. Front. Mar. Sci. 10, 1118226 (2023).
Buchanan, C., Lacouture, R. V., Marshall, H. G., Olson, M. & Johnson, J. M. Phytoplankton reference communities for Chesapeake Bay and its tidal tributaries. Estuaries 28 (1), 138–159 (2005).
Moradi, M. & Moradi, N. Correlation between concentrations of chlorophyll-a and satellite derived climatic factors in the Persian Gulf. Mar. Pollut. Bull. 161, 111728 (2020).
Fu, D. et al. Vcr-graphormer: A mini-batch graph transformer via virtual connections. The Twelfth International Conference on Learning Representations. (2024).
Sugihara, G. et al. Detecting Causality in Complex Ecosystems. Science 338 (6106), 496–500. https://doi.org/10.1126/science.1227079 (2012).
Ghaemi, M., Abtahi, B. & Gholamipour, S. Spatial distribution of nutrients and chlorophyll a across the Persian Gulf and the Gulf of Oman. Ocean. Coastal. Manage. 201, 105476. https://doi.org/10.1016/j.ocecoaman.2020.105476 (2021).
Al-Yamani, F., Saburova, M. & Polikarpov, I. A preliminary assessment of harmful algal blooms in Kuwait’s marine environment. Aquat. Ecosyst. Health Manag. 15 (sup1), 64–72 (2012).
Wang, H., Galbraith, E. & Convertino, M. Algal bloom ties: Spreading network inference and extreme eco-environmental feedback. Entropy 25 (4), 636 (2023).
Al-Wardy, M., Zarei, E., Nikoo, M. R. & Nazari, R. Climate-driven projections of cyanobacterial harmful algal bloom expansion in coastal waters. Sci. Total Environ. 992, 179940. https://doi.org/10.1016/J.SCITOTENV.2025.179940 (2025).
Raulo, S. et al. Determining chlorophyll-a thresholds for characterizing algal bloom conditions: An ocean colour remote sensing approach. Sci. Total Environ. 961, 178353. https://doi.org/10.1016/J.SCITOTENV.2024.178353 (2025).
Yang, M. et al. Two-decade variability and trend of chlorophyll-a in the Arabian Sea and Persian Gulf based on reconstructed satellite data. Front. Mar. Sci. 11, 1520775 (2024).
Ying, C., Xiao, L., Xueliang, Z., Wenyang, S. & Chongxuan, X. Marine chlorophyll-a prediction based on deep auto-encoded temporal convolutional network model. Ocean Model. 186, 102263 (2023).
Cen, H., Han, G., Lin, X., Liu, Y. & Zhang, H. Deep learning-based forecasting model for chlorophyll-a response to tropical cyclones in the Western North Pacific. Ocean Model. 189, 102345. https://doi.org/10.1016/j.ocemod.2024.102345 (2024).
Ding, W. & Li, C. Algal blooms forecasting with hybrid deep learning models from satellite data in the Zhoushan fishery. Ecol. Inf. 82, 102664. https://doi.org/10.1016/j.ecoinf.2024.102664 (2024).
Cao, H., Han, L. & Li, L. A deep learning method for cyanobacterial harmful algae blooms prediction in Taihu Lake, China. Harmful Algae. 113, 102189. https://doi.org/10.1016/j.hal.2022.102189 (2022).
Mattei, F. & Scardi, M. Mining satellite data for extracting chlorophyll a spatio-temporal patterns in the Mediterranean Sea. Environ. Model. Softw. 150, 105353. https://doi.org/10.1016/j.envsoft.2022.105353 (2022).
Abbas, A., Park, M., Baek, S. S. & Cho, K. H. Deep learning-based algorithms for long-term prediction of chlorophyll-a in catchment streams. J. Hydrol. 626, 130240. https://doi.org/10.1016/J.JHYDROL.2023.130240 (2023).
Zhou, G., Chen, J., Liu, M. & Ma, L. A spatiotemporal attention-augmented ConvLSTM model for ocean remote sensing reflectance prediction. Int. J. Appl. Earth Obs. Geoinf. 129, 103815. https://doi.org/10.1016/j.jag.2024.103815 (2024).
Chen, Y. et al. Multi-step prediction of chlorophyll concentration based on adaptive graph-temporal convolutional network with series decomposition. Meas. Sci. Technol. 35 (3), 035801 (2023).
Zhu, Q. et al. Temporal-relational graph neural network for nearshore seawater quality parameters multivariate multi-step prediction and correlation modelling. Expert Syst. Appl. 265, 126020 (2025).
Qian, J. et al. Identification of driving factors of algal growth in the South-to-North Water Diversion Project by Transformer-based deep learning. Water Biology Secur. 2 (3), 100184. https://doi.org/10.1016/j.watbs.2023.100184 (2023).
Kim, K. M. & Ahn, J. H. Machine learning predictions of chlorophyll-a in the Han river basin, Korea. J. Environ. Manage. 318, 115636. https://doi.org/10.1016/j.jenvman.2022.115636 (2022).
Fennel, K. et al. Advancing marine biogeochemical and ecosystem reanalyses and forecasts as tools for monitoring and managing ecosystem health. Front. Mar. Sci. 6, 89 (2019).
Zibordi, G., Voss, K. J., Johnson, B. C. & Mueller, J. L. IOCCG Ocean optics and biogeochemistry protocols for satellite ocean colour sensor validation. IOCCG Protocols Series, 3, 67. https://doi.org/10.25607/OBP-691 (2019).
Zarbipour, P., Nikoo, H. & Akbari, &, M. R Quantum machine learning for wave overtopping estimation: Integrating with causal inference and uncertainty quantification. Ocean. Eng. 343, 123482. https://doi.org/10.1016/j.oceaneng.2025.123482 (2026a).
Author information
Authors and Affiliations
Contributions
Pouya Zarbipour and Hassan Akbari defined the methodology and concept of the study. Pouya Zarbipour and Atefe Kazemi Choolanak done Data Curation and Pouya Zarbipour done Formal Analysis under the supervision of Hassan Akbari and Mohammad Reza Nikoo. He wrote the original draft of the manuscript, and all the authors reviewed and revised the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zarbipour, P., Akbari, H., Nikoo, M.R. et al. Spatiotemporal prediction of chlorophyll-a in semi-enclosed gulfs using a hybrid graph neural network-transformer framework with satellite data and causal analysis. Sci Rep (2026). https://doi.org/10.1038/s41598-026-42388-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-026-42388-0