Abstract
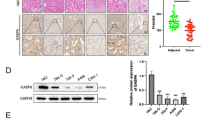
Clear cell renal cell carcinoma (ccRCC) is resistant to conventional radiotherapy and chemotherapy, creating an urgent need for novel therapeutic strategies. Although GABA transaminase (ABAT) is involved in metabolic reprogramming as reported, its precise function and molecular mechanisms in ccRCC remain unclear. Here, we demonstrate that ABAT overexpression suppresses tumor growth both in vitro and in vivo. Mechanistically, ABAT mediates the cGAS–STING signaling pathway, and interacts with protein arginine methyltransferase 5 (PRMT5), thereby enhances interferon signaling. Besides, ABAT was found to reduce the infiltration of regulatory T cells within the tumor microenvironment. Collectively, these results suggest that ABAT represents a potential therapeutic target in clear cell renal cell carcinoma.
Similar content being viewed by others
Data availability
The datasets generated and analysed during the current study are available in the NCBI Sequence Read Archive (SRR36638439, SRR36638436, SRR36638441, SRR36638438, SRR36638440, SRR36638437).
References
Huang, J. et al. A global trend analysis of kidney cancer incidence and mortality and their associations with smoking, alcohol consumption, and metabolic syndrome. Eur. Urol. Focus 8, 200. https://doi.org/10.1016/j.euf.2020.12.020 (2022).
Lindgren, D., Sjölund, J. & Axelson, H. Tracing renal cell carcinomas back to the nephron. Trends Cancer 4, 472. https://doi.org/10.1016/j.trecan.2018.05.003 (2018).
Jonasch, E., Gao, J. & Rathmell, W. K. Renal cell carcinoma. BMJ 349, g4797. https://doi.org/10.1136/bmj.g4797 (2014).
Chen, F. et al. Multilevel genomics-based taxonomy of renal cell carcinoma. Cell Rep. 14, 2476. https://doi.org/10.1016/j.celrep.2016.02.024 (2016).
Hsieh, J. J. et al. Renal cell carcinoma. Nat. Rev. Dis. Primers 3, 17009. https://doi.org/10.1038/nrdp.2017.9 (2017).
He, Y. et al. Perioperative circulating tumor DNA in colorectal liver metastases: Concordance with metastatic tissue and predictive value for tumor burden and prognosis. Cancer Manag Res 12, 1621. https://doi.org/10.2147/cmar.S240869 (2020).
Siegel, R. L., Miller, K. D. & Jemal, A. Cancer statistics, 2015. CA Cancer J. Clin. 65, 5. https://doi.org/10.3322/caac.21254 (2015).
Zhang, B. et al. B cell-derived GABA elicits IL-10(+) macrophages to limit anti-tumour immunity. Nature 599, 471. https://doi.org/10.1038/s41586-021-04082-1 (2021).
Tagore, M. et al. GABA regulates electrical activity and tumor initiation in melanoma. Cancer Discov. 13, 2270. https://doi.org/10.1158/2159-8290.Cd-23-0389 (2023).
Xie, M. et al. GABA regulates metabolic reprogramming to mediate the development of brain metastasis in non-small cell lung cancer. J. Exp. Clin. Cancer Res. 44, 61. https://doi.org/10.1186/s13046-025-03315-9 (2025).
Huang, D. et al. Cancer-cell-derived GABA promotes β-catenin-mediated tumour growth and immunosuppression. Nat. Cell Biol. 24, 230. https://doi.org/10.1038/s41556-021-00820-9 (2022).
Zhang, Q. et al. Insights and progress on the biosynthesis, metabolism, and physiological functions of gamma-aminobutyric acid (GABA): A review. PeerJ 12, e18712. https://doi.org/10.7717/peerj.18712 (2024).
Lu, J. et al. ABAT and ALDH6A1, regulated by transcription factor HNF4A, suppress tumorigenic capability in clear cell renal cell carcinoma. J. Transl. Med. 18, 101. https://doi.org/10.1186/s12967-020-02268-1 (2020).
Zhao, B. et al. Structural basis for concerted recruitment and activation of IRF-3 by innate immune adaptor proteins. Proc. Natl. Acad. Sci. U. S. A. 113, E3403. https://doi.org/10.1073/pnas.1603269113 (2016).
Zhuang, Q. et al. RNA methyltransferase FTSJ3 regulates the Type I Interferon Pathway to promote hepatocellular carcinoma immune evasion. Cancer Res. 84, 405. https://doi.org/10.1158/0008-5472.Can-23-2049 (2024).
Haag, S. M. et al. Targeting STING with covalent small-molecule inhibitors. Nature 559, 269. https://doi.org/10.1038/s41586-018-0287-8 (2018).
Rawling, D. C. et al. Small-molecule antagonists of the RIG-I innate immune receptor. ACS Chem. Biol. 15, 311. https://doi.org/10.1021/acschembio.9b00810 (2020).
Engstrom, L. D. et al. MRTX1719 is an MTA-cooperative PRMT5 inhibitor that exhibits synthetic lethality in preclinical models and patients with MTAP-deleted cancer. Cancer Discov. 13, 2412. https://doi.org/10.1158/2159-8290.Cd-23-0669 (2023).
Kalev, P. et al. MAT2A inhibition blocks the growth of MTAP-deleted cancer cells by reducing PRMT5-dependent mRNA splicing and inducing DNA damage. Cancer Cell 39, 209. https://doi.org/10.1016/j.ccell.2020.12.010 (2021).
Lacroix, M. et al. The histone-binding protein COPR5 is required for nuclear functions of the protein arginine methyltransferase PRMT5. EMBO Rep. 9, 452. https://doi.org/10.1038/embor.2008.45 (2008).
Andreu-Pérez, P. et al. Protein arginine methyltransferase 5 regulates ERK1/2 signal transduction amplitude and cell fate through CRAF. Sci. Signal. 4, ra58. https://doi.org/10.1126/scisignal.2001936 (2011).
Guderian, G. et al. RioK1, a new interactor of protein arginine methyltransferase 5 (PRMT5), competes with pICln for binding and modulates PRMT5 complex composition and substrate specificity. J. Biol. Chem. 286, 1976. https://doi.org/10.1074/jbc.M110.148486 (2011).
El-Hattab, A. W., Craigen, W. J. & Scaglia, F. Mitochondrial DNA maintenance defects. Biochim. Biophys. Acta Mol. Basis Dis. 1863, 1539. https://doi.org/10.1016/j.bbadis.2017.02.017 (2017).
Liu, H. et al. TFAM is an autophagy receptor that limits inflammation by binding to cytoplasmic mitochondrial DNA. Nat. Cell Biol. 26, 878. https://doi.org/10.1038/s41556-024-01419-6 (2024).
Lyu, Q. et al. A small proportion of X-linked genes contribute to X chromosome upregulation in early embryos via BRD4-mediated transcriptional activation. Curr. Biol. 32, 4397. https://doi.org/10.1016/j.cub.2022.08.059 (2022).
George, D. J., Lee, C. H. & Heng, D. New approaches to first-line treatment of advanced renal cell carcinoma. Ther. Adv. Med. Oncol. 13, 17588359211034708. https://doi.org/10.1177/17588359211034708 (2021).
Wang, Z. et al. PRMT5 reduces immunotherapy efficacy in triple-negative breast cancer by methylating KEAP1 and inhibiting ferroptosis. J. Immunother. Cancer. https://doi.org/10.1136/jitc-2023-006890 (2023).
Liu, A. et al. PRMT5 methylating SMAD4 activates TGF-β signaling and promotes colorectal cancer metastasis. Oncogene 42, 1572. https://doi.org/10.1038/s41388-023-02674-x (2023).
Jiang, Y. et al. PRMT5 disruption drives antitumor immunity in cervical cancer by reprogramming T cell-mediated response and regulating PD-L1 expression. Theranostics 11, 9162. https://doi.org/10.7150/thno.59605 (2021).
Yan, W. et al. PRMT5-mediated FUBP1 methylation accelerates prostate cancer progression. J. Clin. Invest. https://doi.org/10.1172/jci175023 (2024).
Frau, M., Feo, F. & Pascale, R. M. Pleiotropic effects of methionine adenosyltransferases deregulation as determinants of liver cancer progression and prognosis. J. Hepatol. 59, 830. https://doi.org/10.1016/j.jhep.2013.04.031 (2013).
Belmontes, B. et al. AMG 193, a clinical stage MTA-cooperative PRMT5 inhibitor, drives antitumor activity preclinically and in patients with MTAP-deleted cancers. Cancer Discov. 15, 139. https://doi.org/10.1158/2159-8290.Cd-24-0887 (2025).
Drizyte-Miller, K. et al. Combination of the MTA-cooperative PRMT5 inhibitor BMS-986504 and KRAS inhibitors is an effective treatment strategy for MTAP-deleted KRAS-mutant pancreatic cancer. Cancer Res. 85, 3540. https://doi.org/10.1158/0008-5472.Can-25-1507 (2025).
Rodon, J. et al. First-in-human study of AMG 193, an MTA-cooperative PRMT5 inhibitor, in patients with MTAP-deleted solid tumors: Results from phase I dose exploration. Ann. Oncol. 35, 1138. https://doi.org/10.1016/j.annonc.2024.08.2339 (2024).
Zhang, C. & Zhuang, S. The role of protein arginine methyltransferases in kidney diseases. Clin. Sci. 134, 2037. https://doi.org/10.1042/cs20200680 (2020).
Che, P. et al. Integrin αvβ3 engagement regulates glucose metabolism and migration through focal adhesion kinase (FAK) and protein arginine methyltransferase 5 (PRMT5) in glioblastoma cells. Cancers https://doi.org/10.3390/cancers13051111 (2021).
Huang, L. et al. Inhibition of protein arginine methyltransferase 5 enhances hepatic mitochondrial biogenesis. J. Biol. Chem. 293, 10884. https://doi.org/10.1074/jbc.RA118.002377 (2018).
Şenbabaoğlu, Y. et al. Tumor immune microenvironment characterization in clear cell renal cell carcinoma identifies prognostic and immunotherapeutically relevant messenger RNA signatures. Genome Biol. 17, 231. https://doi.org/10.1186/s13059-016-1092-z (2016).
Thompson, R. H. et al. PD-1 is expressed by tumor-infiltrating immune cells and is associated with poor outcome for patients with renal cell carcinoma. Clin. Cancer Res. 13, 1757. https://doi.org/10.1158/1078-0432.Ccr-06-2599 (2007).
Rathmell, W. K. et al. Management of metastatic clear cell renal cell carcinoma: ASCO guideline. J. Clin. Oncol. 40, 2957. https://doi.org/10.1200/jco.22.00868 (2022).
Stupichev, D. et al. AI-driven multimodal algorithm predicts immunotherapy and targeted therapy outcomes in clear cell renal cell carcinoma. Cell Rep. Med. 6, 102299. https://doi.org/10.1016/j.xcrm.2025.102299 (2025).
Shah, A. Y. et al. Outcomes of patients with metastatic clear-cell renal cell carcinoma treated with second-line VEGFR-TKI after first-line immune checkpoint inhibitors. Eur. J. Cancer. 114, 67. https://doi.org/10.1016/j.ejca.2019.04.003 (2019).
Li, M. et al. ID1 boosts antiviral immunity by countering PRMT5-mediated STING methylation. Cell Rep. 44, 116547. https://doi.org/10.1016/j.celrep.2025.116547 (2025).
Zhang, H. et al. Epigenetically targeting PRMT5 promotes antitumor immunity by inducing endogenous retroviruses expression and triggering viral mimicry response. Transl. Res. 281, 55. https://doi.org/10.1016/j.trsl.2025.05.007 (2025).
Kim, H. et al. PRMT5 control of cGAS/STING and NLRC5 pathways defines melanoma response to antitumor immunity. Sci. Transl. Med. https://doi.org/10.1126/scitranslmed.aaz5683 (2020).
Zhang, Q. et al. Decoding sunitinib resistance in ccRCC: Metabolic-reprogramming-induced ABAT and GABAergic system shifts. iScience 27, 110415. https://doi.org/10.1016/j.isci.2024.110415 (2024).
Funding
This study received financial support from the Ministry of Science and Technology of the People’s Republic of China (Hongzhao Li, 2022YFC3602901).
Author information
Authors and Affiliations
Contributions
Yi Feng was responsible for conceptualization, data curation, formal analysis, investigation, methodology, and writing of the original draft. Senming Cao was responsible for investigation and methodology. Tianwei Cai was responsible for formal analysis, validation, and methodology. Yin Lu was responsible for formal analysis and investigation. Bin Jiang was responsible for formal analysis and investigation. Zexuan Lv was responsible for formal analysis and investigation. Jinlu Tang was responsible for investigation. Chunyu Liu was responsible for investigation. Qi Wang was responsible for formal analysis. Ji Feng was responsible for investigation. Zheng Wang was responsible for resources. Qi Ai was responsible for resources. Xupeng Zhao was responsible for resources. KL was responsible for conceptualization and data curation. Qiang Cheng was responsible for conceptualization, data curation, and funding acquisition. Wenmei Fan was responsible for conceptualization, resources, supervision, and funding acquisition. Hongzhao Li was responsible for conceptualization, resources, data curation, formal analysis, supervision, funding acquisition, and writing of the original draft.
Corresponding authors
Ethics declarations
Ethics approval
Renal tissue samples were acquired from the Kidney Biobank of the Department of Urology at the Chinese PLA General Hospital during the period spanning January to December 2019. The study protocol involving human specimens received ethical approval from the Chinese PLA General Hospital Ethics Committee (Approval No. S2013-065-01). Confirming The study is reported in accordance with ARRIVE guidelines. All methods were carried out in accordance with relevant guidelines and regulations.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Feng, Y., Cao, S., Cai, T. et al. Gamma-aminobutyric acid transaminase mediates tumor suppression in renal cell carcinoma through the cGAS-STING–interferon-β axis. Sci Rep (2026). https://doi.org/10.1038/s41598-026-42861-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-026-42861-w